Abstract
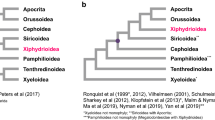
The remarkable antiquity, diversity and ecological significance of arthropods have inspired numerous attempts to resolve their deep phylogenetic history, but the results of two decades of intensive molecular phylogenetics have been mixed1,2,3,4,5,6,7. The discovery that terrestrial insects (Hexapoda) are more closely related to aquatic Crustacea than to the terrestrial centipedes and millipedes2,8 (Myriapoda) was an early, if exceptional, success. More typically, analyses based on limited samples of taxa and genes have generated results that are inconsistent, weakly supported and highly sensitive to analytical conditions7,9,10. Here we present strongly supported results from likelihood, Bayesian and parsimony analyses of over 41 kilobases of aligned DNA sequence from 62 single-copy nuclear protein-coding genes from 75 arthropod species. These species represent every major arthropod lineage, plus five species of tardigrades and onychophorans as outgroups. Our results strongly support Pancrustacea (Hexapoda plus Crustacea) but also strongly favour the traditional morphology-based Mandibulata11 (Myriapoda plus Pancrustacea) over the molecule-based Paradoxopoda (Myriapoda plus Chelicerata)2,5,12. In addition to Hexapoda, Pancrustacea includes three major extant lineages of ‘crustaceans’, each spanning a significant range of morphological disparity. These are Oligostraca (ostracods, mystacocarids, branchiurans and pentastomids), Vericrustacea (malacostracans, thecostracans, copepods and branchiopods) and Xenocarida (cephalocarids and remipedes). Finally, within Pancrustacea we identify Xenocarida as the long-sought sister group to the Hexapoda, a result confirming that ‘crustaceans’ are not monophyletic. These results provide a statistically well-supported phylogenetic framework for the largest animal phylum and represent a step towards ending the often-heated, century-long debate on arthropod relationships.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Data deposits
All sequences generated for this publication have been deposited in GenBank under the accession numbers given in Supplementary Tables 4 and 5. Full data matrices are available in Supplementary Information.
References
Abele, L. G., Kim, W. & Felgenhauer, B. E. Molecular evidence for inclusion of the phylum Pentastomida in the Crustacea. Mol. Biol. Evol. 6, 685–691 (1989)
Friedrich, M. & Tautz, D. Ribosomal DNA phylogeny of the major extant arthropod classes and the evolution of myriapods. Nature 376, 165–167 (1995)
Regier, J. C. & Shultz, J. W. Molecular phylogeny of the major arthropod groups indicates polyphyly of crustaceans and a new hypothesis for the origin of hexapods. Mol. Biol. Evol. 14, 902–913 (1997)
Giribet, G., Edgecombe, G. D. & Wheeler, W. C. Arthropod phylogeny based on eight molecular loci and morphology. Nature 413, 157–161 (2001)
Hwang, U. W., Friedrich, M., Tautz, D., Park, C. J. & Kim, W. Mitochondrial protein phylogeny joins myriapods with chelicerates. Nature 413, 154–157 (2001)
Mallatt, J. & Giribet, G. Further use of nearly complete 28S and 18S rRNA genes to classify Ecdysozoa: 37 more arthropods and a kinorhynch. Mol. Phylogenet. Evol. 40, 772–794 (2006)
Budd, G. E. & Telford, M. J. The origin and evolution of arthropods. Nature 457, 812–817 (2009)
Boore, J. L., Lavrov, D. V. & Brown, W. M. Gene translocation links insects and crustaceans. Nature 392, 667–668 (1998)
Phillips, M. J., Delsuc, F. & Penny, D. Genome-scale phylogeny and the detection of systematic biases. Mol. Biol. Evol. 21, 1455–1458 (2004)
Rota-Stabelli, O. & Telford, M. J. A multi criterion approach for the selection of optimal outgroups in phylogeny: recovering some support for Mandibulata over Myriochelata using mitogenomics. Mol. Phylogenet. Evol. 48, 103–111 (2008)
Snodgrass, R. E. Evolution of the Annelida, Onychophora and Arthropoda (Smithsonian Inst. Press, 1938)
Mallatt, J. M., Garey, J. R. & Shultz, J. W. Ecdysozoan phylogeny and Bayesian inference: first use of nearly complete 28S and 18S rRNA gene sequences to classify the arthropods and their kin. Mol. Phylogenet. Evol. 31, 178–191 (2004)
Dunn, C. W. et al. Broad phylogenomic sampling improves resolution of the animal tree of life. Nature 452, 745–749 (2008)
Timmermans, M. J. T. N., Roelofs, D., Mariën, J. & Van Straalen, N. M. Revealing pancrustacean relationships: phylogenetic analysis of ribosomal protein genes places Collembola (springtails) in a monophyletic Hexapoda and reinforces the discrepancy between mitochondrial and nuclear DNA markers. BMC Evol. Biol. 8, 83 (2008)
Philippe, H. et al. Phylogenomics revives traditional views on deep animal relationships. Curr. Biol. 19, 706–712 (2009)
Regier, J. C. et al. Resolving arthropod phylogeny: exploring phylogenetic signal within 41 kb of protein-coding nuclear gene sequence. Syst. Biol. 57, 920–938 (2008)
Holder, M. T., Zwickl, D. J. & Dessimoz, C. Evaluating the robustness of phylogenetic methods to among-site variability in substitution processes. Phil. Trans. R. Soc. B 363, 4013–4021 (2008)
Seo, T. & Kishino, H. Statistical comparison of nucleotide, amino acid, and codon substitution models for evolutionary analysis of protein-coding sequences. Syst. Biol. 58, 199–210 (2009)
Podsiadlowski, L., Braband, A. & Mayer, G. The complete mitochondrial genome of the onychophoran Epiperipatus biolleyi reveals a unique transfer RNA set and provides further support for the Ecdysozoa hypothesis. Mol. Biol. Evol. 25, 42–51 (2008)
Shultz, J. W. A phylogenetic analysis of the arachnid orders based on morphological characters. Zool. J. Linn. Soc. 150, 221–265 (2007)
Schram, F. R. ed. Crustacean Phylogeny (Balkema, 1983)
Fanenbruck, M., Harzsch, S. & Wagele, J. The brain of the Remipedia (Crustacea) and an alternative hypothesis on their phylogenetic relationships. Proc. Natl Acad. Sci. USA 101, 3868–3873 (2004)
Harzsch, S. Neurophylogeny: architecture of the nervous system and a fresh view on arthropod phyologeny. Integr. Comp. Biol. 46, 162–194 (2006)
Boxshall, G. A. Crustacean classification: on-going controversies and unresolved problems. Zootaxa 1668, 313–325 (2007)
Carapelli, A., Liò, P., Nardi, F., van der Wath, E. & Frati, F. Phylogenetic analysis of mitochondrial protein coding genes confirms the reciprocal paraphyly of Hexapoda and Crustacea. BMC Evol. Biol. 7 (suppl. 2). S8 (2007)
Hennig, W. Insect Phylogeny (Wiley, 1981)
Zrzavy, J., Hypsa, V. & Vlaskova, M. in Arthropod Relationships (eds Fortey, R. A. & Thomas, R. H.) 97–107 (Chapman and Hall, 1997)
Gai, Y.-H., Song, D., Sun, H. & Zhou, K. Myriapod monophyly and relationships among myriapod classes based on nearly complete 28S and 18S rDNA sequences. Zool. Sci. 23, 1101–1108 (2006)
Zwickl, D. J. Genetic Algorithm Approaches for the Phylogenetic Analysis of Large Biological Sequence Datasets under the Maximum Likelihood Criterion. PhD thesis, Univ. Texas Austin (2006)
Goldman, N., Thorne, J. L. & Jones, D. T. Using evolutionary trees in protein secondary structure prediction and other comparative sequence analyses. J. Mol. Biol. 263, 196–208 (1996)
Remm, M., Storm, C. E. & Sonnhammer, E. L. Automatic clustering of orthologs and in-paralogs from pairwise species comparisons. J. Mol. Biol. 314, 1041–1052 (2001)
Regier, J. C. & Shi, D. Increased yield of PCR product from degenerate primers with nondegenerate, nonhomologous 5′ tails. Biotechniques 38, 34–38 (2005)
Staden, R., Judge, D. & Bonfield, J. Sequence assembly and finishing methods. Methods Biochem. Anal. 43, 303–322 (2001)
Smith, S. W., Overbeck, R., Woese, C. R., Gilbert, W. & Gillevet, P. M. The genetic data environment and expandable GUI for multiple sequence analysis. Comp. Appl. Biosci. 10, 671–675 (1994)
Katoh, K., Kuma, K., Toh, H. & Miyata, T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–518 (2005)
PAUP*. v.4.0 (Sinauer Associates, Sunderland, Massachusetts, 2002)
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogeny. Bioinformatics 17, 754–755 (2001)
Ronquist, F. & Huelsenbeck, J. P. MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003)
Altekar, G., Dwarkadas, S., Huelsenbeck, J. P. & Ronquist, F. Parallel Metropolis-coupled Markov chain Monte Carlo for Bayesian phylogenetic inference. Bioinformatics 20, 407–415 (2004)
MRMODELTEST. v.2 (Evolutionary Biology Centre, Uppsala University, 2004)
Regier, J. C. & Shultz, J. W. Elongation factor-2: a useful gene for arthropod phylogenetics. Mol. Phylogenet. Evol. 20, 136–148 (2001)
Cummings, M. & Huskamp, J. Grid computing. EDUCAUSE Rev. 40, 116–117 (2005)
Bazinet, A. L. & Cummings, M. C. in Distributed & Grid Computing–Science Made Transparent for Everyone. Principles, Applications and Supporting Communities (ed. Weber, M. H. W.) 2–13 (Tectum, 2009)
Acknowledgements
C.W.C. thanks W. Hartman for early insight into questions of arthropod phylogeny and D. Percy for sequencing. J.W.M. and R.W. thank N. Tait, G. Hampson and R. Hessler for help collecting samples. J.C.R. and A.Z. thank M. Cummings and A. Bazinet for making available grid computing, and the DNA Sequencing Facility at the Center for Biosystems Research, University of Maryland Biotechnology Institute. J.W.S. was supported by the Maryland Agricultural Experiment Station. C.W.C. was supported by the Whiteley Center. This work was funded by two programmes at the US National Science Foundation, namely Biocomplexity in the Environment: Genome-Enabled Environmental Science and Engineering, and Assembling the Tree of Life.
Author Contributions C.W.C., J.C.R., J.W.S., A.Z. and J.W.M. designed the project. J.W.S., J.W.M., R.W. and J.C.R. designed and carried out taxon sampling and collection. J.C.R. and C.W.C. supervised DNA sequencing and editing, with PCR templates generated by J.C.R., B.B. and others. J.C.R., A.Z., C.W.C. and J.W.S. decided on the strategy for data analysis and its implementation, with the degen1 coding method developed and implemented by J.C.R., A.H. and A.Z. J.C.R. and A.Z. assembled the Supplementary Information and submitted sequences to GenBank. J.W.S. and J.W.M. proposed the names for the new, strongly supported clades in the Pancrustacea. C.W.C. wrote the first draft of the manuscript, with major additions by J.C.R. and J.W.S. and additional contributions by J.W.M. and A.Z. All authors commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-7 with Legends and Supplementary Tables 1-5. (PDF 918 kb)
Supplementary Data
This file contains the nucleotide data matrix, 80 taxa. (TXT 4150 kb)
Supplementary Data
This file contains the degen1 data matrix, 80 taxa. (TXT 4072 kb)
Supplementary Data
This file contains the amino acid data matrix, 85 taxa. (TXT 1087 kb)
Supplementary Data
This file contains the perl script to generate degen1 data matrices. (TXT 9 kb)
Supplementary Information
This file contains explanation of the Degen1_v1_2.pl script. (PDF 29 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Regier, J., Shultz, J., Zwick, A. et al. Arthropod relationships revealed by phylogenomic analysis of nuclear protein-coding sequences. Nature 463, 1079–1083 (2010). https://doi.org/10.1038/nature08742
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08742
This article is cited by
-
The first Cyclida from the Triassic of Italy
Swiss Journal of Palaeontology (2024)
-
Macroevolutionary dynamics of gene family gain and loss along multicellular eukaryotic lineages
Nature Communications (2024)
-
A network analysis of early arthropod evolution and the potential of the primitive
Scientific Reports (2024)
-
Bergmann-Rensch Continuum Under Shell: Gender-Specific Trend in Response to Latitudinal Gradient
Evolutionary Biology (2024)
-
Establishment of a new subfamily of Idyanthidae Lang, 1944 with the description of a new species of Pseudometeorina George & Wiest, 2015 (Copepoda, Harpacticoida) from the Eratosthenes Seamount (eastern Mediterranean Sea)
Marine Biodiversity (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.