Abstract
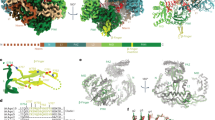
RNA silencing is a conserved regulatory mechanism in fungi, plants and animals that regulates gene expression and defence against viruses and transgenes1. Small silencing RNAs of ∼20–30 nucleotides and their associated effector proteins, the Argonaute family proteins, are the central components in RNA silencing2. A subset of small RNAs, such as microRNAs and small interfering RNAs (siRNAs) in plants, Piwi-interacting RNAs in animals and siRNAs in Drosophila, requires an additional crucial step for their maturation; that is, 2′-O-methylation on the 3′ terminal nucleotide3,4,5,6. A conserved S-adenosyl-l-methionine-dependent RNA methyltransferase, HUA ENHANCER 1 (HEN1), and its homologues are responsible for this specific modification3,4,5,7,8. Here we report the 3.1 Å crystal structure of full-length HEN1 from Arabidopsis in complex with a 22-nucleotide small RNA duplex and cofactor product S-adenosyl-l-homocysteine. Highly cooperative recognition of the small RNA substrate by multiple RNA binding domains and the methyltransferase domain in HEN1 measures the length of the RNA duplex and determines the substrate specificity. Metal ion coordination by both 2′ and 3′ hydroxyls on the 3′-terminal nucleotide and four invariant residues in the active site of the methyltransferase domain suggests a novel Mg2+-dependent 2′-O-methylation mechanism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Ghildiyal, M. & Zamore, P. D. Small silencing RNAs: an expanding universe. Nature Rev. Genet. 10, 94–108 (2009)
Farazi, T. A., Juranek, S. A. & Tuschl, T. The growing catalog of small RNAs and their association with distinct Argonaute/Piwi family members. Development 135, 1201–1214 (2008)
Yu, B. et al. Methylation as a crucial step in plant microRNA biogenesis. Science 307, 932–935 (2005)
Horwich, M. D. et al. The Drosophila RNA methyltransferase, DmHen1, modifies germline piRNAs and single-stranded siRNAs in RISC. Curr. Biol. 17, 1265–1272 (2007)
Saito, K. et al. Pimet, the Drosophila homolog of HEN1, mediates 2'-O-methylation of Piwi-interacting RNAs at their 3′ ends. Genes Dev. 21, 1603–1608 (2007)
Kirino, Y. & Mourelatos, Z. Mouse Piwi-interacting RNAs are 2'-O-methylated at their 3′ termini. Nature Struct. Mol. Biol. 14, 347–348 (2007)
Kirino, Y. & Mourelatos, Z. The mouse homolog of HEN1 is a potential methylase for Piwi-interacting RNAs. RNA 13, 1397–1401 (2007)
Kurth, H. M. & Mochizuki, K. 2'-O-methylation stabilizes Piwi-associated small RNAs and ensures DNA elimination in Tetrahymena . RNA 15, 675–685 (2009)
Chen, X., Liu, J., Cheng, Y. & Jia, D. HEN1 functions pleiotropically in Arabidopsis development and acts in C function in the flower. Development 129, 1085–1094 (2002)
Park, W., Li, J., Song, R., Messing, J. & Chen, X. CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana . Curr. Biol. 12, 1484–1495 (2002)
Yang, Z., Ebright, Y. W., Yu, B. & Chen, X. HEN1 recognizes 21–24 nt small RNA duplexes and deposits a methyl group onto the 2' OH of the 3′ terminal nucleotide. Nucleic Acids Res. 34, 667–675 (2006)
Li, J., Yang, Z., Yu, B., Liu, J. & Chen, X. Methylation protects miRNAs and siRNAs from a 3′-end uridylation activity in Arabidopsis . Curr. Biol. 15, 1501–1507 (2005)
Ramachandran, V. & Chen, X. Degradation of microRNAs by a family of exoribonucleases in Arabidopsis . Science 321, 1490–1492 (2008)
Chen, X. A marked end. Nature Struct. Mol. Biol. 14, 259–260 (2007)
Tkaczuk, K., Obarska, A. & Bujnicki, J. Molecular phylogenetics and comparative modeling of HEN1, a methyltransferase involved in plant microRNA biogenesis. BMC Evol. Biol. 6, 6 (2006)
Kang, C. B., Dhe-Paganon, S. & Yoon, H. S. FKBP family proteins: immunophilins with versatile biological functions. Neurosignals 16, 318–325 (2008)
Tian, B., Bevilacqua, P. C., Diegelman-Parente, A. & Mathews, M. B. The double-stranded-RNA-binding motif: interference and much more. Nature Rev. Mol. Cell Biol. 5, 1013–1023 (2004)
Curry, S. & Conte, M. R. A terminal affair: 3′-end recognition by the human La protein. Trends Biochem. Sci. 31, 303–305 (2006)
Maraia, R. J. & Bayfield, M. A. The La protein-RNA complex surfaces. Mol. Cell 21, 149–152 (2006)
Teplova, M. et al. Structural basis for recognition and sequestration of UUUOH 3′ temini of nascent RNA polymerase III transcripts by La, a rheumatic disease autoantigen. Mol. Cell 21, 75–85 (2006)
Vargason, J. M., Szittya, G., Burgyán, J. & Hall, T. M. T. Size selective recognition of siRNA by an RNA silencing suppressor. Cell 115, 799–811 (2003)
Ye, K., Malinina, L. & Patel, D. J. Recognition of small interfering RNA by a viral suppressor of RNA silencing. Nature 426, 874–878 (2003)
Yu, B., Chapman, E. J., Yang, Z., Carrington, J. C. & Chen, X. Transgenically expressed viral RNA silencing suppressors interfere with microRNA methylation in Arabidopsis . FEBS Lett. 580, 3117–3120 (2006)
Schubert, H. L., Blumenthal, R. M. & Cheng, X. Many paths to methyltransfer: a chronicle of convergence. Trends Biochem. Sci. 28, 329–335 (2003)
Ryter, J. M. & Schultz, S. C. Molecular basis of double-stranded RNA-protein interactions: structure of a dsRNA-binding domain complexed with dsRNA. EMBO J. 17, 7505–7513 (1998)
MacRae, I. J. et al. Structural basis for double-stranded RNA processing by Dicer. Science 311, 195–198 (2006)
Lingel, A., Simon, B., Izaurralde, E. & Sattler, M. Nucleic acid 3′-end recognition by the Argonaute2 PAZ domain. Nature Struct. Mol. Biol. 11, 576–577 (2004)
Ma, J.-B., Ye, K. & Patel, D. J. Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature 429, 318–322 (2004)
Yang, Z. et al. Approaches for studying microRNA and small interfering RNA methylation in vitro and in vivo . Methods Enzymol. 427, 139–154 (2007)
Stump, W. T. & Hall, K. B. Crosslinking of an iodo-uridine-RNA hairpin to a single site on the human U1A N-terminal RNA binding domain. RNA 1, 55–63 (1995)
Doublié, S. & Carter, C. W. Jr. Preparation of selenomethionyl proteins for phase determination. Methods Enzymol. 276, 523–530 (1997)
Otwinowski, Z., Minor, W. & Carter, C. W. Jr. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Hendrickson, W. A. Determination of macromolecular structures from anomalous diffraction of synthrotron radiation. Science 254, 51–58 (1991)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in eletron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Brunger, A. T. Version 1.2 of the Crystallography and NMR system. Nature Protocols 2, 2728–2733 (2007)
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993)
Acknowledgements
We thank K. Sergiy for assistance with the in-house X-ray generator operation, and the staff at Advanced Photon Source beamlines 19ID and 23ID, Argonne National Laboratory, for help with data collection. We thank T. Townes and H. Wang for critical reading of the manuscript. This work was supported by a start-up fund (to J.-B.M.) and partly by a grant from the V Foundation for Cancer Research (to J.-B.M.) and a grant from the National Science Foundation (MCB-0718029 to X.C.). D.G.V. is supported by grants from National Institutes of Health (R01 GM074252 and R01 GM074840).
Author Contributions Y.H. expressed and purified proteins, grew crystals, solved structure and wrote the manuscript. J.-B.M. collected data, solved structure, performed crosslinking assays and wrote the manuscript. L.J. performed small RNA methyltransferase assays and wrote the manuscript. Q.H. and D.G.V. were involved in data processing and refinement. X.C. analysed data and wrote the manuscript. The overall project management and manuscript preparation were by Y.H., X.C. and J.-B.M.
Author information
Authors and Affiliations
Corresponding author
Additional information
Atomic coordinates and structural factors for the reported crystal structure have been deposited in the Protein Data Bank under access code 3HTX.
Supplementary information
Supplementary Information
This file contains Supplementary Notes, Supplementary References, Supplementary Tables 1-3 and Supplementary Figures S1-S10 with Legends. (PDF 7558 kb)
Rights and permissions
About this article
Cite this article
Huang, Y., Ji, L., Huang, Q. et al. Structural insights into mechanisms of the small RNA methyltransferase HEN1. Nature 461, 823–827 (2009). https://doi.org/10.1038/nature08433
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08433
This article is cited by
-
Phylogenetic and Evolutionary Analysis of Plant Small RNA 2′-O-Methyltransferase (HEN1) Protein Family
Journal of Molecular Evolution (2023)
-
Exogenous miRNA: A Perspective Role as Therapeutic in Rheumatoid Arthritis
Current Rheumatology Reports (2021)
-
Ectopic overexpression of bol-miR390a from broccoli (B. oleracea L var. italica) increases lateral branches in Arabidopsis
Plant Growth Regulation (2020)
-
Mechanisms of microRNA-mediated gene regulation in unicellular model alga Chlamydomonas reinhardtii
Biotechnology for Biofuels (2018)
-
Messages on small RNA duplexes in plants
Journal of Plant Research (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.