Abstract
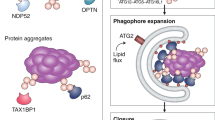
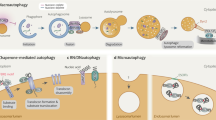
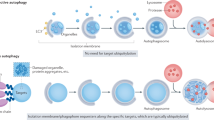
Macroautophagy is a process that leads to the bulk degradation of subcellular constituents by producing autophagosomes/autolysosomes1,2,3. It is believed that Atg5 (ref. 4) and Atg7 (ref. 5) are essential genes for mammalian macroautophagy. Here we show, however, that mouse cells lacking Atg5 or Atg7 can still form autophagosomes/autolysosomes and perform autophagy-mediated protein degradation when subjected to certain stressors. Although lipidation of the microtubule-associated protein light chain 3 (LC3, also known as Map1lc3a) to form LC3-II is generally considered to be a good indicator of macroautophagy6, it did not occur during the Atg5/Atg7-independent alternative process of macroautophagy. We also found that this alternative process of macroautophagy was regulated by several autophagic proteins, including Unc-51-like kinase 1 (Ulk1) and beclin 1. Unlike conventional macroautophagy, autophagosomes seemed to be generated in a Rab9-dependent manner by the fusion of isolation membranes with vesicles derived from the trans-Golgi and late endosomes. In vivo, Atg5-independent alternative macroautophagy was detected in several embryonic tissues. It also had a function in clearing mitochondria during erythroid maturation. These results indicate that mammalian macroautophagy can occur through at least two different pathways: an Atg5/Atg7-dependent conventional pathway and an Atg5/Atg7-independent alternative pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
20 January 2016
A Correction to this paper has been published: https://doi.org/10.1038/nature16538
References
Mizushima, N., Levine, B., Cuervo, A. M. & Klionsky, D. J. Autophagy fights disease through cellular self-digestion. Nature 451, 1069–1075 (2008)
Xie, Z. & Klionsky, D. J. Autophagosome formation: core machinery and adaptations. Nature Cell Biol. 9, 1102–1109 (2007)
Mizushima, N., Ohsumi, Y. & Yoshimori, T. Autophagosome formation in mammalian cells. Cell Struct. Funct. 27, 421–429 (2002)
Kuma, A. et al. The role of autophagy during the early neonatal starvation period. Nature 432, 1032–1036 (2004)
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169, 425–434 (2005)
Kabeya, Y. et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 19, 5720–5728 (2000)
Yue, Z., Jin, S., Yang, C., Levine, A. J. & Heintz, N. Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc. Natl Acad. Sci. USA 100, 15077–15082 (2003)
Qu, X. et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J. Clin. Invest. 112, 1809–1820 (2003)
Rosenbluth, J. M. & Pietenpol, J. A. mTOR regulates autophagy-associated genes downstream of p73. Autophagy 5, 114–116 (2009)
Liou, W., Geuze, H. J., Geelen, M. J. H. & Slot, J. W. The autophagic and endocytic pathways converge at the nascent autophagic vacuoles. J. Cell Biol. 136, 61–70 (1997)
Razi, M., Chan, E. Y. W. & Tooze, S. A. Early endosomes and endosomal coatomer are required for autophagy. J. Cell Biol. 185, 305–321 (2009)
Yoshimori, T., Yamamoto, A., Moriyama, Y., Futai, M. & Tashiro, Y. Bafilomycin A1, a specific inhibitor of vacuolar-type H+-ATPase, inhibits acidification and protein degradation in lysosomes of cultured cells. J. Biol. Chem. 266, 17707–17712 (1991)
Bursch, W. The autophagosomal–lysosomal compartment in programmed cell death. Cell Death Differ. 8, 569–581 (2001)
Cuervo, A. M. Autophagy: many paths to the same end. Mol. Cell. Biochem. 263, 55–72 (2004)
Mizushima, N. et al. Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells. J. Cell Biol. 152, 657–668 (2001)
Kabeya, Y. et al. LC3, GABARAP and GATE16 localize to autophagosomal membrane depending on form-II formation. J. Cell Sci. 117, 2805–2812 (2004)
Young, A. R. et al. Starvation and ULK1-dependent cycling of mammalian Atg9 between the TGN and endosomes. J. Cell Sci. 119, 3888–3900 (2006)
Hara, T. et al. FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells. J. Cell Biol. 181, 497–510 (2008)
Gagnon, E. Endoplasmic reticulum-mediated phagocytosis is a mechanism of entry into macrophages. Cell 110, 119–131 (2002)
Riederer, M. A., Soldati, T., Shapiro, A. D., Lin, J. & Pfeffer, S. R. Lysosome biogenesis requires Rab9 function and receptor recycling from endosomes to the trans-Golgi network. J. Cell Biol. 125, 573–582 (1994)
Fader, C. M. & Colombo, M. I. Multivesicular bodies and autophagy in erythrocyte maturation. Autophagy 2, 122–125 (2006)
Sandoval, H. et al. Essential role for Nix in autophagic maturation of erythroid cells. Nature 454, 232–235 (2008)
Matsui, M., Yamamoto, A., Kuma, A., Ohsumi, Y. & Mizushima, N. Organelle degradation during the lens and erythroid differentiation is independent of autophagy. Biochem. Biophys. Res. Commun. 339, 485–489 (2006)
Kundu, M. et al. Ulk1 plays a critical role in the autophagic clearance of mitochondria and ribosomes during reticulocyte maturation. Blood 112, 1493–1502 (2008)
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006)
Komatsu, M. et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441, 880–884 (2006)
Shimizu, S. et al. Role of Bcl-2 family proteins in a non-apoptotic programmed cell death dependent on autophagy genes. Nature Cell Biol. 6, 1221–1228 (2004)
Ogier-Denis, E., Houri, J. J., Bauvy, C. & Codogno, P. Guanine nucleotide exchange on heterotrimeric Gi3 protein controls autophagic sequestration in HT-29 cells. J. Biol. Chem. 271, 28593–28600 (1996)
Acknowledgements
We thank M. Narita and A. R. J. Young for critical reading of the manuscript; N. Mizushima for providing Atg5+/- mice and the expression plasmid of GFP–LC3; T. Yoshimori for the human beclin 1 expression plasmid; and T. Kitamura for providing Plat-E cells. This study was supported in part by the Program for Promotion of Fundamental Studies in Health Sciences of the National Institute of Biomedical Innovation (NIBIO), a grant for Creative Scientific Research, a grant for the 21st Century COE Program from the Japanese Ministry of Education, Science, Sports and Culture, a grant for Comprehensive Research on Aging and Health from the Japanese Ministry of Health, Labor and Welfare, and a grant for Solution-Oriented Research for Science and Technology (SORST) from the Japan Science and Technology Corporation. This study was also supported by grants from the Uehara Memorial Foundation, the Sagawa Foundation for Promotion of Cancer Research, the YASUDA Medical Foundation, the Astellas foundation for research on metabolic disorders, and the Foundation for Promotion of Cancer Research.
Author Contributions Y.N. performed the biochemical analyses. S.A. and T.K. performed the electron microscopy analyses. K.F. and H.Y. performed the Rab9 study. T.M. developed the Lamp2 immunofluorescence assay. M.K. provided the Atg7-/- cells. K.O. contributed data analysis. Y.T. supervised data interpretation. S.S. designed the research and wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-21 with Legends. (PDF 10346 kb)
Rights and permissions
About this article
Cite this article
Nishida, Y., Arakawa, S., Fujitani, K. et al. Discovery of Atg5/Atg7-independent alternative macroautophagy. Nature 461, 654–658 (2009). https://doi.org/10.1038/nature08455
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08455
This article is cited by
-
Quercetin promotes ATG5-mediating autophagy-dependent ferroptosis in gastric cancer
Journal of Molecular Histology (2024)
-
The impact of VPS35 D620N mutation on alternative autophagy and its reversal by estrogen in Parkinson's disease
Cellular and Molecular Life Sciences (2024)
-
NDP52 mediates an antiviral response to hepatitis B virus infection through Rab9-dependent lysosomal degradation pathway
Nature Communications (2023)
-
Pancreatic β-cell mitophagy as an adaptive response to metabolic stress and the underlying mechanism that involves lysosomal Ca2+ release
Experimental & Molecular Medicine (2023)
-
Obesity impairs cardiolipin-dependent mitophagy and therapeutic intercellular mitochondrial transfer ability of mesenchymal stem cells
Cell Death & Disease (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.