Abstract
The zebra finch is an important model organism in several fields1,2 with unique relevance to human neuroscience3,4. Like other songbirds, the zebra finch communicates through learned vocalizations, an ability otherwise documented only in humans and a few other animals and lacking in the chicken5—the only bird with a sequenced genome until now6. Here we present a structural, functional and comparative analysis of the genome sequence of the zebra finch (Taeniopygia guttata), which is a songbird belonging to the large avian order Passeriformes7. We find that the overall structures of the genomes are similar in zebra finch and chicken, but they differ in many intrachromosomal rearrangements, lineage-specific gene family expansions, the number of long-terminal-repeat-based retrotransposons, and mechanisms of sex chromosome dosage compensation. We show that song behaviour engages gene regulatory networks in the zebra finch brain, altering the expression of long non-coding RNAs, microRNAs, transcription factors and their targets. We also show evidence for rapid molecular evolution in the songbird lineage of genes that are regulated during song experience. These results indicate an active involvement of the genome in neural processes underlying vocal communication and identify potential genetic substrates for the evolution and regulation of this behaviour.
Similar content being viewed by others
Main
As in all songbirds, singing in the zebra finch is under the control of a discrete neural circuit that includes several dedicated centres in the forebrain termed the ‘song control nuclei’ (for an extensive series of reviews see ref. 8). Neurophysiological studies in these nuclei during singing have yielded some of the most illuminating examples of how vocalizations are encoded in the motor system of a vertebrate brain9,10. In the zebra finch, these nuclei develop more fully in the male than in the female (who does not sing), and they change markedly in size and organization during the juvenile period when the male learns to sing11. Analysis of the underlying cellular mechanisms of plasticity led to the unexpected discovery of neurogenesis in adult songbirds and life-long replacement of neurons12. Sex steroid hormones also contribute to songbird neural plasticity, in part by influencing the survival of new neurons13. Some of these effects are probably caused by oestrogen and/or testosterone synthesized within the brain itself rather than just in the gonads14.
Song perception and memory also involve auditory centres that are present in both sexes, and the mere experience of hearing a song activates gene expression in these auditory centres15. The gene response itself changes as a song becomes familiar over the course of a day16 or as the context of the experience changes17. The act of singing induces gene expression in the male song control nuclei, and these patterns of gene activation also vary with the context of the experience18. The function of this changing genomic activity is not yet understood, but it may support or suppress learning and help integrate information over periods of hours to days19.
The chicken genome is the only other bird genome analysed to date6. The chicken and zebra finch lineages diverged about 100 million years ago near the base of the avian radiation7. By comparing their genomes we can now discern features that are shared (and thus generally characteristic of birds), and features that are most conspicuously different between the two lineages—some of which will be related to the distinctive neural and behavioural traits of songbirds.
We sequenced and assembled a male zebra finch genome using methods described previously6,20. A male (the homogametic sex in birds) was chosen to maximize coverage of the Z chromosome. Of the 1.2 gigabase (Gb) draft assembly, 1.0 Gb has been assigned to 33 chromosomes and three linkage groups, by using zebra finch genetic linkage21 and bacterial artificial chromosome (BAC) fingerprint maps. The genome assembly is of sufficient quality for the analysis presented here (see Supplementary Note 1 and Supplementary Table 1). A total of 17,475 protein-coding genes were predicted from the zebra finch genome assembly using the Ensembl pipeline supplemented by Gpipe gene models (Supplementary Note 1). To extend further the characterization of genes relevant to brain and behaviour, we also sequenced complementary DNAs from the forebrain of zebra finches at 50 (juvenile, during the critical song learning period) and 850 (adult) days post-hatch, mapping these reads (Illumina GA2) to the protein-coding models (Supplementary Note 1). Of the 17,475 protein-coding gene models we find 9,872 (56%) and 10,106 (57%) genes expressed in the forebrain at these two ages (90.7% overlap), respectively. In addition to evidence for developmental regulation, these reads show further splice forms, new exons and untranslated sequences (Supplementary Figs 1 and 2).
To address issues of large-scale genome structure and evolution, we compared the chromosomes of zebra finch and chicken using both sequence alignment and fluorescent in situ hybridization. These analyses showed overall conservation of synteny and karyotype in the two species, although the rate of intrachromosomal rearrangement was high (Supplementary Note 2). We were also surprised to see genes of the major histocompatibility complex (MHC) dispersed across several chromosomes in the zebra finch, in contrast to the syntenic organization of both chicken and human MHCs (Supplementary Note 2).
We assessed specific gene losses and expansions in the zebra finch lineage by constructing phylogenies of genes present in the last common ancestor of birds and mammals (Supplementary Note 2 and Supplementary Fig. 3). Both the zebra finch and the chicken genome assemblies lack genes encoding vomeronasal receptors, casein milk proteins, salivary-associated proteins and enamel proteins—not surprisingly, as birds lack vomeronasal organs, mammary glands and teeth. Unexpectedly, both species lack the gene for the neuronal protein synapsin 1 (SYN1); comparative analyses suggest that the loss of SYN1 and flanking genes probably occurred in an ancestor to modern birds, possibly within the dinosaur lineage (Supplementary Note 2, Supplementary Table 2 and Supplementary Fig. 4). Both zebra finch and chicken have extensive repertoires of olfactory receptor-like sequences (Supplementary Note 2 and Supplementary Fig. 5), proteases (Supplementary Table 3), and a rich repertoire of neuropeptide and pro-hormone genes.
Compared to mammals, zebra finch has duplications of genes encoding several proteins with known neural functions, including growth hormone, (Supplementary Fig. 3), caspase-3 and β-secretase (Supplementary Table 3). Two large expansions of gene families expressed in the brain seem to have occurred in the zebra finch lineage after the split from mammals. One involves a family related to the PAK3 (p21-activated kinase) gene. Thirty-one uninterrupted PAK3-like sequences have been identified in the zebra finch genome, of which 29 are expressed in testis and/or brain (Supplementary Note 2). The second involves the PHF7 gene, which encodes a zinc-finger-containing transcriptional control protein. Humans only have a single PHF7 gene, but remarkably the gene has been duplicated independently, many times in both the zebra finch and chicken lineages to form species-specific clades of 17 and 18 genes, respectively (Supplementary Fig. 6). In the zebra finch these genes are expressed in the brain (Supplementary Note 2).
An intriguing puzzle in avian genomics has been the evident lack of a chromosome-wide dosage compensation mechanism to balance the expression of genes on the Z sex chromosome, which is present in two copies in males but only one in females22,23. The chicken has been suspected of exerting dosage compensation on a more local level, by the non-coding RNA MHM (male hypermethylated)24,25, to cause a characteristic variation of gene expression along the Z chromosome. The zebra finch genome assembly, however, lacks an MHM sequence, and genes adjacent to the comparable MHM chromosomal position show no special cluster of dosage compensation (Fig. 1 and Supplementary Note 2). Thus, the putative MHM-mediated mechanism of restricted Z-chromosome dosage compensation is not common to all birds. Chromosomal sex differences in the brain could have a direct role in the sex differences so evident in zebra finch neuroanatomy and singing behaviour.
a, b, The male to female (M/F) ratio of gene expression, measured by species-specific microarrays, is plotted along the Z chromosome of chicken (a) and zebra finch (b). Each point represents the average M/F ratio of a sliding window of 30 genes plotted at the median gene position and stepping one gene at a time along the chromosome. Note region of lower M/F ratios in chicken surrounding the locus of the MHM (male hypermethylated) ncRNA. In zebra finch, genes adjacent to the comparable MHM position (asterisk) show no special cluster of dosage compensation (low M/F ratios), and no MHM sequence appears in the genome assembly. bp, base pairs.
In mammals, as much as half of their genomes represent interspersed repeats derived from mobile elements, whereas the interspersed repeat content of the chicken genome is only 8.5%. We find that the zebra finch genome also has a low overall interspersed repeat content (7.7%), containing a little over 200,000 mobile elements (Supplementary Tables 4 and 5). The zebra finch, however, has about three times as many retrovirus-derived long terminal repeat (LTR) element copies as the chicken, and a low copy number of short interspersed elements (SINEs), which the chicken lacks altogether. Expressed sequence tag (EST) analysis shows that mobile elements are present in about 4% of the transcripts expressed in the zebra finch brain, and some of these transcripts are regulated by song exposure (next section, Table 1). Figure 2 shows an example of an RNA that was identified in a microarray screening for genes specifically enriched in song control nuclei26 and now seems to represent a long non-coding RNA (ncRNA) containing a CR1-like mobile element. These results indicate that further experiments investigating a possible role of mobile-element-derived repeated sequences in vocal communication are warranted.
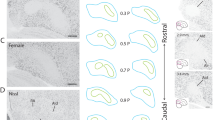
a, Genomic alignment of an RNA containing a CR1-like retrotransposon element (in blue) and adjacent ESTs, with respective GenBank accession numbers. b–d, DV949717 is expressed in the brain of adult males with enrichment in song nuclei HVC (letter-based name) and LMAN (lateral magnocellular nucleus of the anterior nidopallium), as revealed by in situ hybridization. The diagram in b indicates areas shown in photomicrographs in c and d. Cb, cerebellum; Hp, hippocampus; Meso, mesopallium; Nido, nidopallium; Shelf, nidopallial shelf region; St, striatum. Scale bars, 0.1 mm.
A large portion of the genome is directly engaged by vocal communication. A recent study27 defined distinct sets of RNAs in the auditory forebrain that respond in different ways to song playbacks during the process of song-specific habituation, a form of learning16. We now map each of these song-responsive RNAs to the genome assembly (Table 1 and Supplementary Note 3). Notably, we find evidence that ∼40% of transcripts in the unstimulated auditory forebrain are non-coding and derive from intronic or intergenic loci (Table 1). Among the RNAs that are rapidly suppressed in response to new vocal signals (‘novel down’), two-thirds are ncRNAs.
The robust involvement of ncRNAs in the response to song led us to ask whether song exposure alters the expression of microRNAs—small ncRNAs that regulate gene expression by binding to target messenger RNAs. Indeed we find that miR-124, a conserved microRNA implicated in neurological function in other species28, is rapidly suppressed in response to song playbacks (Fig. 3). We independently measured this effect by direct Illumina sequencing of small RNAs in the auditory forebrain, and also identified other known and new microRNAs, several of which also change in expression after song stimulation (Supplementary Note 2).
TaqMan assays comparing samples from the auditory lobule of adult male zebra finches in silence (open bars) or 30 min after onset of new song playback (filled bars). a, Comparison of two sample pools, each containing auditory forebrains of 20 birds. b, Comparisons of paired individual subjects, n = 6 pairs (P = 0.03, Wilcoxon paired test). Error bars denote s.e.m. of triplicate TaqMan assays. Parallel TaqMan analyses of the small RNA RNU6B were performed with all samples and showed no significant effect of treatment for this control RNA.
A potential site of action for microRNAs was shown by genomic mapping of transcripts that increase rapidly after new song exposure (Table 1, ‘novel up’). Two of the cDNA clones that measured the most robust increases27 align to an unusually long (3 kilobases (kb)) 3′ untranslated region (UTR) in the human gene that encodes the NR4A3 transcription factor protein (Fig. 4a). The entire UTR is similar in humans and zebra finches, with several long segments of >80% identity (Fig. 4b). Within these segments we find conserved predicted binding sites for 11 different microRNAs, including five new microRNAs found by direct sequencing of small RNAs from the zebra finch forebrain (Fig. 4b). These findings indicate that this NR4A3 transcript element may function in both humans and songbirds to integrate many conserved microRNA regulatory pathways.
a, zPicture alignment of 3′ portion of zebra finch to human gene35 showing UTR region of high similarity beyond the coding exons. Dark red bars, regions with the highest sequence conservation; black rectangles, position of song-regulated ESTs27 within the conserved UTR but outside the Ensembl gene model (ENSTGUG00000008853). b, Alignment of zebra finch and human 3′ UTR sequences showing the per cent sequence identity for each evolutionarily conserved region. Dots indicate positions of conserved new (‘n-’) or established (‘miR-’) microRNA-binding sites in both species within these regions.
The act of singing also alters gene expression in song control nuclei29, and we used the genome assembly to analyse the transcriptional control structure of this response. Using oligonucleotide microarrays, we identified 807 genes in which expression significantly changed as a result of singing. These were grouped by k-means clustering into 20 distinct expression profile clusters (Fig. 5a and Supplementary Note 3). Gene regulatory sequences (transcription-factor-binding sites) were predicted across the genome using a new motif-scanning approach (Supplementary Note 1), and we observed significant correlation between changes in expression of transcription factor genes and their predicted targets (Fig. 5b and Supplementary Table 6). Thus, the experience of singing and hearing song engages complex gene regulatory networks in the forebrain, altering the expression of microRNAs, transcription factor genes, and their targets, as well as of non-coding RNA elements that may integrate transcriptional and post-transcriptional control systems.
a, Clustered (1–20) temporal expression profiles of 807 genes (rows) that change with time and amount of singing; red, increases; blue, decreases; white, no change relative to average 0-h control. Grey/coloured bars on left, clusters with enrichment of specific promoter motifs (P < 0.01). b, Enriched transcription-factor-binding motifs (abbreviations) found in the promoters of late response genes, clusters 9–12 (coloured as in a); bold, binding sites for known activity-dependent transcription factors (for example, CREBP1) or transcription factor complexes (for example, CREBP1–CJUN); black, sites for post-translationally activated transcription factors; brown, sites for transcriptionally activated transcription factors including by singing (for example, in cluster 1). Graph shows time course of average expression of all genes in the late response clusters, normalized to average 0 h for that cluster. Also plotted is the average expression of the C-FOS transcription factor mRNA, which binds to the AP-1 site over-represented in the promoters of cluster 10 genes.
Learned vocal communication is crucial to the reproductive success of a songbird, and this behaviour evolved after divergence of the songbird lineage5. Thus, it seems likely that genes involved in the neurobiology of vocal communication have been influenced by positive selection in songbirds. With this in mind, we examined the intersection of two sets of genes: (1) those that respond to song exposure in the auditory forebrain as discussed in the previous section; and (2) those that contain residues that seem to have been positively selected in the zebra finch lineage, as determined using phylogenetic analysis by maximum likelihood (PAML) (Supplementary Note 4). There are 214 genes that are common to both lists. Of these, 49 are suppressed by song exposure (Supplementary Table 7), and 6 of these 49 are explicitly annotated for ion channel activity (Table 2). This yields a highly significant statistical enrichment for the term ‘ion channel activity’ (P = 0.0016, false discovery rate (FDR) adjusted Fisher’s exact test) and other related terms in this subset of genes (Supplementary Tables 8 and 9). Independent evidence has also demonstrated differential anatomical expression of ion channel genes in song control nuclei26,30. Ion channel genes have important roles in many aspects of behaviour, neurological function and disease31. This class of genes is highly likely to be linked to song behaviour and should be a major target for future functional studies.
Passerines represent one of the most successful and complex radiations of terrestrial animals7. Here we present the first, to our knowledge, analysis of the genome of a passerine bird. The zebra finch was chosen because of its well-developed status as a model organism for a number of fields in biology, including neurobiology, ethology, ecology, biogeography and evolution. In the zebra finch as in the chicken, we see a smaller, tighter genome compared to mammals, with a marked reduction of interspersed repeats. The zebra finch presents a picture of greater genomic plasticity than might have been expected from the chicken and other precedents, with a high degree of intrachromosomal rearrangements between the two avian species, gene copy number variations and transcribed mobile elements. Yet we also see an overall similarity to mammals in protein-coding gene content and core transcriptional control systems.
Our analysis suggests several channels through which evolution may have acted to produce the unique neurobiological properties of songbirds compared to the chicken and other animals. These include the management of sex chromosome gene expression, accelerated evolution of neuronal ion transport genes, gene duplications to produce new variants of PHF7, PAK3 and other neurobiologically important genes, and a new arrangement of MHC genes. Most notably, our analyses suggest a large recruitment of the genome during vocal communication, including the extensive involvement of ncRNAs. It has been proposed that ncRNAs have a contributing role in enabling or driving the evolution of greater complexity in humans and other complex eukaryotes32. Seeing that learned vocal communication itself is a phenomenon that has emerged only in some of the most complex organisms, perhaps ncRNAs are a nexus of this phenomenon.
Much work will be needed to establish the actual functional significance of many of these observations and to determine when they arose in avian evolution. This work can now be expedited with the recent development of a method for transgenesis in the zebra finch33. An important general lesson, however, is that dynamic and serendipitous aspects of the genome may have unexpected roles in the elaborate vocal communicative capabilities of songbirds.
Methods Summary
Sequence assembly
Sequenced reads were assembled and attempts were made to assign the largest contiguous blocks of sequence to chromosomes using a genetic linkage map21, fingerprint map and synteny with the chicken genome assembly Gallus_gallus-2.1, a revised version of the original draft6 (Supplementary Note 1).
Genes
Gene orthology assignment was performed using the EnsemblCompara GeneTrees pipeline and the OPTIC pipeline (Supplementary Note 1). Orthology rate estimation was performed with PAML (pairwise model = 0, Nssites = 0). In all cases, codon frequencies were estimated from the nucleotide composition at each codon position (F3X4 model).
Gene expression and evolution
Methods for Illumina read counting, in situ hybridization, TaqMan RT–PCR, microarrays, regulatory motif and evolutionary rate analyses are given in Supplementary Notes 1–4.
Accession codes
Primary accessions
DDBJ/GenBank/EMBL
Data deposits
The Taeniopygia guttata whole-genome shotgun project has been deposited in DDBJ/EMBL/GenBank under the project accession ABQF00000000.
References
Zann, R. A. The Zebra Finch: A Synthesis of Field and Laboratory Studies (Oxford Univ, Press, 1996)
Clayton, D. F., Balakrishnan, C. N. & London, S. E. Integrating genomes, brain and behavior in the study of songbirds. Curr. Biol. 19, R865–R873 (2009)
Nottebohm, F. in Hope For a New Neurology (ed. Nottebohm, F.) (New York Academy of Science, 1985)
Doupe, A. J. & Kuhl, P. K. Birdsong and human speech: common themes and mechanisms. Annu. Rev. Neurosci. 22, 567–631 (1999)
Jarvis, E. D. Learned birdsong and the neurobiology of human language. Ann. NY Acad. Sci. 1016, 749–777 (2004)
Hillier, L. W. et al. Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 432, 695–716 (2004)
Hackett, S. J. et al. A phylogenomic study of birds reveals their evolutionary history. Science 320, 1763–1768 (2008)
Zeigler, H. P. & Marler, P. Behavioral Neurobiology of Bird Song Vol. 1016 (New York Academy of Sciences, 2004)
Hahnloser, R. H., Kozhevnikov, A. A. & Fee, M. S. An ultra-sparse code underlies the generation of neural sequences in a songbird. Nature 419, 65–70 (2002)
Mooney, R. Neural mechanisms for learned birdsong. Learn. Mem. 16, 655–669 (2009)
Konishi, M. & Akutagawa, E. Neuronal growth, atrophy and death in a sexually dimorphic song nucleus in the zebra finch brain. Nature 315, 145–147 (1985)
Goldman, S. A. & Nottebohm, F. Neuronal production, migration, and differentiation in a vocal control nucleus of the adult female canary brain. Proc. Natl Acad. Sci. USA 80, 2390–2394 (1983)
Nottebohm, F. The road we travelled: discovery, choreography, and significance of brain replaceable neurons. Ann. NY Acad. Sci. 1016, 628–658 (2004)
London, S. E., Remage-Healey, L. & Schlinger, B. A. Neurosteroid production in the songbird brain: A re-evaluation of core principles. Front. Neuroendocrinol. 30, 302–314 (2009)
Mello, C. V., Vicario, D. S. & Clayton, D. F. Song presentation induces gene expression in the songbird forebrain. Proc. Natl Acad. Sci. USA 89, 6818–6822 (1992)
Dong, S. & Clayton, D. F. Habituation in songbirds. Neurobiol. Learn. Mem. 92, 183–188 (2009)
Woolley, S. C. & Doupe, A. J. Social context-induced song variation affects female behavior and gene expression. PLoS Biol. 6, e62 (2008)
Jarvis, E. D., Scharff, C., Grossman, M. R., Ramos, J. A. & Nottebohm, F. For whom the bird sings: context-dependent gene expression. Neuron 21, 775–788 (1998)
Clayton, D. F. The genomic action potential. Neurobiol. Learn. Mem. 74, 185–216 (2000)
Warren, W. C. et al. Genome analysis of the platypus reveals unique signatures of evolution. Nature 453, 175–183 (2008)
Stapley, J., Birkhead, T. R., Burke, T. & Slate, J. A linkage map of the zebra finch Taeniopygia guttata provides new insights into avian genome evolution. Genetics 179, 651–667 (2008)
Itoh, Y. et al. Dosage compensation is less effective in birds than in mammals. J. Biol. 6, 2 (2007)
Ellegren, H. et al. Faced with inequality: chicken do not have a general dosage compensation of sex-linked genes. BMC Biol. 5, 40 (2007)
Teranishi, M. et al. Transcripts of the MHM region on the chicken Z chromosome accumulate as non-coding RNA in the nucleus of female cells adjacent to the DMRT1 locus. Chromosome Res. 9, 147–165 (2001)
Arnold, A. P., Itoh, Y. & Melamed, E. A bird’s-eye view of sex chromosome dosage compensation. Annu. Rev. Genomics Hum. Genet. 9, 109–127 (2008)
Lovell, P. V., Clayton, D. F., Replogle, K. L. & Mello, C. V. Birdsong “transcriptomics”: neurochemical specializations of the oscine song system. PLoS One 3, e3440 (2008)
Dong, S. et al. Discrete molecular states in the brain accompany changing responses to a vocal signal. Proc. Natl Acad. Sci. USA 106, 11364–11369 (2009)
Makeyev, E. V. & Maniatis, T. Multilevel regulation of gene expression by microRNAs. Science 319, 1789–1790 (2008)
Wada, K. et al. A molecular neuroethological approach for identifying and characterizing a cascade of behaviorally regulated genes. Proc. Natl Acad. Sci. USA 103, 15212–15217 (2006)
Wada, K., Sakaguchi, H., Jarvis, E. D. & Hagiwara, M. Differential expression of glutamate receptors in avian neural pathways for learned vocalization. J. Comp. Neurol. 476, 44–64 (2004)
Cooper, E. C. & Jan, L. Y. Ion channel genes and human neurological disease: recent progress, prospects, and challenges. Proc. Natl Acad. Sci. USA 96, 4759–4766 (1999)
Mattick, J. S. RNA regulation: a new genetics? Nature Rev. Genet. 5, 316–323 (2004)
Agate, R. J., Scott, B. B., Haripal, B., Lois, C. & Nottebohm, F. Transgenic songbirds offer an opportunity to develop a genetic model for vocal learning. Proc. Natl Acad. Sci. USA 106, 17963–17967 (2009)
Replogle, K. et al. The Songbird Neurogenomics (SoNG) Initiative: community-based tools and strategies for study of brain gene function and evolution. BMC Genomics 9, 131 (2008)
Ovcharenko, I., Loots, G. G., Hardison, R. C., Miller, W. & Stubbs, L. zPicture: dynamic alignment and visualization tool for analyzing conservation profiles. Genome Res. 14, 472–477 (2004)
Acknowledgements
The sequencing of zebra finch was funded by the National Human Genome Research Institute (NHGRI). Further research support included grants to D.F.C. (NIH RO1 NS045264 and RO1 NS051820), H.E. (Swedish Research Council and Knut and Alice Wallenberg Foundation), E.D.J. (HHMI, NIH Directors Pioneer Award and R01 DC007218), M.A.B. (NIH RO1 GM59290) and J.S. (Biotechnology and Biological Sciences Research Council grant number BBE0175091). Resources for exploring the sequence and annotation data are available on browser displays available at UCSC (http://genome.ucsc.edu), Ensembl (http://www.ensembl.org), the NCBI (http://www.ncbi.nlm.nih.gov) and http://aviangenomes.org. We thank K. Lindblad-Toh for permission to use the green anole lizard genome assembly, the Production Sequencing Group of The Genome Center at Washington University School of Medicine for generating all the sequence reads used for genome assembly, and the Clemson University Genome Institute for the construction of the BAC library. We would like to recognize all the important published work that we were unable to cite owing to space limitations.
Author Contributions W.C.W., D.F.C., H.E. and A.P.A. comprise the organizing committee of the zebra finch genome sequencing project. Project planning, management and data analysis: W.C.W., D.F.C., H.E. and A.P.A. Assembly annotation and analysis: L.W.H., P.M., S.-P.Y., L.Y., J.N., A.C., S.H., J.Sl., J.St., D.B. and S.-P.Y. Protein coding and non-coding gene prediction: S.S., C.B., P.F., S.W., A.H., C.P.P. and L.K. SNP analysis: P.F. and W.M.M. Orthology prediction and analysis: A.J.V., A.H., C.P.P., S.F. and L.K. Repeat element analysis: M.A.B., A.F.A.S., R.H., M.K.K., J.A.W., W.G. and D.D.P. Segmental duplication and gene duplication analysis: L.C., Z.C., E.E.E., L.K., C.P.P., M.F., C.N.B., R.E., J.G. and S.E.L. Protease annotation and analysis: X.S.P., V.Q., G.V. and C.L.-O. Neuropeptide hormone annotation: J.Sw. and B.S. Small non-coding RNA analysis: Y-C.L., Y.L., P.G., M.W. and X.L. Comparative mapping: D.K.G., M.V. and B.M.S. Singing induced gene network analysis: E.D.J., A.R.P., O.W. and J.H. Z-chromosome analysis: Y.I. and A.P.A. Gene expression and in situ analysis and synapsin synteny/loss analysis: C.V.M., P.L. and T.A.F.V. Adaptive evolution analysis: A.K., K.N., N.B., L.S., B.N. and C.N.B. Gene expression in the brain analysis: C.S., I.A., A.S., H.L., H.R. and M.S. MHC analysis: S.E., C.N.B. and R.E. Olfactory receptor analysis: T.O., D.L. and L.K. Sequencing management: R.K.W., E.R.M. and L.F. Physical map construction: T.G. Zebra finch tissue resources: T.Bu. and T.Bi. Zebra finch cDNA resources: D.F.C., E.D.J. and X.L.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Notes S1-S4, Supplementary Tables 1-10, Supplementary Figures 1-9 with legends and Supplementary References. (PDF 2523 kb)
Supplementary Data
This file contains the annotated non-coding RNA data. (XLS 64 kb)
Supplementary Accessory Figures
This file contains Supplementary Accessory Figures 1-10 with legends. (PDF 6371 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Warren, W., Clayton, D., Ellegren, H. et al. The genome of a songbird. Nature 464, 757–762 (2010). https://doi.org/10.1038/nature08819
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08819
This article is cited by
-
High heterogeneity in genomic differentiation between phenotypically divergent songbirds: a test of mitonuclear co-introgression
Heredity (2023)
-
Integrative comparative analysis of avian chromosome evolution by in-silico mapping of the gene ontology of homologous synteny blocks and evolutionary breakpoint regions
Genetica (2023)
-
False gene and chromosome losses in genome assemblies caused by GC content variation and repeats
Genome Biology (2022)
-
Recurrent chromosome reshuffling and the evolution of neo-sex chromosomes in parrots
Nature Communications (2022)
-
Prenatal testosterone triggers long-term behavioral changes in male zebra finches: unravelling the neurogenomic mechanisms
BMC Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.