Abstract
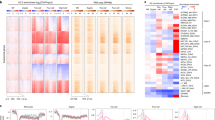
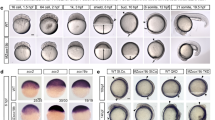
After fertilization the embryonic genome is inactive until transcription is initiated during the maternal–zygotic transition1,2,3. This transition coincides with the formation of pluripotent cells, which in mammals can be used to generate embryonic stem cells. To study the changes in chromatin structure that accompany pluripotency and genome activation, we mapped the genomic locations of histone H3 molecules bearing lysine trimethylation modifications before and after the maternal–zygotic transition in zebrafish. Histone H3 lysine 27 trimethylation (H3K27me3), which is repressive, and H3K4me3, which is activating, were not detected before the transition. After genome activation, more than 80% of genes were marked by H3K4me3, including many inactive developmental regulatory genes that were also marked by H3K27me3. Sequential chromatin immunoprecipitation demonstrated that the same promoter regions had both trimethylation marks. Such bivalent chromatin domains also exist in embryonic stem cells and are thought to poise genes for activation while keeping them repressed4,5,6,7,8. Furthermore, we found many inactive genes that were uniquely marked by H3K4me3. Despite this activating modification, these monovalent genes were neither expressed nor stably bound by RNA polymerase II. Inspection of published data sets revealed similar monovalent domains in embryonic stem cells. Moreover, H3K4me3 marks could form in the absence of both sequence-specific transcriptional activators and stable association of RNA polymerase II, as indicated by the analysis of an inducible transgene. These results indicate that bivalent and monovalent domains might poise embryonic genes for activation and that the chromatin profile associated with pluripotency is established during the maternal–zygotic transition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Newport, J. & Kirschner, M. A major developmental transition in early Xenopus embryos: II. Control of the onset of transcription. Cell 30, 687–696 (1982)
Schier, A. F. The maternal-zygotic transition: death and birth of RNAs. Science 316, 406–407 (2007)
Tadros, W. & Lipshitz, H. D. The maternal-to-zygotic transition: a play in two acts. Development 136, 3033–3042 (2009)
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006)
Azuara, V. et al. Chromatin signatures of pluripotent cell lines. Nature Cell Biol. 8, 532–538 (2006)
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007)
Zhao, X. D. et al. Whole-genome mapping of histone H3 Lys4 and 27 trimethylations reveals distinct genomic compartments in human embryonic stem cells. Cell Stem Cell 1, 286–298 (2007)
Pan, G. et al. Whole-genome analysis of histone H3 lysine 4 and lysine 27 methylation in human embryonic stem cells. Cell Stem Cell 1, 299–312 (2007)
Schier, A. F. & Talbot, W. S. Molecular genetics of axis formation in zebrafish. Annu. Rev. Genet. 39, 561–613 (2005)
Schuettengruber, B., Chourrout, D., Vervoort, M., Leblanc, B. & Cavalli, G. Genome regulation by polycomb and trithorax proteins. Cell 128, 735–745 (2007)
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007)
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007)
Song, J. S. et al. Model-based analysis of two-color arrays (MA2C). Genome Biol. 8, R178 (2007)
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007)
Akkers, R. C. et al. A hierarchy of H3K4me3 and H3K27me3 acquisition in spatial gene regulation in Xenopus embryos. Dev. Cell 17, 425–434 (2009)
Stock, J. K. et al. Ring1-mediated ubiquitination of H2A restrains poised RNA polymerase II at bivalent genes in mouse ES cells. Nature Cell Biol. 9, 1428–1435 (2007)
Core, L. J. & Lis, J. T. Transcription regulation through promoter-proximal pausing of RNA polymerase II. Science 319, 1791–1792 (2008)
Guenther, M. G., Levine, S. S., Boyer, L. A., Jaenisch, R. & Young, R. A. A chromatin landmark and transcription initiation at most promoters in human cells. Cell 130, 77–88 (2007)
Zeitlinger, J. et al. RNA polymerase stalling at developmental control genes in the Drosophila melanogaster embryo. Nature Genet. 39, 1512–1516 (2007)
Muse, G. W. et al. RNA polymerase is poised for activation across the genome. Nature Genet. 39, 1507–1511 (2007)
Boettiger, A. N. & Levine, M. Synchronous and stochastic patterns of gene activation in the Drosophila embryo. Science 325, 471–473 (2009)
Dreijerink, K. M. et al. Menin links estrogen receptor activation to histone H3K4 trimethylation. Cancer Res. 66, 4929–4935 (2006)
Vermeulen, M. et al. Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell 131, 58–69 (2007)
Hammoud, S. S. et al. Distinctive chromatin in human sperm packages genes for embryo development. Nature 460, 473–478 (2009)
Sha, K. & Boyer, L. A. The chromatin signature of pluripotent cells. The Stem Cell Research Community, StemBook. 10.3824/stembook.1.45.1 〈http://www.stembook.org〉 (2009)
Yuzyuk, T., Fakhouri, T. H., Kiefer, J. & Mango, S. E. The polycomb complex protein mes-2/E(z) promotes the transition from developmental plasticity to differentiation in C. elegans embryos. Dev. Cell 16, 699–710 (2009)
Barski, A. et al. Chromatin poises miRNA- and protein-coding genes for expression. Genome Res. 19, 1742–1751 (2009)
Wardle, F. C. et al. Zebrafish promoter microarrays identify actively transcribed embryonic genes. Genome Biol. 7, R71 (2006)
O’Geen, H., Nicolet, C. M., Blahnik, K., Green, R. & Farnham, P. J. Comparison of sample preparation methods for ChIP-chip assays. Biotechniques 41, 577–580 (2006)
Phatnani, H. P. & Greenleaf, A. L. Phosphorylation and functions of the RNA polymerase II CTD. Genes Dev. 20, 2922–2936 (2006)
Sagasti, A., Guido, M. R., Raible, D. W. & Schier, A. F. Repulsive interactions shape the morphologies and functional arrangement of zebrafish peripheral sensory arbors. Curr. Biol. 15, 804–814 (2005)
Link, V., Shevchenko, A. & Heisenberg, C. Proteomics of early zebrafish embryos. BMC Dev. Biol. 6, 1 (2006)
Shin, H., Liu, T., Manrai, A. K. & Liu, X. S. CEAS: Cis-regulatory Element Annotation System. Bioinformatics 25, 2605–2606 (2009)
Dennis, G. et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 4, 3 (2003)
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols 4, 44–57 (2009)
Acknowledgements
We thank members of the Schier laboratory for help and advice; H. G. Shin, L. Taing and Z. J. Wu for computational analysis and discussions; N. Follmer and B. Lilley for technical advice; and J. Dubrulle, N. Francis, R. Losick, S. Mango, T. van Opijnen and W. Talbot for discussions and critical reading of the manuscript. This work was supported by NIH grants to X.S.L. (1R01 HG004069) and A.F.S. (5R01 GM56211), and by EMBO and HFSP (LT-00090/2007) fellowships to N.L.V.
Author Contributions N.L.V. and A.F.S. designed the study. N.L.V. performed the experiments. Y.Z. performed computational analysis. N.L.V., Y.Z., J.R., X.S.L. and A.F.S. designed and performed data analysis. I.G.W. provided technical support. F.I. provided RNA profiling data. A.R. provided analytical advice. N.L.V. and A.F.S. interpreted the data and wrote the paper with support from co-authors.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Discussions 1-5, Supplementary Figures S1-S9 with legends and Supplementary References. (PDF 1626 kb)
Supplementary Table 1
This file contains Supplementary Table 1, which includes the list of analyzed genes and their status for H3K4me3, H3K27me3, H3K36me3 and RNA polymerase II in zebrafish blastomeres. (XLS 110 kb)
Rights and permissions
About this article
Cite this article
Vastenhouw, N., Zhang, Y., Woods, I. et al. Chromatin signature of embryonic pluripotency is established during genome activation. Nature 464, 922–926 (2010). https://doi.org/10.1038/nature08866
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08866
This article is cited by
-
Regulation, functions and transmission of bivalent chromatin during mammalian development
Nature Reviews Molecular Cell Biology (2023)
-
Endothelial Brg1 fine-tunes Notch signaling during zebrafish heart regeneration
npj Regenerative Medicine (2023)
-
Pluripotency factors determine gene expression repertoire at zygotic genome activation
Nature Communications (2022)
-
Context-specific Polycomb mechanisms in development
Nature Reviews Genetics (2022)
-
Insights into epigenetic patterns in mammalian early embryos
Protein & Cell (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.