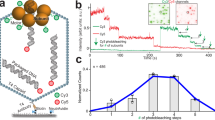
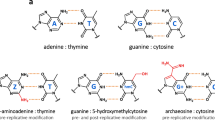
Abstract
The ASCE (additional strand, conserved E) superfamily of proteins consists of structurally similar ATPases associated with diverse cellular activities involving metabolism and transport of proteins and nucleic acids in all forms of life1. A subset of these enzymes consists of multimeric ringed pumps responsible for DNA transport in processes including genome packaging in adenoviruses, herpesviruses, poxviruses and tailed bacteriophages2. Although their mechanism of mechanochemical conversion is beginning to be understood3, little is known about how these motors engage their nucleic acid substrates. Questions remain as to whether the motors contact a single DNA element, such as a phosphate or a base, or whether contacts are distributed over several parts of the DNA. Furthermore, the role of these contacts in the mechanochemical cycle is unknown. Here we use the genome packaging motor of the Bacillus subtilis bacteriophage ϕ29 (ref. 4) to address these questions. The full mechanochemical cycle of the motor, in which the ATPase is a pentameric-ring5 of gene product 16 (gp16), involves two phases—an ATP-loading dwell followed by a translocation burst of four 2.5-base-pair (bp) steps6 triggered by hydrolysis product release7. By challenging the motor with a variety of modified DNA substrates, we show that during the dwell phase important contacts are made with adjacent phosphates every 10-bp on the 5′–3′ strand in the direction of packaging. As well as providing stable, long-lived contacts, these phosphate interactions also regulate the chemical cycle. In contrast, during the burst phase, we find that DNA translocation is driven against large forces by extensive contacts, some of which are not specific to the chemical moieties of DNA. Such promiscuous, nonspecific contacts may reflect common translocase–substrate interactions for both the nucleic acid and protein translocases of the ASCE superfamily1.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Iyer, L. M., Makarova, K. S., Koonin, E. V. & Aravind, L. Comparative genomics of the FtsK-HerA superfamily of pumping ATPases: implications for the origins of chromosome segregation, cell division and viral capsid packaging. Nucleic Acids Res. 32, 5260–5279 (2004)
Burroughs, A. M., Iyer, L. M. & Aravind, L. Comparative Genomics and Evolutionary Trajectories of Viral ATP Dependent DNA-Packaging Systems 48–65 (Basel, 2007)
Hopfner, K. P. & Michaelis, J. Mechanisms of nucleic acid translocases: lessons from structural biology and single-molecule biophysics. Curr. Opin. Struct. Biol. 17, 87–95 (2007)
Grimes, S., Jardine, P. J. & Anderson, D. Bacteriophage φ29 DNA packaging. Adv. Virus Res. 58, 255–294 (2002)
Morais, M. C. et al. Defining molecular and domain boundaries in the bacteriophage φ29 DNA packaging motor. Structure 16, 1267–1274 (2008)
Moffitt, J. R. et al. Intersubunit coordination in a homomeric ring ATPase. Nature 457, 446–450 (2009)
Chemla, Y. R. et al. Mechanism of force generation of a viral DNA packaging motor. Cell 122, 683–692 (2005)
Thiviyanathan, V. et al. Structure of hybrid backbone methylphosphonate DNA heteroduplexes: effect of R and S stereochemistry. Biochemistry 41, 827–838 (2002)
Strauss, J. K. & Maher, L. J. DNA bending by asymmetric phosphate neutralization. Science 266, 1829–1834 (1994)
Eoff, R. L., Spurling, T. L. & Raney, K. D. Chemically modified DNA substrates implicate the importance of electrostatic interactions for DNA unwinding by Dda helicase. Biochemistry 44, 666–674 (2005)
Kawaoka, J., Jankowsky, E. & Pyle, A. M. Backbone tracking by the SF2 helicase NPH-II. Nature Struct. Biol. 11, 526–530 (2004)
SenGupta, D. J. & Borowiec, J. A. Strand-specific recognition of a synthetic DNA replication fork by the SV40 large tumor antigen. Science 256, 1656–1661 (1992)
Mancini, E. J. et al. Atomic snapshots of an RNA packaging motor reveal conformational changes linking ATP hydrolysis to RNA translocation. Cell 118, 743–755 (2004)
Dillingham, M. S., Soultanas, P. & Wigley, D. B. Site-directed mutagenesis of motif III in PcrA helicase reveals a role in coupling ATP hydrolysis to strand separation. Nucleic Acids Res. 27, 3310–3317 (1999)
Howard, J. Mechanics of Motor Proteins and the Cytoskeleton 1st edn 62 (Sinauer Associates, 2001)
Stanley, L. K. et al. When a helicase is not a helicase: dsDNA tracking by the motor protein EcoR124I. EMBO J. 25, 2230–2239 (2006)
Enemark, E. J. & Joshua-Tor, L. Mechanism of DNA translocation in a replicative hexameric helicase. Nature 442, 270–275 (2006)
Pearson, R. K. & Fox, M. S. Effects of DNA heterologies on bacteriophage lambda packaging. Genetics 118, 5–12 (1988)
Moll, W. D. & Guo, P. Translocation of nicked but not gapped DNA by the packaging motor of bacteriophage φ29. J. Mol. Biol. 351, 100–107 (2005)
Oram, M., Sabanayagam, C. & Black, L. W. Modulation of the packaging reaction of bacteriophage t4 terminase by DNA structure. J. Mol. Biol. 381, 61–72 (2008)
Agresti, A. Categorical Data Analysis Chs 5–6 (Wiley-Interscience, 2003)
Shen, J., Gai, D., Patrick, A., Greenleaf, W. B. & Chen, X. S. The roles of the residues on the channel β-hairpin and loop structures of simian virus 40 hexameric helicase. Proc. Natl Acad. Sci. USA 102, 11248–11253 (2005)
Barkow, S. R., Levchenko, I., Baker, T. A. & Sauer, R. T. Polypeptide translocation by the AAA+ ClpXP protease machine. Chem. Biol. 16, 605–612 (2009)
Mulkidjanian, A. Y., Makarova, K. S., Galperin, M. Y. & Koonin, E. V. Inventing the dynamo machine: the evolution of the F-type and V-type ATPases. Nature Rev. Microbiol. 5, 892–899 (2007)
Smith, D. E. et al. The bacteriophage φ29 portal motor can package DNA against a large internal force. Nature 413, 748–752 (2001)
Lewis, J. & Sauro, J. When 100% really isn’t 100%: improving the accuracy of small-sample estimates of completion rates. J. Usability Stud. 1, 136–150 (2006)
Agresti, A. & Coull, B. Approximate is better than “exact” for interval estimation of binomial proportions. Am. Stat. 52, 119–126 (1998)
Rickgauer, J. P. et al. Portal motor velocity and internal force resisting viral DNA packaging in bacteriophage ϕ29. Biophys. J. 94, 159–167 (2008)
Smith, S. B., Cui, Y. & Bustamante, C. Optical-trap force transducer that operates by direct measurement of light momentum. Methods Enzymol. 361, 134–162 (2003)
Moffitt, J. R., Chemla, Y. R., Izhaky, D. & Bustamante, C. Differential detection of dual traps improves the spatial resolution of optical tweezers. Proc. Natl Acad. Sci. USA 103, 9006–9011 (2006)
Acknowledgements
We thank C. L. Hetherington, M. Kopaczynska, A. Spakowitz and J. M. Berger for critical discussions, and D. Reid, M. T. Couvillon and N. L. S. Chavez for preliminary work leading to this publication. K.A. acknowledges the PMMB fellowship through the Burroughs Wellcome Fund, A.T.P. the NIH Molecular Biophysics Training Grant, A.K. the Human Frontier Science Program Cross-Disciplinary Fellowship, J.R.M. the NSF Graduate Research Fellowship, and Y.R.C. the Burroughs Wellcome Fund Career Award at the Scientific Interface for funding. This research was supported in part by the National Institutes of Health (NIH) grants GM-071552, DE-003606 and GM-059604. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author Contributions K.A., A.T.P., A.K. and J.R.M. conducted the experiments; K.A., A.T.P., A.K., J.R.M. and Y.R.C. performed the analysis; S.G., P.J.J. and D.L.A. prepared and provided experimental materials; and K.A., A.T.P., A.K., J.R.M., Y.R.C., S.G., P.J.J. and C.B. wrote the paper. K.A., A.T.P., A.K. and J.R.M. contributed equally to this work.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-6, a Supplementary Discussion, Supplementary Data, Supplementary Figures 1-2 with Legends and Supplementary References. (PDF 596 kb)
Rights and permissions
About this article
Cite this article
Aathavan, K., Politzer, A., Kaplan, A. et al. Substrate interactions and promiscuity in a viral DNA packaging motor. Nature 461, 669–673 (2009). https://doi.org/10.1038/nature08443
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08443
This article is cited by
-
Optical tweezers in single-molecule biophysics
Nature Reviews Methods Primers (2021)
-
A DNA packaging motor inchworms along one strand allowing it to adapt to alternative double-helical structures
Nature Communications (2021)
-
Helical inchworming: a novel translocation mechanism for a ring ATPase
Biophysical Reviews (2021)
-
Progress towards revealing the mechanism of herpesvirus capsid maturation and genome packaging
Protein & Cell (2020)
-
Nucleotide-dependent DNA gripping and an end-clamp mechanism regulate the bacteriophage T4 viral packaging motor
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.