Abstract
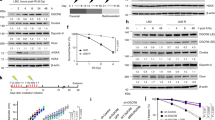
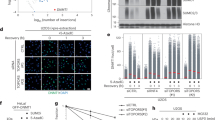
DNA double-strand breaks (DSBs) are highly cytotoxic lesions that are generated by ionizing radiation and various DNA-damaging chemicals. Following DSB formation, cells activate the DNA-damage response (DDR) protein kinases ATM, ATR and DNA-PK (also known as PRKDC). These then trigger histone H2AX (also known as H2AFX) phosphorylation and the accumulation of proteins such as MDC1, 53BP1 (also known as TP53BP1), BRCA1, CtIP (also known as RBBP8), RNF8 and RNF168/RIDDLIN into ionizing radiation-induced foci (IRIF) that amplify DSB signalling and promote DSB repair1,2. Attachment of small ubiquitin-related modifier (SUMO) to target proteins controls diverse cellular functions3,4,5,6. Here, we show that SUMO1, SUMO2 and SUMO3 accumulate at DSB sites in mammalian cells, with SUMO1 and SUMO2/3 accrual requiring the E3 ligase enzymes PIAS4 and PIAS1. We also establish that PIAS1 and PIAS4 are recruited to damage sites via mechanisms requiring their SAP domains, and are needed for the productive association of 53BP1, BRCA1 and RNF168 with such regions. Furthermore, we show that PIAS1 and PIAS4 promote DSB repair and confer ionizing radiation resistance. Finally, we establish that PIAS1 and PIAS4 are required for effective ubiquitin-adduct formation mediated by RNF8, RNF168 and BRCA1 at sites of DNA damage7,8,9,10,11. These findings thus identify PIAS1 and PIAS4 as components of the DDR and reveal how protein recruitment to DSB sites is controlled by coordinated SUMOylation and ubiquitylation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Stucki, M. & Jackson, S. P. MDC1/NFBD1: a key regulator of the DNA damage response in higher eukaryotes. DNA Repair (Amst.) 3, 953–957 (2004)
Downs, J. A., Nussenzweig, M. C. & Nussenzweig, A. Chromatin dynamics and the preservation of genetic information. Nature 447, 951–958 (2007)
Hay, R. T. SUMO: a history of modification. Mol. Cell 18, 1–12 (2005)
Meulmeester, E. & Melchior, F. Cell biology: SUMO. Nature 452, 709–711 (2008)
Bergink, S. & Jentsch, S. Principles of ubiquitin and SUMO modifications in DNA repair. Nature 458, 461–467 (2009)
Geoffroy, M. C. & Hay, R. T. An additional role for SUMO in ubiquitin-mediated proteolysis. Nature Rev. Mol. Cell Biol. 10, 564–568 (2009)
Doil, C. et al. RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell 136, 435–446 (2009)
Huen, M. S. et al. RNF8 transduces the DNA-damage signal via histone ubiquitylation and checkpoint protein assembly. Cell 131, 901–914 (2007)
Mailand, N. et al. RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell 131, 887–900 (2007)
Morris, J. R. & Solomon, E. BRCA1: BARD1 induces the formation of conjugated ubiquitin structures, dependent on K6 of ubiquitin, in cells during DNA replication and repair. Hum. Mol. Genet. 13, 807–817 (2004)
Stewart, G. S. et al. The RIDDLE syndrome protein mediates a ubiquitin-dependent signaling cascade at sites of DNA damage. Cell 136, 420–434 (2009)
Evdokimov, E., Sharma, P., Lockett, S. J., Lualdi, M. & Kuehn, M. R. Loss of SUMO1 in mice affects RanGAP1 localization and formation of PML nuclear bodies, but is not lethal as it can be compensated by SUMO2 or SUMO3. J. Cell Sci. 121, 4106–4113 (2008)
Lukas, C., Falck, J., Bartkova, J., Bartek, J. & Lukas, J. Distinct spatiotemporal dynamics of mammalian checkpoint regulators induced by DNA damage. Nature Cell Biol. 5, 255–260 (2003)
Limoli, C. L. & Ward, J. F. A new method for introducing double-strand breaks into cellular DNA. Radiat. Res. 134, 160–169 (1993)
Matsuoka, S. et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 316, 1160–1166 (2007)
Hickson, I. et al. Identification and characterization of a novel and specific inhibitor of the ataxia-telangiectasia mutated kinase ATM. Cancer Res. 64, 9152–9159 (2004)
Sartori, A. A. et al. Human CtIP promotes DNA end resection. Nature 450, 509–514 (2007)
Potts, P. R. & Yu, H. Human MMS21/NSE2 is a SUMO ligase required for DNA repair. Mol. Cell. Biol. 25, 7021–7032 (2005)
Kolas, N. K. et al. Orchestration of the DNA-damage response by the RNF8 ubiquitin ligase. Science 318, 1637–1640 (2007)
Lou, Z. et al. MDC1 maintains genomic stability by participating in the amplification of ATM-dependent DNA damage signals. Mol. Cell 21, 187–200 (2006)
Stewart, G. S., Wang, B., Bignell, C. R., Taylor, A. M. & Elledge, S. J. MDC1 is a mediator of the mammalian DNA damage checkpoint. Nature 421, 961–966 (2003)
Wang, B. & Elledge, S. J. Ubc13/Rnf8 ubiquitin ligases control foci formation of the Rap80/Abraxas/Brca1/Brcc36 complex in response to DNA damage. Proc. Natl Acad. Sci. USA 104, 20759–20763 (2007)
Xie, A. et al. Distinct roles of chromatin-associated proteins MDC1 and 53BP1 in mammalian double-strand break repair. Mol. Cell 28, 1045–1057 (2007)
Aravind, L. & Koonin, E. V. SAP – a putative DNA-binding motif involved in chromosomal organization. Trends Biochem. Sci. 25, 112–114 (2000)
Durant, S. T. & Nickoloff, J. A. Good timing in the cell cycle for precise DNA repair by BRCA1. Cell Cycle 4, 1216–1222 (2005)
Gudmundsdottir, K. & Ashworth, A. The roles of BRCA1 and BRCA2 and associated proteins in the maintenance of genomic stability. Oncogene 25, 5864–5874 (2006)
Pierce, A. J., Hu, P., Han, M., Ellis, N. & Jasin, M. Ku DNA end-binding protein modulates homologous repair of double-strand breaks in mammalian cells. Genes Dev. 15, 3237–3242 (2001)
Stucki, M. et al. MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell 123, 1213–1226 (2005)
Polanowska, J., Martin, J. S., Garcia-Muse, T., Petalcorin, M. I. & Boulton, S. J. A conserved pathway to activate BRCA1-dependent ubiquitylation at DNA damage sites. EMBO J. 25, 2178–2188 (2006)
Tahk, S. et al. Control of specificity and magnitude of NF-κB and STAT1-mediated gene activation through PIASy and PIAS1 cooperation. Proc. Natl Acad. Sci. USA 104, 11643–11648 (2007)
Acknowledgements
We thank S.P.J. lab members for support, in particular J. Harrigan, P. Huertas, S. Gravel, K. Dry and R. Chapman. We also thank C. Lukas for U2OS cells expressing GFP–BRCA1/Flag–BARD1, D. Durocher and G. Stewart for hTERT RIDDLE syndrome fibroblasts complemented with vector or HA–RNF168 and RNF168 antibody, T. Halazonetis for RNF8 antibody, R. Baer for the Flag–BARD1 construct, P. Harkin for the HA–BRCA1 construct, K. Iwabuchi for HA-tagged, full length, N, C, CΔBRCT and BRCT 53BP1 constructs, R. Walker for help with FACS, and J. R. Morris for sharing results before publication. Research in the S.P.J. lab is supported by grants from Cancer Research UK and the European Union (Integrated Project DNA repair, LSHG-CT-2005-512113, and Genomic Instability in Cancer and Precancer, HEALTH-F2-2007-201630).
Author Contributions R.B. cloned the PIAS cDNAs, tested the original siRNA efficiencies and provided help with processing of laser experiments. J.C. intensively helped with cell survival, homologous recombination and NHEJ experiments and provided support with tissue culture maintenance and stable-cell-line generation. S.P. set up the laser system in the laboratory and helped perform and analyse the FRAP experiments. K.M.M. provided the initial results on 53BP1 IRIF in PIAS4-depleted cells and constructed siRNA-resistant RFP–PIAS4. Y.G. initiated the project, led the teamwork and performed all other experiments described in the manuscript. Y.G. and S.P.J. conceived the study and wrote the paper. All authors discussed and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 Legend for Supplementary Movies 1a -1d, Supplementary Figures 2-10 with Legends, Supplementary Tables 1-3 and Supplementary References. (PDF 31609 kb)
Supplementary Movie 1
This file contains Supplementary Movie 1a (see Supplementary Information file- Supplementary Figure 1 Legend for details). (MOV 2894 kb)
Supplementary Movie 2
This file contains Supplementary Movie 1b (see Supplementary Information file- Supplementary Figure 1 Legend for details). (MOV 1977 kb)
Supplementary Movie 3
This file contains Supplementary Movie 1c (see Supplementary Information file- Supplementary Figure 1 Legend for details). (MOV 3032 kb)
Supplementary Movie 4
This file contains Supplementary Movie 1d (see Supplementary Information file-Supplementary Figure 1 Legend for details). (MOV 3379 kb)
Rights and permissions
About this article
Cite this article
Galanty, Y., Belotserkovskaya, R., Coates, J. et al. Mammalian SUMO E3-ligases PIAS1 and PIAS4 promote responses to DNA double-strand breaks. Nature 462, 935–939 (2009). https://doi.org/10.1038/nature08657
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08657
This article is cited by
-
SENP6 regulates localization and nuclear condensation of DNA damage response proteins by group deSUMOylation
Nature Communications (2023)
-
The ubiquitin-dependent ATPase p97 removes cytotoxic trapped PARP1 from chromatin
Nature Cell Biology (2022)
-
Crosstalk between SUMOylation and ubiquitylation controls DNA end resection by maintaining MRE11 homeostasis on chromatin
Nature Communications (2022)
-
Unique evolutionary trajectories of breast cancers with distinct genomic and spatial heterogeneity
Scientific Reports (2021)
-
Zinc finger protein ZNF384 is an adaptor of Ku to DNA during classical non-homologous end-joining
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.