Abstract
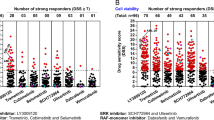
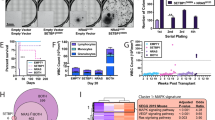
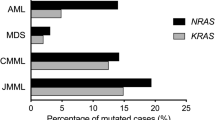
The cascade comprising Raf, mitogen-activated protein kinase kinase (MEK) and extracellular signal-regulated kinase (ERK) is a therapeutic target in human cancers with deregulated Ras signalling, which includes tumours that have inactivated the Nf1 tumour suppressor1,2. Nf1 encodes neurofibromin, a GTPase-activating protein that terminates Ras signalling by stimulating hydrolysis of Ras–GTP. We compared the effects of inhibitors of MEK in a myeloproliferative disorder (MPD) initiated by inactivating Nf1 in mouse bone marrow and in acute myeloid leukaemias (AMLs) in which cooperating mutations were induced by retroviral insertional mutagenesis. Here we show that MEK inhibitors are ineffective in MPD, but induce objective regression of many Nf1-deficient AMLs. Drug resistance developed because of outgrowth of AML clones that were present before treatment. We cloned clone-specific retroviral integrations to identify candidate resistance genes including Rasgrp1, Rasgrp4 and Mapk14, which encodes p38α. Functional analysis implicated increased RasGRP1 levels and reduced p38 kinase activity in resistance to MEK inhibitors. This approach represents a robust strategy for identifying genes and pathways that modulate how primary cancer cells respond to targeted therapeutics and for probing mechanisms of de novo and acquired resistance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Downward, J. Targeting RAS signalling pathways in cancer therapy. Nature Rev. Cancer 3, 11–22 (2003)
Schubbert, S., Shannon, K. & Bollag, G. Hyperactive Ras in developmental disorders and cancer. Nature Rev. Cancer 7, 295–308 (2007)
Van Etten, R. A. & Shannon, K. M. Focus on myeloproliferative diseases and myelodysplastic syndromes. Cancer Cell 6, 547–552 (2004)
Lauchle, J. O., Braun, B. S., Loh, M. L. & Shannon, K. Inherited predispositions and hyperactive Ras in myeloid leukemogenesis. Pediatr. Blood Cancer 46, 579–585 (2006)
Shannon, K. M. et al. Loss of the normal NF1 allele from the bone marrow of children with type 1 neurofibromatosis and malignant myeloid disorders. N. Engl. J. Med. 330, 597–601 (1994)
Bollag, G. et al. Loss of NF1 results in activation of the Ras signaling pathway and leads to aberrant growth in murine and human hematopoietic cells. Nature Genet. 12, 144–148 (1996)
Side, L. et al. Homozygous inactivation of the NF1 gene in bone marrow cells from children with neurofibromatosis type 1 and malignant myeloid disorders. N. Engl. J. Med. 336, 1713–1720 (1997)
Le, D. T. et al. Somatic inactivation of Nf1 in hematopoietic cells results in a progressive myeloproliferative disorder. Blood 103, 4243–4250 (2004)
Wolff, L., Koller, R., Hu, X. & Anver, M. R. A Moloney murine leukemia virus-based retrovirus with 4070A long terminal repeat sequences induces a high incidence of myeloid as well as lymphoid neoplasms. J. Virol. 77, 4965–4971 (2003)
Sebolt-Leopold, J. S. et al. Blockade of the MAP kinase pathway suppresses growth of colon tumors in vivo . Nature Med. 5, 810–816 (1999)
Brown, A. P., Carlson, T. C., Loi, C. M. & Graziano, M. J. Pharmacodynamic and toxicokinetic evaluation of the novel MEK inhibitor, PD0325901, in the rat following oral and intravenous administration. Cancer Chemother. Pharmacol. 59, 671–679 (2007)
Dupuy, A. J. et al. Mammalian mutagenesis using a highly mobile somatic Sleeping Beauty transposon system. Nature 436, 221–226 (2005)
Stone, J. C. Regulation of Ras in lymphocytes: get a GRP. Biochem. Soc. Trans. 34, 858–861 (2006)
Das, J. et al. Digital signaling and hysteresis characterize ras activation in lymphoid cells. Cell 136, 337–351 (2009)
Bain, J. et al. The selectivity of protein kinase inhibitors: a further update. Biochem. J. 408, 297–315 (2007)
Fabian, M. A. et al. A small molecule-kinase interaction map for clinical kinase inhibitors. Nature Biotechnol. 23, 329–336 (2005)
Druker, B. J. et al. Activity of a specific inhibitor of the BCR-ABL tyrosine kinase in the blast crisis of chronic myeloid leukemia and acute lymphoblastic leukemia with the Philadelphia chromosome. N. Engl. J. Med. 344, 1038–1042 (2001)
Shah, N. P. et al. Multiple BCR-ABL kinase domain mutations confer polyclonal resistance to the tyrosine kinase inhibitor imatinib (STI571) in chronic phase and blast crisis chronic myeloid leukemia. Cancer Cell 2, 117–125 (2002)
Choi, S. et al. Relapse in children with acute lymphoblastic leukemia involving selection of a preexisting drug-resistant subclone. Blood 110, 632–639 (2007)
Mullighan, C. G. et al. Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science 322, 1377–1380 (2008)
Sharma, S. V. et al. A common signaling cascade may underlie ‘addiction’ to the Src, BCR-ABL, and EGF receptor oncogenes. Cancer Cell 10, 425–435 (2006)
Parmar, S. et al. Role of the p38 mitogen-activated protein kinase pathway in the generation of the effects of imatinib mesylate (STI571) in BCR-ABL-expressing cells. J. Biol. Chem. 279, 25345–2532 (2004)
Kogan, S. C. et al. Bethesda proposals for classification of nonlymphoid hematopoietic neoplasms in mice. Blood 100, 238–245 (2002)
Akagi, K. et al. RTCGD: retroviral tagged cancer gene database. Nucleic Acids Res. 32 (Database issue). D523–D527 (2004)
Curtiss, N. P. et al. Isolation and analysis of candidate myeloid tumor suppressor genes from a commonly deleted segment of 7q22. Genomics 85, 600–607 (2005)
Acknowledgements
We are grateful to B. Braun, J. Downing, S. Lowe and C. Sawyers for discussion and advice throughout the project. We are thankful for C. Hartzell for studies on RasGRP protein expression. This work was supported by National Institutes of Health grants U01 CA84221, R37 CA72614, T32 CA09043, T32 HD044331 and K08 CA119105, by a Specialized Center of Research award from the Leukemia and Lymphoma Society (LLS 7019-04), by the US Army Neurofibromatosis Research Program (Project DAMD 17-02-1-0638), by the Ronald McDonald House Charities of Southern California/Couples Against Leukemia, by the Jeffrey and Karen Peterson Family Foundation and by the Frank A. Campini Foundation. S.C.K. is a Scholar of the Leukemia and Lymphoma Society of America. J.P.R. is a Kimmel Foundation Scholar. The Intramural Research Program of the National Cancer Institute’s Center for Cancer Research supports research in the laboratory of L.W. at the National Institutes of Health.
Author Contributions J.O.L and K.S. designed and performed experiments, analysed data and wrote the paper. D.K., D.T.L., M.C., K.K., K.W. and J.M.B. designed experiments, conducted studies, analysed data and provided input to the manuscript. K.A. performed bioinformatics analysis of retroviral insertion sequences. Q.L., K.M.C., E.D.-F., M.G. and M.T. designed and conducted experiments. J.P.R. provided critical reagents, assisted in experimental design and edited the manuscript. N.C., N.J. and L.W. provided retroviral reagents and training in experimental protocols. L.P. provided the mouse strain and input into the manuscript. J.S.-L. and S.P. developed protocols for suspending and administering the MEK inhibitor and provided the drug used in all studies.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-5 with Legends and Supplementary Tables 1-3. (PDF 875 kb)
Rights and permissions
About this article
Cite this article
Lauchle, J., Kim, D., Le, D. et al. Response and resistance to MEK inhibition in leukaemias initiated by hyperactive Ras. Nature 461, 411–414 (2009). https://doi.org/10.1038/nature08279
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08279
This article is cited by
-
RasGRP1 promotes the acute inflammatory response and restricts inflammation-associated cancer cell growth
Nature Communications (2022)
-
Clinical Pharmacokinetics and Pharmacodynamics of Selumetinib
Clinical Pharmacokinetics (2021)
-
Evaluation of ERK as a therapeutic target in acute myelogenous leukemia
Leukemia (2020)
-
Current status of MEK inhibitors in the treatment of plexiform neurofibromas
Child's Nervous System (2020)
-
Loss of glucocorticoid receptor expression mediates in vivo dexamethasone resistance in T-cell acute lymphoblastic leukemia
Leukemia (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.