Abstract
Recent advances in sequencing technologies have initiated an era of personal genome sequences. To date, human genome sequences have been reported for individuals with ancestry in three distinct geographical regions: a Yoruba African, two individuals of northwest European origin, and a person from China1,2,3,4. Here we provide a highly annotated, whole-genome sequence for a Korean individual, known as AK1. The genome of AK1 was determined by an exacting, combined approach that included whole-genome shotgun sequencing (27.8× coverage), targeted bacterial artificial chromosome sequencing, and high-resolution comparative genomic hybridization using custom microarrays featuring more than 24 million probes. Alignment to the NCBI reference, a composite of several ethnic clades5,6, disclosed nearly 3.45 million single nucleotide polymorphisms (SNPs), including 10,162 non-synonymous SNPs, and 170,202 deletion or insertion polymorphisms (indels). SNP and indel densities were strongly correlated genome-wide. Applying very conservative criteria yielded highly reliable copy number variants for clinical considerations. Potential medical phenotypes were annotated for non-synonymous SNPs, coding domain indels, and structural variants. The integration of several human whole-genome sequences derived from several ethnic groups will assist in understanding genetic ancestry, migration patterns and population bottlenecks.
Similar content being viewed by others
Main
A bacterial artificial chromosome (BAC) library was constructed from AK1 genomic DNA. The genomic locations of about 100,000 AK1 BAC clones were ascertained by end-sequencing (Supplementary Table 1). Massively parallel DNA sequencing was performed using sequencing-by-synthesis with reversible-terminator chemistry on Illumina Genome Analyzers using two complementary strategies (Table 1, Supplementary Table 2 and Supplementary Fig. 1). First, selected genomic regions were sequenced at very high depth using overlapping BAC clones. Chromosome 20 was sequenced in this manner at 155× coverage, as were 390 other regions of the genome that are commonly affected by copy number variants (CNVs) (at an average of 151× coverage). Second, whole-genome sequencing was performed for the entire genome to an average depth of 27.8× using libraries of AK1 genomic DNA with different insert sizes to provide even coverage. Some sequences were generated using a reformulated cleavage reagent that removed thymine fluorophores more completely. This improved phasing and reduced background signals, error rates, and GC bias in longer reads (Supplementary Fig. 2), resulting in increased sequence yields and read lengths to 18 gigabases (Gb) per flow cell and 106 nucleotides, respectively. The average sequence quality was 24 (Q score3), and 74.4% of sequences aligned to the human genome reference (NCBI build 36.3) using the GSNAP alignment tool tolerating 5% mismatches7,8,9. A total of 99.8% of the reference genome was represented, and no coverage bias was appreciated apart from expected gaps at centromeres and other heterochromatic regions (Supplementary Discussion).
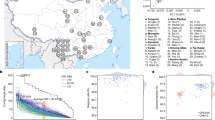
Bioinformatic filters were trained to detect and genotype SNPs in the aligned sequences. Filters ascertained by comparing SNP genotypes derived from sequencing with results from Illumina 370K genotyping array gave a positive predictive value and sensitivity of SNP detection of 99.9% and 95.0%, respectively (Supplementary Fig. 3 and Supplementary Table 3), and SNP genotype accuracy of 99.1% (Supplementary Fig. 4 and Supplementary Table 4). These filters detected 3,453,653 SNPs in the genome of AK1 (density of 1.21 per kilobase (kb)), of which 17.1% were new and 10,162 were non-synonymous (Supplementary Tables 5 and 6). These results were verified by hybridization of genomic DNA from AK1 to an Illumina 610K genotyping array, deep sequencing of chromosome 20 BAC clones, and Sanger resequencing of the AK1 genome (Supplementary Tables 7, 8 and Supplementary Discussion). The number of SNPs detected in the genome of AK1 was similar to that of James Watson, higher than Craig Venter and the Chinese YH, and less than the Yoruba African, NA18507 (Fig. 1a, b and Supplementary Table 9), which may reflect differences in technical procedures or inter-individual variability1,2,3,4. Overlap among 9,527,824 SNPs detected in these five sequenced genomes indicated that 21% of AK1’s SNPs were unique, and 8% were shared by all (Fig. 1b). A total of 2,110,403 AK1 SNPs were heterozygous, yielding a higher SNP diversity than in the Venter, Watson or YH genomes, but less than the Yoruba individual (heterozygous/homozygous SNP ratio of 1.57, and nucleotide diversity (π) of 7.40 × 10-4; Supplementary Discussion). Sequencing of other genomes using uniform technical procedures is warranted to evaluate the proportion of genetic variance explained by differences within and between human populations.
a, Geographic map showing the regions of ancestry of five sequenced genomes. MT type, mitochondrial haplogroup. b, The number of SNPs overlapping between five genomes. c, Correlation between SNP–indel densities on chromosome 6 (per 10-kb window). From top: SNP density, indel density, SNP–indel density (moving average of ten 10-kb windows), SNP density in a portion of chromosome 6, and indel density along the same portion of chromosome 6. The x axis represents the nucleotide position in Mb.
We applied the same bioinformatic filters to the genome sequence of AK1 to detect indels (Supplementary Methods). The NCBI reference genome contained 7,910 exonic indel mismatches in comparison with the reference transcript (Supplementary Discussion). Excluding these, the SNP filters detected 170,202 indels (density of 0.060 per kb), of which 71,995 were homo- or hemizygous. Sixty-two per cent of indels were new and 55.9% were deletions (Supplementary Tables 10 and 11). The size range detected was -29 to +5 nucleotides, with approximately normal frequency distribution (Supplementary Fig. 5). Two-hundred-and-twelve AK1 indels mapped to coding domains, which was three times greater than that reported for the YH genome and one-fourth of that reported for the Venter genome1,4 (Supplementary Fig. 6 and Supplementary Table 12). These marked differences reflect substantial differences either between individuals or between technical procedures, highlighting the need for definition of foundational data standards. Indel results were confirmed by Sanger resequencing of AK1 genomic DNA and deep sequencing of chromosome 20 BAC clones, showing that the sensitivity of indel detection was less than 80%, whereas the positive predictive value was 100% (Supplementary Discussion). Indel underestimation was unavoidable in local repetitive or homopolymeric sequences containing indels at, or near, the ends of reads (Supplementary Discussion). Seventy coding-domain indels were homozygous, of which 26 were in genes with Online Mendelian Inheritance in Man (OMIM) entries, 13 of which had medical phenotypes (Supplementary Table 13).
Highly significant pairwise correlations of SNP and indel densities were observed throughout the genome (Pearson’s correlation10 was 0.40 genome-wide, P < 10-300; Fig. 1c, Supplementary Fig. 7 and Supplementary Table 14). This SNP–indel correlation seems to be a general phenomenon in individual human genomes, rather than a technical artefact, because it was also detected in the YH genome (Supplementary Table 14) and has been reported for other eukaryotes, including primates11,12,13,14,15,16. SNP–indel density covariation was not a function of coverage depth or gene density (Supplementary Table 14). Genome-wide correlation of SNP and indel densities in individual human genomes is a new finding, and suggests that unifying molecular or temporal considerations underpin the generation and/or removal of both types of variants11,12,13,14,15,16.
Several, complementary approaches were used for CNV detection in AK1 (Supplementary Fig. 8). For large deletions of the AK1 genome, we used events identified in deep sequencing of 1,132 BAC clones as a training set (Supplementary Fig. 9). Most showed reduced coverage, predominance of homozygous SNPs (for heterozygous deletions), and alignment of ‘stretched’ paired-end reads in whole-genome sequences (Supplementary Fig. 10). We used these criteria to filter candidate deletions detected by a new, custom-designed 24 million probe set array comparative genomic hybridization (CGH), which found 1,237 CNV regions in total (Supplementary Fig. 11 and Supplementary Table 15), as well as genotyping microarrays (Supplementary Table 16). Figure 2a shows an example of a deletion in the genome of AK1 detected both by BAC sequencing and by CGH with the characteristics mentioned above. Figure 2b shows another deletion where the coverage drop in whole-genome sequencing was not as prominent as in BAC sequencing. However, the latter region contains all other features of deletions, illustrating the usefulness of BAC sequencing for CNV detection. The 238 regions that met these conservative criteria represent the most reliable list of true deletions yet identified in an individual genome sequence (Supplementary Fig. 10 and Supplementary Table 17). Deletions in AK1 ranged from 277 to 196,900 bases in length, and totalled 2.4 Mb. One-hundred-and-forty-eight of these had not previously been described in the Database of Genomic Variants as of 10 November 2008 (DGV; http://projects.tcag.ca/variation/).
a, Homozygous deletion identified by targeted haploid sequencing (top) and diploid sequencing (bottom). Stretched sequencing pairs and a drop in sequencing coverage define the deletion in both panels. Chr, chromosome. b, Heterozygous deletion identified by targeted haploid sequencing (top) and confirmed by diploid sequencing (bottom). Stretched pairs confirm the deletion in the diploid sequence but complete coverage drop is not detected. c, Copy number gain is identified by CGH microarray (top) and confirmed by increased coverage for the corresponding genomic region by diploid sequencing (bottom). For all panels: blue, fold coverage; horizontal red lines, stretched sequence pairs; green, CNV region in the DGV; grey, gene; vertical red bars, homozygous SNPs; vertical black bars, heterozygous SNPs; and broken vertical grey lines define the boundaries of the structural variants.
Copy number gains in AK1 were selected conservatively with three approaches for different sizes of insertions: (1) array CGH described above yielded insertions ranging from 2.15 kb to 1.06 Mb, (2) aligned end-sequences of BACs yielded insertions of 16.8 to 357.1 kb, and (3) aligned long-insert paired-end reads yielded insertions of 0.9 to 2.2 kb (Supplementary Tables 18–20). These regions were confirmed by increased sequencing coverage of these genomic regions (Supplementary Figs 12–14). An example is shown in Fig. 2c, in which an increased signal on a microarray coincided with a corresponding significant gain of sequencing coverage. The AK1 genome contained 77 copy number gains, totalling 7.0 Mb. Thirty-three (42.8%) of these were absent from the DGV and were therefore considered to be new.
Non-synonymous SNPs detected in AK1 were compared with those identified in the YH and the Yoruban genomes3,4 (which were ascertained using technical approaches similar to those used here) (Fig. 3a). Although only 37% of AK1 SNPs were shared among these three genomes, 57% of genes that contained non-synonymous SNPs in AK1 were common to all three (Fig. 3a and Supplementary Discussion). These data indicate that a subset of genes is enriched for non-synonymous SNPs in these individuals. Ontology analysis of this gene subset showed enrichment for functions associated with environmental adaptation, such as sensory function, immunological function, and signal transduction (Supplementary Table 21). Possibly, these genes have heightened diversity and/or many pseudogenes.
a, Top, the numbers of non-synonymous SNPs (nsSNPs) and genes containing non-synonymous SNPs are compared between the Korean (AK1), Han Chinese (YH) and Yoruban (NA18507) genomes. Bottom, comparison of non-synonymous SNPs and genes containing non-synonymous SNPs in AK1 with those in the YH and Yoruban genomes. Common denotes shared by three genomes. Left axis: number of nsSNPs (blue) or genes containing nsSNPs (red); right axis: ratio (%) of the number of nsSNP genes to the number of nsSNPs (green). b, Seven-hundred-and-seventy-three SNPs potentially associated with clinical phenotypes derived from the database of human gene mutation data (HGMD), OMIM, SNPedia and other hypotheses. DM, diabetes mellitus; NIDDM, non-insulin-dependent diabetes mellitus; TB, tuberculosis. c, Genes affected by large homozygous and heterozygous chromosomal deletions.
Using Trait-o-matic—an algorithm for high-throughput variant annotation—773 SNPs that were potentially associated with clinical phenotypes were identified (J. V. Thakuria and G. M. Church, manuscript in preparation; Supplementary Table 22). Of these, 269 were relatively common SNPs previously associated with risk of complex disorders or traits. For example, the genome of AK1 contained 90 SNPs that have shown associations with susceptibility to various cancers, 34 SNPs with type II diabetes mellitus, 13 with Alzheimer’s disease, and seven with rheumatoid arthritis. These data should be interpreted cautiously, however, because risk factors for complex diseases, for example, rheumatoid arthritis, differ in northwest European and Korean populations17, and because the translation of genetic burden into risk assessment for polygenic traits is rudimentary. The genome of AK1 also contained 504 non-synonymous SNPs in genes associated with complex or Mendelian disorders or traits. Of these, 22 were stop codons and five were homozygous. Among Mendelian traits, AK1 was homozygous for a variant conferring dry earwax18 that has a high allele frequency in Koreans. Eighteen variants of pharmacogenetic relevance were identified, potentially affecting dosing, efficacy and/or toxicity of β-2-adrenoceptor agonists, statins, rosiglitazone, warfarin, citalopram, abacavir, debrisoquine, bleomycin, fluorouracil and aramycin-C (Fig. 3b).
CNVs have shown associations with common, complex disorders in humans. One-hundred-and-six genes were affected by CNV losses in AK1 (Fig. 3c). One gene deleted in the genome of AK1 was leukocyte immunoglobulin-like receptor (LILRA3). Most northeast Asians have functional loss or deletion of this locus, which has been suggested to be under positive or balancing selection19.
We have obtained the genome sequence of a Korean individual by a unique combination of whole-genome shotgun sequencing, targeted BAC sequencing, and custom-designed high-resolution array CGH. This combination of approaches improved the accuracy of SNP, indel and CNV detection, and will assist in the assembly of contiguous sequences. Agreement on technical standards for individual genome sequences will aid in comparisons between genomes and, ultimately, to associations with phenotypic differences.
Methods Summary
Genomic DNA samples were obtained from an anonymous healthy Korean adult male (AK1) with normal karyotype (Supplementary Fig. 15), using guidelines approved by the Institutional Review Board of Seoul National University (approval C-0806-023-246). AK1 provided written consent for public release of genomic data. A BAC clone library was prepared using standard methods20. End sequences of 96,768 BAC clones were generated with an ABI 3730xl DNA analyser. A minimally overlapping BAC tiling path of chromosome 20, and 1,132 BAC clones from 390 common CNV regions were selected for haploid targeted sequencing. Genome-wide genotyping and CNV detection were performed using Agilent custom 24 million feature CGH array set, as well as Illumina Human cnv370- and 610-quad Beadchip. Short and long insert paired-end read libraries were generated from pooled BAC clone DNA or genomic DNA as described3. Paired-end and singleton, 36–106 nucleotide reads were generated using Illumina Genome Analyser (GA) and GAII instruments as described3. Long reads were obtained by use of multiple 36 nucleotide sequencing kits with reformulated cleavage reagent provided by Illumina for evaluation. This reformulated reagent has now been made available in all new Illumina sequencing-by-synthesis (SBS) reagent kits. The total sequencing cost took less than 200,000 US dollars with total run time of 6 weeks using three GA instruments. Sequences and runs were used in analyses if the average Q scores were ≥20. Sequences were aligned to NCBI build 36.3 using GSNAP21. SNPs and indels were identified using optimized filters through Alpheus8. Custom scripts were developed to identify CNVs in short, paired-end reads by the occurrence of clusters of reads with aligned insert sizes deviating from the mean by >2 standard deviations, and based on contiguous regions with significantly increased or depressed coverage. Putative SNPs, indels and deletions were validated by targeted Sanger sequencing (Supplementary Fig. 16 and Supplementary Table 23). Variants that were previously associated with a clinical phenotype or risk in other studies were identified with Trait-o-matic. Statistical analysis was performed using JMP-Genomics (SAS Institute) or R (http://www.R-project.org).
Online Methods
Sample preparation
Sperm samples and peripheral blood were obtained from an anonymous healthy Korean adult male in his thirties, following the guidelines by the Institutional Review Board of Seoul National University (approval number C-0806-023-246). High molecular mass genomic DNA was extracted from sperm and peripheral blood using standard protocols.
BAC clone library preparation and end sequencing
The BAC library was prepared from a partial HindIII digest of high molecular weight genomic DNA from sperm in the vector pECBAC1 using standard methods20. BAC clone DNA was extracted from a single colony by a standard alkaline lysis extraction method. Inserts of BAC clones were sequenced from both ends using universal primers, an ABI 3730xl DNA analyser and an ABI BigDye Terminator cycle sequencing (Applied Biosystems).
High-depth coverage data of single chromosome and frequent CNV region (CNVR) was obtained using targeted BAC GA sequencing. Minimally overlapping BAC clone tiling path of chromosome 20 was constructed using BAC end sequences22. Also, we selected 390 CNV regions from DGV at which the frequency of variants among the population was more than 20% (including genes). From the CNVRs, 1,132 BACs were selected. BAC clones were amplified and DNA was extracted for GA library preparation.
Genome Analyser library preparation
We constructed libraries for massive parallel sequencing under the manufacturer’s standard protocol (Illumina, Inc.)3.
Genome sequencing
Paired-end and singleton 36 nucleotide-long reads were generated using Illumina Genome Analyser (GA) and GAII instruments using standard protocols3.
Long read (2 × 88 bp, 2 × 106 bp) GA sequencing
Longer reads were enabled by an improved cleavage mix increasing the efficiency of thiamine fluorophore cleavage, thereby decreasing background noise. Paired 88-bp reads and 106-bp reads were generated with improved cleavage mix on Illumina GA II instruments using standard protocols3. Sequences passing standard Illumina GA pipeline filters (v1.1) were retained. Long insert paired reads were trimmed to 50 nucleotides for insertion detection.
Sequence alignments
Short-reads were aligned using the GSNAP21 alignment tool (Supplementary Methods).
Validation studies
Sixty-five SNPs, 60 indels and ten large deletions were selected for Sanger validation (Supplementary Discussion). PCR was performed in 50 μl containing 50 ng genomic DNA, 10 pmol of forward and reverse primer each, using Ex Taq (Takara), at 95 °C for 10 min, 35 cycles of 95 °C for 30 s, 60 °C for 30 s and 72 °C for 30 s, and finally 72 °C for 10 min, using genomic DNA (Supplementary Table 24). PCR products were subsequently purified with AccuPrep PCR purification kit (BioNeer). Sanger sequencing was performed using an ABI 3730xl DNA analyser and ABI BigDye Terminator cycle sequencing.
SNP and indel sensitivity
To estimate the sensitivity of our SNP and indel detection, we used high-depth GA data from 742 BAC clones targeting chromosome 20. We selected 22.255 Mb of chromosome 20 with single BAC coverage for the identification of highly reliable SNPs and indels. Filter conditions were identical as homozygous SNP and indel detection used in autosomes. We estimated the sensitivity by comparing them with SNPs and indels identified by whole-genome shotgun data (Supplementary Discussion).
SNP and indel correlation
Binned counts observed for both SNPs and indel start positions (bin size = 10 kb) were computed, and pairwise correlations were calculated. Bins with no coverage were removed from the analysis. To investigate the potential confounding effect of coverage and gene density on the sample SNP–indel correlation, we computed the partial correlation10 between SNP and indel counts, correcting for the effects of coverage variation and/or gene density (bin size = 10 kb). All pairwise correlations and partial correlations were computed using pcor and pcor.test from the ggm library in R. Moving average plots were generated in R using the lowess (locally weighted robust scatterplot smoothing) smooting function. Each 10-kb window was smoothed using the 100 surrounding windows. Lowess calculations were done with three iterations and without any linear interpolation of adjacent points.
Detection of true positive deletions with paired end GA data from 1,132 BACs
A sequence read pair with an insert size of more than 500 bp was defined as a stretched pair. Two criteria were used to finalize deletion list: the presence of 15 or more stretched pairs with the reads of either side clustering within 500-bp region, and the coverage of the candidate region between the clusters of stretched-pair reads less than half of those in the flanking regions of both sides. See Supplementary Methods for more details.
Detection of structural variations using microarrays
Putative structural variations were detected with microarrays (Illumina BeadChip 370K, 610K, and Agilent 24M aCGH; Supplementary Methods). For Illumina BeadChips, normalized bead intensity data and genotype calls were obtained with Illumina BeadStudio 3.1 software. Results from Agilent 24M aCGH were analysed on Nexus software (BioDiscovery Inc.). Each aberration call was manually checked to confirm the accuracy of the calls.
Detection of amplification with BAC end sequencing
BACs less than 40 kb were considered as significantly short. The co-localization of two or more short BACs was considered as the candidate region of amplification.
Detection of amplification with LIPE
Compressed pair was defined when it had a shorter than 2,000 bp insert size. The criteria used to finalize amplification list was the occurrence of ten or more de-redundant compressed pairs with the reads of either side clustering within 4 kb region. See Supplementary Methods for more details.
Confirmation of structural variants on diploid GA sequencing
The putative structural variants were confirmed using diploid GA. To confirm reliable large deletions, three criteria had to be met: (1) relative coverage drops compared to those of the flanking regions; (2) the existence of stretched paired-end sequence; and (3) the loss of heterozygous SNPs under the regions. Relative increase of sequencing coverage was used as a confirmation tool for copy number gains.
High-throughput annotation; Trait-o-matic
Trait-o-matic system was used for annotating 3.45 million SNPs of AK1 (Supplementary Methods). A working reference is available at: http://wiki.github.com/xwu/trait-o-matic/download-installation.
Accession codes
Data deposits
Data have been deposited in the NCBI short read archive under accession number SRA008370. These data are also available freely from http://gmi.ac.kr. SNPs and indels are deposited in the dbSNP database under handle GMI.
References
Levy, S. et al. The diploid genome sequence of an individual human. PLoS Biol. 5, e254 (2007)
Wheeler, D. A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008)
Bentley, D. R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008)
Wang, J. et al. The diploid genome sequence of an Asian individual. Nature 456, 60–65 (2008)
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001)
Sugarbaker, D. J. et al. Transcriptome sequencing of malignant pleural mesothelioma tumors. Proc. Natl Acad. Sci. USA 105, 3521–3526 (2008)
Miller, N. A. et al. Management of high-throughput DNA sequencing projects: alpheus. J. Comput. Sci. Syst. Biol. 1, 132–148 (2008)
Mudge, J. et al. Genomic convergence analysis of schizophrenia: mRNA sequencing reveals altered synaptic vesicular transport in post-mortem cerebellum. PLoS One 3, e3625 (2008)
Sokal, R. R. & Rohlf, F. J. Biometry: the Principles and Practice of Statistics in Biological Research 3rd edn (W.H. Freeman, 1995)
Kondrashov, A. S. Direct estimates of human per nucleotide mutation rates at 20 loci causing Mendelian diseases. Hum. Mutat. 21, 12–27 (2003)
Hardison, R. C. et al. Covariation in frequencies of substitution, deletion, transposition, and recombination during eutherian evolution. Genome Res. 13, 13–26 (2003)
Petrov, D. A. DNA loss and evolution of genome size in Drosophila . Genetica 115, 81–91 (2002)
Longman-Jacobsen, N., Williamson, J. F., Dawkins, R. L. & Gaudieri, S. In polymorphic genomic regions indels cluster with nucleotide polymorphism: Quantum Genomics. Gene 312, 257–261 (2003)
Silva, J. C. & Kondrashov, A. S. Patterns in spontaneous mutation revealed by human-baboon sequence comparison. Trends Genet. 18, 544–547 (2002)
Tian, D. et al. Single-nucleotide mutation rate increases close to insertions/deletions in eukaryotes. Nature 455, 105–108 (2008)
Lee, H. S. et al. Genetic risk factors for rheumatoid arthritis differ in Caucasian and Korean populations. Arthritis Rheum. 60, 364–371 (2009)
Yoshiura, K. et al. A SNP in the ABCC11 gene is the determinant of human earwax type. Nature Genet. 38, 324–330 (2006)
Hirayasu, K. et al. Evidence for natural selection on leukocyte immunoglobulin-like receptors for HLA class I in Northeast Asians. Am. J. Hum. Genet. 82, 1075–1083 (2008)
Frijters, A. C. J. et al. Construction of a bacterial artificial chromosome library containing large EcoRI and HindIII genomic fragments of lettuce. Theor. Appl. Genet. 94, 390–399 (1997)
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005)
Chen, R., Sodergren, E., Weinstock, G. M. & Gibbs, R. A. Dynamic building of a BAC clone tiling path for the Rat Genome Sequencing Project. Genome Res. 14, 679–684 (2004)
Acknowledgements
We acknowledge J. Yi for her help during this work. We would like to thank D. Bentley and M. Ross for providing genotypes of NA18507, and S. Ham for his comment on BAC analysis. This work has been supported by the Korean Ministry of Education, Science, and Technology (grant M10305030000), Green Cross Therapeutics (0411-20080023), and Macrogen Inc. (MG2009001). This work was also supported in part by National Institutes of Health (NIH) grants RR016480 (S.F.K.) and HG004221 (C.L.). We acknowledge G. D. May, J. E. Woodward and L. E. Clendenen for generation of some of the sequence data, and T. Ramaraj for assistance with data analysis.
Author Contributions J.-S.S. managed the project. J.-S.S., W.-Y.P., H.K., H.K., H.-S.K., I.-S.C., W.-C.L. and J.-S.L. prepared the BAC library. Seon.L., S.-H.S., J.-Y.Y., H.N.W., R.W.K., J.J.H., Sh.L., G.P.S., H.K., K.-S.Y. and H.K. executed sequencing of the genome. J.-I.K., Y.S.J., S.K., J.M., N.A.M., D.H., C.J.B., H.L., H.-J.K., M.K.L. and T.D.W. analysed sequence data. Y.S.J., Ha.P., D.S., Seung.L., M.Y., M.K., Y.Z., Hy.P. and J.Y.K. validated DNA sequence variations. Ha.P., O.G., R.E.M. and C.L. were involved with microarray experiments. A.W.Z., J.T., X.W. and G.M.C. performed Trait-o-matic analysis. J.-S.S., J.-I.K., Y.S.J., J.-H.Y. and S.F.K. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Methods with References, Supplementary Discussion with References, Supplementary Figures 1-20 with Legends, and Supplementary Tables 1-24 with Legends. Note that multi-page figures and tables are shown only as previews; the full versions are contained in the accompanying zip folders. (PDF 2760 kb)
Supplementary Figures
This zip file contains full versions of Supplementary Figures 7, 9, 10, 12, 13, 14, 16, 18 and 19 (PDFs). (ZIP 26247 kb)
Supplementary Tables
This zip file contains full versions of Supplementary Tables 1, 3, 6, 12, 14, 15, 17, 21, 22, 34 and 24 (Excel spreadsheets) and Supplementary Table 11 (text file). (ZIP 6194 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Kim, JI., Ju, Y., Park, H. et al. A highly annotated whole-genome sequence of a Korean individual. Nature 460, 1011–1015 (2009). https://doi.org/10.1038/nature08211
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08211
This article is cited by
-
A biallelic multiple nucleotide length polymorphism explains functional causality at 5p15.33 prostate cancer risk locus
Nature Communications (2023)
-
Chromatin alternates between A and B compartments at kilobase scale for subgenic organization
Nature Communications (2023)
-
Is it time to change the reference genome?
Genome Biology (2019)
-
dnAQET: a framework to compute a consolidated metric for benchmarking quality of de novo assemblies
BMC Genomics (2019)
-
The poplar pangenome provides insights into the evolutionary history of the genus
Communications Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.