Abstract
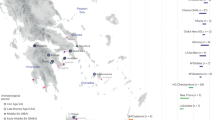
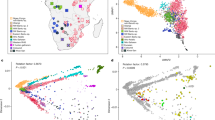
India has been underrepresented in genome-wide surveys of human variation. We analyse 25 diverse groups in India to provide strong evidence for two ancient populations, genetically divergent, that are ancestral to most Indians today. One, the ‘Ancestral North Indians’ (ANI), is genetically close to Middle Easterners, Central Asians, and Europeans, whereas the other, the ‘Ancestral South Indians’ (ASI), is as distinct from ANI and East Asians as they are from each other. By introducing methods that can estimate ancestry without accurate ancestral populations, we show that ANI ancestry ranges from 39–71% in most Indian groups, and is higher in traditionally upper caste and Indo-European speakers. Groups with only ASI ancestry may no longer exist in mainland India. However, the indigenous Andaman Islanders are unique in being ASI-related groups without ANI ancestry. Allele frequency differences between groups in India are larger than in Europe, reflecting strong founder effects whose signatures have been maintained for thousands of years owing to endogamy. We therefore predict that there will be an excess of recessive diseases in India, which should be possible to screen and map genetically.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Majumdar, D. N. & Rao, C. R. Race Elements in Bengal: a Quantitative Study (Asia Publishing House, 1960)
Roychoudhury, A. K. & Nei, M. Genetic relationships between Indians and their neighboring populations. Hum. Hered. 35, 201–206 (1985)
Das, B. M., Das, P. B., Das, R., Walter, H. & Danker-Hopfe, H. Anthropological studies in Assam, India. Anthropol. Anz. 44, 239–248 (1986)
Zerjal, T. et al. Y-chromosomal insights into the genetic impact of the caste system in India. Hum. Genet. 121, 137–144 (2007)
Bamshad, M. et al. Genetic evidence on the origins of Indian caste populations. Genome Res. 11, 994–1004 (2001)
Wells, R. S. et al. The Eurasian heartland: a continental perspective on Y-chromosome diversity. Proc. Natl Acad. Sci. USA 98, 10244–10249 (2001)
Thanseem I et al. Genetic affinities among the lower castes and tribal groups of India: inference from Y chromosome and mitochondrial DNA. BMC Genet. 7, 42 (2006)
Basu, A. et al. Ethnic India: a genomic view, with special reference to peopling and structure. Genome Res. 13, 2277–2290 (2003)
Thangaraj, K. et al. In situ origin of deep rooting lineages of mitochondrial Macrohaplogroup ‘M’ in India. BMC Genomics 7, 151 (2006)
Indian Genome Variation Consortium. Genetic landscape of the people of India: a canvas for disease gene exploration. J. Genet. 87, 3–20 (2008)
Rosenberg, N. A. et al. Low levels of genetic divergence across geographically and linguistically diverse populations from India. PLoS Genet. 2, e215 (2006)
Abbi, A. Is Great Andamanese genealogically and typologically distinct from Onge and Jarawa? Language Sciences 10.1016/j.langsci.2008.02.002 (22 April 2008)
The International HapMap Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007)
Li, J. Z. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008)
Jakobsson, M. et al. Genotype, haplotype and copy-number variation in worldwide human populations. Nature 451, 998–1003 (2008)
Menozzi, P., Piazza, A. & Cavalli-Sforza, L. Synthetic maps of human gene frequencies in Europeans. Science 201, 786–792 (1978)
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006)
Thangaraj, K., Ramana, G. V. & Singh, L. Y-chromosome and mitochondrial DNA polymorphisms in Indian populations. Electrophoresis 20, 1743–1747 (1999)
Thangaraj, K. et al. Genetic affinities of the Andaman Islanders, a vanishing human population. Curr. Biol. 13, 86–93 (2003)
Lao, O. et al. Correlation between genetic and geographic structure in Europe. Curr. Biol. 18, 1241–1248 (2008)
Novembre, J. et al. Genes mirror geography within Europe. Nature 456, 98–101 (2008)
Dronamraju, K. R. Mating systems of the Andhra Pradesh people. Cold Spring Harb. Symp. Quant. Biol. 29, 81–84 (1964)
Nei, M. & Chesser, R. K. Estimation of fixation indices and gene diversities. Ann. Hum. Genet. 47, 253–259 (1983)
Karve, I. Hindu Society—an Interpretation (S. R. Deshmukh, 1968)
Boivin, N. in The Evolution and History of Human Populations in South Asia (eds Petraglia, M. D. & Allchin, B.) 341–362 (Springer, 2007)
Dirks, N. B. Castes of Mind: Colonialism and the Making of Modern India (Princeton Univ. Press, 2001)
Bhasin, M. K. & Walter, H. Genetics of Castes and Tribes of India (Kamla-Raj Enterprises, 2001)
Index of /genotypes/2008-07_phaseIII. 〈http://ftp.hapmap.org/genotypes/2008-07_phaseIII/〉
Campbell, C. D. et al. Demonstrating stratification in a European American population. Nature Genet. 37, 868–872 (2005)
Haldane, J. B. S. A defense of beanbag genetics. Perspect. Biol. Med. 7, 343–359 (1964)
Dhandapany, P. S. et al. A common Cardiac Myosin Binding Protein C variant associated with cardiomyopathies in South Asia. Nature Genet. 41, 187–191 (2009)
Pemberton, T. J. et al. Using population mixtures to optimize the utility of genomic databases: linkage disequilibrium and association study design in India. Ann. Hum. Genet. 72, 535–546 (2008)
Künsch, H. R. The jackknife and the bootstrap for general stationary observations. Ann. Stat. 17, 1217–1241 (1989)
Keinan, A., Mullikin, J. C., Patterson, N. & Reich, D. Measurement of the human allele frequency spectrum demonstrates greater genetic drift in East Asians than in Europeans. Nature Genet. 39, 1251–1255 (2007)
Cavalli-Sforza, L. L. & Edwards, A. W. Phylogenetic analysis. Models and estimation procedures. Am. J. Hum. Genet. 19, 233–257 (1967)
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000)
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003)
Mallory, J. P. & Adams, D. O. The Oxford Introduction to Proto-Indo-European and the Proto-Indo-European World (Oxford Univ. Press, 2006)
Barik, S. S. et al. Detailed mtDNA genotypes permit a reassessment of the settlement and population structure of the Andaman Islands. Am. J. Phys. Anthropol. 136, 19–27 (2008)
Palanichamy, M. G. et al. Comment on “Reconstructing the Origin of Andaman Islanders”. Science 311, 470 (2006)
Watkins, W. S. et al. Genetic variation in South Indian castes: evidence from Y-chromosome, mitochondrial, and autosomal polymorphisms. BMC Genet. 9, 86 (2008)
Southworth, F. C. Linguistic archaeology of South Asia (Routledge-Curzon, 2005)
Cordaux, R. et al. Mitochondrial DNA analysis reveals diverse histories of tribal populations from India. Eur. J. Hum. Genet. 11, 253–264 (2003)
Kivisild, T. et al. Deep common ancestry of Indian and western-Eurasian mitochondrial DNA lineages. Curr. Biol. 9, 1331–1334 (1999)
Falush, D. et al. Traces of human migrations in Helicobacter pylori populations. Science 299, 1582–1585 (2003)
Baird, S. J. E. Phylogenetics: Fisher’s markers of admixture. Heredity 97, 81–83 (2006)
Chikhi, L., Bruford, M. W. & Beaumont, M. A. Estimation of admixture proportions: a likelihood-based approach using Markov chain Monte Carlo. Genetics 158, 1347–1362 (2001)
Hellenthal, G., Auton, A. & Falush, D. Inferring human colonization history using a copying model. PLoS Genet. 4, e1000078 (2008)
Lohmueller, K. E., Bustamante, C. D. & Clark, A. G. Methods for human demographic inference using haplotype patterns from genomewide single-nucleotide polymorphism data. Genetics 182, 217–231 (2009)
Singh, K. S. People of India, National Series, Volume III, Scheduled Tribes (Oxford Univ. Press, 1994)
Singh, K. S. People of India, National Series, Volume III, Scheduled Castes (Oxford Univ. Press, 1993)
McCarroll, S. A. et al. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nature Genet. 40, 1166–1174 (2008)
Thorburn, D. On the asymptotic normality of the jackknife. Scand. J. Stat. 4, 113–118 (1977)
Acknowledgements
We thank the volunteers from throughout India who donated DNA; A. G. Reddy, A. Shah and R. Tamang for generating the Y chromosome and mtDNA data; J. Neubauer for sample preparation; and A. Tandon for data curation. We thank B. N. Sarkar and A. G. Roy for helping with group census size estimates, and D. Falush, J. Novembre, A. Ruiz-Linares and S. Watkins for comments on the manuscript. D.R., N.P. and A.L.P. were supported by NIH grant HG004168, and D.R. was supported by a Burroughs Wellcome Career Development Award in the Biomedical Sciences. K.T. and L.S. were supported by grants from the Council of Scientific and Industrial Research of the Government of India, and K.T. was supported by a UKIERI Major Award (RG-4772).
Author Contributions K.T. and L.S. collected the DNA samples, D.R., K.T. and L.S. collected the genetic data, N.P. developed the mathematical theory for f-statistics, and D.R., K.T., N.P. and A.L.P. analysed the data. D.R. wrote the manuscript and Supplementary Information with input from all authors.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Tables S1-S6, Supplementary Figures S1-S7, Supplementary Notes S1-S5 and Supplementary References. (PDF 1777 kb)
Supplementary Appendix
This file contains Supplementary Data, Supplementary Statistics and Supplementary References. (PDF 162 kb)
Rights and permissions
About this article
Cite this article
Reich, D., Thangaraj, K., Patterson, N. et al. Reconstructing Indian population history. Nature 461, 489–494 (2009). https://doi.org/10.1038/nature08365
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08365
This article is cited by
-
Title-molecular diagnostics of dystrophinopathies in Sri Lanka towards phenotype predictions: an insight from a South Asian resource limited setting
European Journal of Medical Research (2024)
-
More than a decade of genetic research on the Denisovans
Nature Reviews Genetics (2024)
-
Case–control association study of congenital heart disease from a tertiary paediatric cardiac centre from North India
BMC Pediatrics (2023)
-
Indigenous Australian genomes show deep structure and rich novel variation
Nature (2023)
-
The Newfoundland and Labrador mosaic founder population descends from an Irish and British diaspora from 300 years ago
Communications Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.