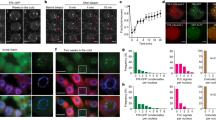
Abstract
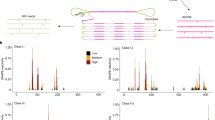
Transcription in eukaryotic genomes generates an extensive array of non-protein-coding RNA, the functional significance of which is mostly unknown1. We are investigating the link between non-coding RNA and chromatin regulation through analysis of FLC — a regulator of flowering time in Arabidopsis and a target of several chromatin pathways. Here we use an unbiased strategy to characterize non-coding transcripts of FLC and show that sense/antisense transcript levels correlate in a range of mutants and treatments, but change independently in cold-treated plants. Prolonged cold epigenetically silences FLC in a Polycomb-mediated process called vernalization2. Our data indicate that upregulation of long non-coding antisense transcripts covering the entire FLC locus may be part of the cold-sensing mechanism. Induction of these antisense transcripts occurs earlier than, and is independent of, other vernalization markers3 and coincides with a reduction in sense transcription. We show that addition of the FLC antisense promoter sequences to a reporter gene is sufficient to confer cold-induced silencing of the reporter. Our data indicate that cold-induced FLC antisense transcripts have an early role in the epigenetic silencing of FLC, acting to silence FLC transcription transiently. Recruitment of the Polycomb machinery then confers the epigenetic memory. Antisense transcription events originating from 3′ ends of genes might be a general mechanism to regulate the corresponding sense transcription in a condition/stage-dependent manner.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ponting, C. P., Oliver, P. L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009)
De Lucia, F. et al. A PHD-Polycomb repressive complex 2 triggers the epigenetic silencing of FLC during vernalization. Proc. Natl Acad. Sci. USA 105, 16831–16836 (2008)
Sung, S. & Amasino, R. M. Vernalization in Arabidopsis thaliana is mediated by the PHD finger protein VIN3. Nature 427, 159–164 (2004)
He, Y. et al. The antisense transcriptomes of human cells. Science 322, 1855–1857 (2008)
Swiezewski, S. et al. Small RNA-mediated chromatin silencing directed to the 3′ region of the Arabidopsis gene encoding the developmental regulator, FLC . Proc. Natl Acad. Sci. USA 104, 3633–3638 (2007)
Liu, F. et al. The Arabidopsis RNA-binding protein FCA requires a lysine-specific demethylase 1 homolog to downregulate FLC . Mol. Cell 28, 398–407 (2007)
Bastow, R. et al. Vernalization requires epigenetic silencing of FLC by histone methylation. Nature 427, 164–167 (2004)
Lee, I., Michaels, S. D., Masshardt, A. S. & Amasino, R. M. The late-flowering phenotype of FRIGIDA and mutations in LUMINIDEPENDENS is suppressed in the Landsberg erecta strain of Arabidopsis . Plant J. 6, 903–909 (1994)
Macknight, R. et al. FCA, a gene controlling flowering time in Arabidopsis, encodes a protein containing RNA-binding domains. Cell 89, 737–745 (1997)
Oeder, S., Mages, J., Flicek, P. & Lang, R. Uncovering information on expression of natural antisense transcripts in Affymetrix MOE430 datasets. BMC Genomics 8, 200 (2007)
RIKEN Genome Exploration Research Group and Genome Science Group (Genome Network Project Core Group) and the FANTOM Consortium Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566 (2005)
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007)
Greb, T. et al. The PHD finger protein VRN5 functions in the epigenetic silencing of Arabidopsis FLC . Curr. Biol. 17, 73–78 (2007)
Petruk, S. et al. Transcription of bxd noncoding RNAs promoted by trithorax represses Ubx in cis by transcriptional interference. Cell 127, 1209–1221 (2006)
Camblong, J. et al. Antisense RNA stabilization induces transcriptional gene silencing via histone deacetylation in S. cerevisiae . Cell 131, 706–717 (2007)
Onodera, Y. et al. Plant nuclear RNA polymerase IV mediates siRNA and DNA methylation-dependent heterochromatin formation. Cell 120, 613–622 (2005)
Pandey, R. R. et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 32, 232–246 (2008)
Sheldon, C. C., Finnegan, J. E. & Peacock, W. J. and Dennis, E. S., Mechanisms of gene repression by vernalization in Arabidopsis . Plant J. 59, 488–498 (2009)
Camblong, J. et al. Trans-acting antisense RNAs mediate transcriptional gene cosuppression in S. cerevisiae . Genes Dev. 23, 1534–1545 (2009)
Sung, S., Schmitz, R. J. & Amasino, R. M. A PHD finger protein involved in both the vernalization and photoperiod pathways in Arabidopsis . Genes Dev. 20, 3244–3248 (2006)
Czechowski, T. et al. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis . Plant Physiol. 139, 5–17 (2005)
Curtis, M. D. & Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 133, 462–469 (2003)
Lee, I., Michaels, S. D., Masshardt, A. S. & Amasino, R. M. The late-flowering phenotype of FRIGIDA and mutations in LUMINIDEPENDENS is suppressed in the Landsberg erecta strain of Arabidopsis . Plant J. 6, 903–909 (1994)
Bechtold, N., Ellis, J. & Pelletier, G. In planta Agrobacterium mediated gene transfer by infiltration of adult Arabidopsis thaliana plants. C.R. Acad. Sci. III 316, 1194–1199 (1993)
Hajdukiewicz, P., Svab, Z. & Maliga, P. The small, versatile pPZP family of Agrobacterium binary vectors for plant transformation. Plant Mol. Biol. 25, 989–994 (1994)
Yamamoto, Y. Y. et al. Gene trapping of the Arabidopsis genome with a firefly luciferase reporter. Plant J. 35, 273–283 (2003)
Mylne, J. S. et al. LHP1, the Arabidopsis homologue of HETEROCHROMATIN PROTEIN1, is required for epigenetic silencing of FLC . Proc. Natl Acad. Sci. USA 103, 5012–5017 (2006)
Völker, A., Stierhof, Y. D. & Jürgens, G. Cell cycle-independent expression of the Arabidopsis cytokinesis-specific syntaxin KNOLLE results in mistargeting to the plasma membrane and is not sufficient for cytokinesis. J. Cell Sci. 114, 3001–3012 (2001)
Liu, F. et al. The Arabidopsis RNA-binding protein FCA requires a lysine-specific demethylase 1 homolog to downregulate FLC . Mol. Cell 28, 398–407 (2007)
De Lucia, F. et al. A PHD-polycomb repressive complex 2 triggers the epigenetic silencing of FLC during vernalization. Proc. Natl Acad. Sci. USA 105, 16831–16836 (2008)
Quesada, V., Macknight, R., Dean, C. & Simpson, G. G. Autoregulation of FCA pre-mRNA processing controls Arabidopsis flowering time. EMBO J. 22, 3142–3152 (2003)
R. Development Core Team. R: a language and environment for statistical computing. 〈http://www.R-project.org〉 (R Foundation for Statistical Computing, 2006)
Smyth, G. K. in Bioinformatics and Computational Biology Solutions using R and Bioconductor (eds Gentleman, R., Carey, V. J., Huber, W., Irizarry, R. A. & Dudoit, S.) 397–420 (Springer, 2005)
Olshen, A. B., Venkatraman, E. S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004)
Pelz, C. R., Kulesz-Martin, M., Bagby, G. & Sears, R. C. Global rank-invariant set normalization (GRSN) to reduce systematic distortions in microarray data. BMC Bioinformatics 9, 520 (2008)
Acknowledgements
We thank all members of the Dean laboratory for discussions, J. Ecker and B. Gregory (Salk Institute) for help with the RiboMinus technique, C. Lister for help with cloning steps, M. Lenhard and H. Breuninger for seeds of LUC control plants. This research was supported by the BBSRC Core Strategic Grant to the John Innes Centre and BBSRC BB/D010799/1, NERC NE/C507629 and EU SIROCCO LSHG-CT-2006-037900 grants to C.D.
Author Contributions C.D. and S.S. designed the study. S.S. performed most experiments. S.S. and F.L. performed the array experiment. A.M. and S.S. analysed microarray data. C.D. and S.S. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-6 with Legends. (PDF 2692 kb)
Supplementary Figure 1
This file shows a landscape view of Supplementary Figure 1. (PDF 3236 kb)
Rights and permissions
About this article
Cite this article
Swiezewski, S., Liu, F., Magusin, A. et al. Cold-induced silencing by long antisense transcripts of an Arabidopsis Polycomb target. Nature 462, 799–802 (2009). https://doi.org/10.1038/nature08618
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08618
This article is cited by
-
Capture of regulatory factors via CRISPR–dCas9 for mechanistic analysis of fine-tuned SERRATE expression in Arabidopsis
Nature Plants (2024)
-
Regulation of coconut somatic embryogenesis: decoding the role of long non-coding RNAs
Plant Biotechnology Reports (2024)
-
Cold induction of nuclear FRIGIDA condensation in Arabidopsis
Nature (2023)
-
Getting to the bottom of lncRNA mechanism: structure–function relationships
Mammalian Genome (2022)
-
A premature stop codon in BrFLC2 transcript results in early flowering in oilseed-type Brassica rapa plants
Plant Molecular Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.