Abstract
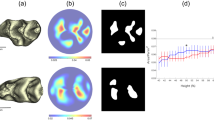
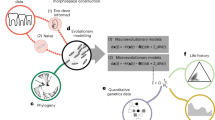
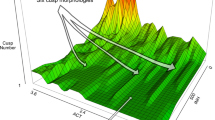
The relationship between the genotype and the phenotype, or the genotype–phenotype map, is generally approached with the tools of multivariate quantitative genetics and morphometrics1,2,3,4. Whereas studies of development5,6,7 and mathematical models of development4,8,9,10,11,12 may offer new insights into the genotype–phenotype map, the challenge is to make them useful at the level of microevolution. Here we report a computational model of mammalian tooth development that combines parameters of genetic and cellular interactions to produce a three-dimensional tooth from a simple tooth primordia. We systematically tinkered with each of the model parameters to generate phenotypic variation and used geometric morphometric analyses to identify, or developmentally ordinate, parameters best explaining population-level variation of real teeth. To model the full range of developmentally possible morphologies, we used a population sample of ringed seals (Phoca hispida ladogensis)13. Seal dentitions show a high degree of variation, typically linked to the lack of exact occlusion13,14,15,16. Our model suggests that despite the complexity of development and teeth, there may be a simple basis for dental variation. Changes in single parameters regulating signalling during cusp development may explain shape variation among individuals, whereas a parameter regulating epithelial growth may explain serial, tooth-to-tooth variation along the jaw. Our study provides a step towards integrating the genotype, development and the phenotype.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Roff, D. A. A centennial celebration for quantitative genetics. Evolution 61, 1017–1032 (2007)
Zeng, Z. B. QTL mapping and the genetic basis of adaptation: recent developments. Genetica 123, 25–37 (2005)
Wagner, G. P. et al. Pleiotropic scaling of gene effects and the ‘cost of complexity’. Nature 452, 470–472 (2008)
Polly, P. D. Developmental dynamics and G-matrices: can morphometric spaces be used to model phenotypic evolution? Evol. Biol. 35, 83–96 (2008)
Prud'homme, B., Gompel, N. & Carroll, S. B. Emerging principles of regulatory evolution. Proc. Natl Acad. Sci. USA 104, 8605–8612 (2007)
McGregor, A. P. et al. Morphological evolution through multiple cis-regulatory mutations at a single gene. Nature 448, 587–590 (2007)
Miller, C. T. et al. cis-Regulatory changes in Kit ligand expression and parallel evolution of pigmentation in sticklebacks and humans. Cell 131, 1179–1189 (2007)
Salazar-Ciudad, I., Newman, S. A. & Solé, R. V. Phenotypic and dynamical transitions in model genetic networks. I. Emergence of patterns and genotype-phenotype relationships. Evol. Dev. 3, 84–94 (2001)
Harris, M. P., Williamson, S., Fallon, J. F., Meinhardt, H. & Prum, R. O. Molecular evidence for an activator-inhibitor mechanism in development of embryonic feather branching. Proc. Natl Acad. Sci. USA 102, 11734–11739 (2005)
Evans, T. M. & Marcus, J. M. A simulation study of the genetic regulatory hierarchy for butterfly eyespot focus determination. Evol. Dev. 8, 273–283 (2006)
Kavanagh, K. D., Evans, A. R. & Jernvall, J. Predicting evolutionary patterns of mammalian teeth from development. Nature 449, 427–432 (2007)
Nahmad, M., Glass, L. & Abouheif, E. The dynamics of developmental system drift in the gene network underlying wing polyphenism in ants: a mathematical model. Evol. Dev. 10, 360–374 (2008)
Jernvall, J. Linking development with generation of novelty in mammalian teeth. Proc. Natl Acad. Sci. USA 97, 2641–2645 (2000)
Lönnberg, E. Några egendomliga variationer i tanduppsättningen hos vikaren, Phoca hispida . Fauna och Flora 18, 116–126 (1923)
Cruwys, L. & Friday, A. Visible supernumerary teeth in pinnipeds. Polar Rec. 42, 83–85 (2006)
Miller, E. H. et al. Variation and integration of the simple mandibular postcanine dentition in two species of phocid seal. J. Mamm. 88, 1325–1334 (2007)
Jernvall, J., Keränen, S. V. E. & Thesleff, I. Evolutionary modification of development in mammalian teeth: Quantifying gene expression patterns and topography. Proc. Natl Acad. Sci. USA 97, 14444–14448 (2000)
Kassai, Y. et al. Regulation of mammalian tooth cusp patterning by ectodin. Science 309, 2067–2070 (2005)
Salazar-Ciudad, I. & Jernvall, J. A gene network model accounting for development and evolution of mammalian teeth. Proc. Natl Acad. Sci. USA 99, 8116–8120 (2002)
Osborn, J. W. A model of growth restraints to explain the development and evolution of tooth shapes in mammals. J. Theor. Biol. 255, 338–343 (2008)
Forgacs, G. & Newman, S. A. Biological Physics of the Developing Embryo (Cambridge Univ. Press, 2005)
Keller, R. Mechanisms of elongation in embryogenesis. Development 133, 2291–2302 (2006)
Lecuit, T. & Lenne, P. F. Cell surface mechanics and the control of cell shape, tissue patterns and morphogenesis. Nature Rev. Mol. Cell Biol. 8, 633–644 (2007)
Kettunen, P. et al. Associations of FGF-3 and FGF-10 with signaling networks regulating tooth morphogenesis. Dev. Dyn. 219, 322–332 (2000)
Rybczynski, N., Dawson, M. R. & Tedford, R. H. A semi-aquatic Arctic mammalian carnivore from the Miocene epoch and origin of Pinnipedia. Nature 458, 1021–1024 (2009)
Luo, Z.-X. Transformation and diversification in early mammal evolution. Nature 450, 1011–1019 (2007)
Stewart, B. E., Innes, S. & Stewart, R. E. A. Mandibular dental ontogeny of ringed seals (Phoca hispida). Mar. Mamm. Sci. 14, 221–231 (1998)
Fraser, G. J. et al. An ancient gene network is co-opted for teeth on old and new jaws. PLoS Biol. 7, e1000031 (2009)
Buchanan, A. V., Sholtis, S., Richtsmeier, J. & Weiss, K. M. What are genes “for” or where are traits “from”? What is the question? Bioessays 31, 198–208 (2009)
Darwin, C. On the Origin of Species by Means of Natural Selection 1st edn (Murray, 1859)
Salazar-Ciudad, I., Garcia-Fernández, J. & Solé, R. V. Gene networks capable of pattern formation: from induction to reaction-diffusion. J. Theor. Biol. 205, 587–603 (2000)
Jaeger, J. et al. Dynamic control of positional information in the early Drosophila embryo. Nature 430, 368–371 (2004)
Newman, T. J. Modeling multi-cellular systems using sub-cellular elements. Math. Biosci. Eng. 2, 611–622 (2005)
Honda, H., Motosugi, N., Nagai, T., Tanemura, M. & Hiiragi, T. Computer simulation of emerging asymmetry in the mouse blastocyst. Development 135, 1407–1414 (2008)
Graner, F. & Glazier, J. A. Simulation of biological cell sorting using a two dimensional extended Potts model. Phys. Rev. Lett. 69, 2013–2016 (1992)
Hammer, Ø. & Bucher, H. Models for the morphogenesis of the molluscan shell. Lethaia 38, 111–122 (2005)
Butler, P. M. The ontogeny of molar pattern. Biol. Rev. Camb. Philos. Soc. 31, 30–69 (1956)
Jernvall, J. Mammalian molar cusp patterns: developmental mechanisms of diversity. Acta Zool. Fenn. 198, 1–61 (1995)
Åberg, T., Wozney, J. & Thesleff, I. Expression patterns of bone morphogenetic proteins (Bmps) in the developing mouse tooth suggest roles in morphogenesis and cell differentiation. Dev. Dyn. 210, 383–396 (1997)
Acknowledgements
We thank I. Corfe, A. R. Evans, J. Fierst, M. Fortelius, B. Julia, S. Sova, J. Hakanen, E. Harjunmaa, N. Navarro, I. Thesleff, P. C. Wright and S. Zohdy for comments, discussions, and support on this work, A. Kangas for help in data collection, and M. Hildén and I. Hanski (Finnish Museum of Natural History) for access to collections. This study was supported by the Ramón y Cajal Program (RYC-2007-00149) and the Academy of Finland.
Author Contributions I.S.-C. and J.J. conceived the study; I.S.-C. constructed the computational model and performed computer simulations; J.J. obtained the empirical data; I.S.-C. and J.J. performed quantitative analyses and wrote the paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-6 with legends, and Supplementary Tables 1-6. (PDF 580 kb)
Rights and permissions
About this article
Cite this article
Salazar-Ciudad, I., Jernvall, J. A computational model of teeth and the developmental origins of morphological variation. Nature 464, 583–586 (2010). https://doi.org/10.1038/nature08838
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08838
This article is cited by
-
Rules of teeth development align microevolution with macroevolution in extant and extinct primates
Nature Ecology & Evolution (2023)
-
Stripe and spot selection in cusp patterning of mammalian molar formation
Scientific Reports (2022)
-
Identifying biological affinities of Holocene northern Iberian populations through the inner structures of the upper first molars
Archaeological and Anthropological Sciences (2022)
-
Using phenotypic plasticity to understand the structure and evolution of the genotype–phenotype map
Genetica (2022)
-
A universal power law for modelling the growth and form of teeth, claws, horns, thorns, beaks, and shells
BMC Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.