Abstract
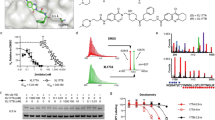
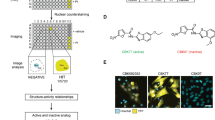
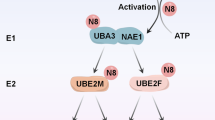
The clinical development of an inhibitor of cellular proteasome function suggests that compounds targeting other components of the ubiquitin–proteasome system might prove useful for the treatment of human malignancies. NEDD8-activating enzyme (NAE) is an essential component of the NEDD8 conjugation pathway that controls the activity of the cullin-RING subtype of ubiquitin ligases, thereby regulating the turnover of a subset of proteins upstream of the proteasome. Substrates of cullin-RING ligases have important roles in cellular processes associated with cancer cell growth and survival pathways. Here we describe MLN4924, a potent and selective inhibitor of NAE. MLN4924 disrupts cullin-RING ligase-mediated protein turnover leading to apoptotic death in human tumour cells by a new mechanism of action, the deregulation of S-phase DNA synthesis. MLN4924 suppressed the growth of human tumour xenografts in mice at compound exposures that were well tolerated. Our data suggest that NAE inhibitors may hold promise for the treatment of cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hershko, A. The ubiquitin system for protein degradation and some of its roles in the control of the cell division cycle. Cell Death Differ. 12, 1191–1197 (2005)
Kane, R. C., Bross, P. F., Farrell, A. T. & Pazdur, R. VELCADE®: U.S. FDA Approval for the treatment of multiple myeloma progressing on prior therapy. Oncologist 8, 508–513 (2003)
Kane, R. C. et al. Bortezomib for the treatment of mantle cell lymphoma. Clin. Cancer Res. 13, 5291–5294 (2007)
Nalepa, G., Rolfe, M. & Harper, J. W. Drug discovery in the ubiquitin proteasome system. Nature Rev. Drug Discov. 5, 596–613 (2006)
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998)
Haas, A. L. & Rose, I. A. The mechanism of ubiquitin activating enzyme: A kinetic and equilibrium analysis. J. Biol. Chem. 257, 10329–10337 (1982)
Wilkinson, K. D. The discovery of ubiquitin-dependent proteolysis. Proc. Natl Acad. Sci. USA 102, 15280–15282 (2005)
Kerscher, O., Felberbaum, R. & Hochstrasser, M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu. Rev. Cell Dev. Biol. 22, 159–180 (2006)
Gong, L. & Yeh, E. T. Identification of the activating and conjugating enzymes of the NEDD8 conjugation pathway. J. Biol. Chem. 274, 12036–12042 (1999)
Pan, Z. Q., Kentsis, A., Dias, D. C., Yamoah, K. & Wu, K. NEDD8 on Cullin: building an expressway to protein destruction. Oncogene 23, 1985–1997 (2004)
Petroski, M. D. & Deshaies, R. J. Function and regulation of cullin-RING ubiquitin ligases. Nature Rev. Mol. Cell Biol. 6, 9–20 (2005)
Chiba, T. & Tanaka, K. Cullin-based ubiquitin ligase and its control by NEDD8 conjugation system. Curr. Protein Pept. Sci. 5, 177–184 (2004)
Read, M. A. et al. Nedd8 modification of Cul-1 activates SCFβTrCP-dependent ubiquitination of IκBα. Mol. Cell. Biol. 20, 2326–2333 (2000)
Podust, V. N. et al. A Nedd8 conjugation pathway is essential for proteolytic targeting of p27KIP1 by ubiquitination. Proc. Natl Acad. Sci. USA 97, 4579–4584 (2000)
Duda, D. M. et al. Structural insights into NEDD8 activation of cullin-RING ligases: conformational control of conjugation. Cell 134, 995–1006 (2008)
Saha, A. & Deshaies, R. J. Multimodal activation of the ubiquitin ligase SCF by NEDD8 conjugation. Mol. Cell 32, 21–31 (2008)
Bohnsack, R. N. & Haas, A. L. Conservation in the mechanism of NEDD8 activation by the human AppBp1-Uba3 heterodimer. J. Biol. Chem. 278, 26823–26830 (2003)
Walden, H., Podgorski, M. S. & Schulman, B. A. Insights into the ubiquitin transfer cascade from the structure of the activating enzyme for NEDD8. Nature 422, 330–334 (2003)
Walden, H. et al. The structure of the APPBP1-UBA3-NEDD8-ATP complex reveals the basis for selective ubiquitin-like protein activation by an E1. Mol. Cell 12, 1427–1437 (2003)
Bloch, A. & Coutsogeorgopoulos, C. Inhibition of protein synthesis by 5′-sulfamoyladenosine. Biochemistry 10, 4394–4398 (1971)
Rock, K. L. et al. Inhibitors of the proteasome block the degradation of most cell proteins and the generation of peptides presented on MHC class I molecules. Cell 78, 761–771 (1994)
Nishitani, H. et al. Two E3 ubiquitin ligases, SCF-Skp2 and DDB1-Cul4 target human Cdt1 for proteolysis. EMBO J. 25, 1126–1136 (2006)
Kondo, T. et al. Rapid degradation of Cdt1 upon UV-induced DNA damage is mediated by SCF Skp2 complex. J. Biol. Chem. 279, 27315–27319 (2004)
Hu, J., McCall, C. M., Ohta, T. & Xiong, Y. Targeted ubiquitination of CDT1 by the DDB1–CUL4A–ROC1 ligase in response to DNA damage. Nature Cell Biol. 6, 1003–1009 (2004)
Carrano, A. C., Eytan, E., Hershko, A. & Pagano, M. SKP2 is required for ubiquitin-mediated degradation of the CDK inhibitor p27. Nature Cell Biol. 1, 193–199 (1999)
Kobayashi, A. et al. Oxidative stress sensor Keap1 functions as an adaptor for Cul3-based E3 ligase to regulate proteasomal degradation of Nrf2. Mol. Cell. Biol. 24, 7130–7139 (2004)
Nateri, A. S., Riera-Sans, L., DaCosta, C. & Behrens, A. The ubiquitin ligase SCFFbw7 antagonizes apoptotic JNK signaling. Science 303, 1374–1378 (2004)
Kamura, T. et al. Activation of HIF1α ubiquitination by a reconstituted von-Hippel-Lindau (VHL) tumor suppressor complex. Proc. Natl Acad. Sci. USA 97, 10430–10435 (2000)
Ye, X. et al. Recognition of phosphodegron motifs in human cyclin E by the SCFFbw7 ubiquitin ligase. J. Biol. Chem. 279, 50110–50119 (2004)
Donzelli, M. et al. Dual mode of degradation of Cdc25A phosphatase. EMBO J. 21, 4875–4884 (2002)
Margottin-Goguet, F. et al. Prophase destruction of Emi1 by the SCFβTrCP/Slimb ubiquitin ligase activates the anaphase promoting complex to allow progression beyond prometaphase. Dev. Cell 4, 813–826 (2003)
Winston, J. T. et al. The SCFβ-TrCP–ubiquitin ligase complex associates specifically with phosphorylated destruction motifs in IκBα and β-catenin and stimulates IκBα ubiquitination in vitro . Genes Dev. 13, 270–283 (1999)
Machida, Y. J., Hamlin, J. L. & Dutta, A. Right place, right time and only once: replication initiation in Metazoans. Cell 123, 13–24 (2005)
Arias, E. E. & Walter, J. C. Replication-dependent destruction of Cdt1 limits DNA replication to a single round per cell cycle in Xenopus egg extracts. Genes Dev. 19, 114–126 (2005)
Li, X., Zhao, Q., Liao, R., Sun, P. & Wu, X. The SCFSkp2 ubiquitin ligase complex interacts with the human replication licensing factor Cdt1 and regulates Cdt1 degradation. J. Biol. Chem. 278, 30854–30858 (2003)
Lovejoy, C. A., Lock, K., Yenamandra, A. & Cortez, D. DDB1 maintains genome integrity through regulation of Cdt1. Mol. Cell. Biol. 26, 7977–7990 (2006)
Higa, L. A., Mihaylov, I. S., Banks, D. P., Zheng, J. & Zhang, H. Radiation-mediated proteolysis of Cdt1 by CUL4–ROC1 and CSN complexes constitutes a new checkpoint. Nature Cell Biol. 5, 1008–1015 (2003)
Saxena, S. & Dutta, A. Geminin and p53: deterrents to rereplication in human cancer cells. Cell Cycle 2, 283–286 (2003)
Melixetian, M. et al. Loss of Geminin induces rereplication in the presence of functional p53. J. Cell Biol. 165, 473–482 (2004)
Vaziri, C. et al. A p53-dependent checkpoint pathway prevents rereplication. Mol. Cell 11, 997–1008 (2003)
Kim, Y. & Kipreos, E. T. Cdt1 degradation to prevent DNA rereplication: conserved and non-conserved pathways. Cell Div. 2, 18–27 (2007)
Archambault, V., Ikui, A. E., Drapkin, B. J. & Cross, F. R. Disruption of mechanisms that prevent rereplication triggers a DNA damage response. Mol. Cell. Biol. 25, 6707–6721 (2005)
Zhu, W., Chen, Y. & Dutta, A. Rereplication by depletion of geminin is seen regardless of p53 status and activates a G2/M checkpoint. Mol. Cell. Biol. 24, 7140–7150 (2004)
Lin, J. J. & Dutta, A. ATR pathway is the primary pathway for activating G2/M checkpoint induction after re-replication. J. Biol. Chem. 282, 30357–30362 (2007)
Handeli, S. & Weintraub, H. The ts41 mutation in Chinese hamster cells leads to successive S phases in the absence of intervening G2, M and G1. Cell 71, 599–611 (1992)
Chen, Y., McPhie, D. L., Hirschberg, J. & Neve, R. L. The amyloid precursor protein-binding protein APP-BP1 drives the cell cycle through the S-M checkpoint and causes apoptosis in neurons. J. Biol. Chem. 275, 8929–8935 (2000)
Acknowledgements
As this project progressed through the discovery phase, many scientists made significant contributions to its success. The authors would like to thank all past contributors to the NAE project including M. H. Foley, V. Chau, T. Gladysheva and S. Sadis. We would also like to thank D. Bowman for technical help with imaging assays, J. Blank for technical help with protein turnover assays and A. Burkhardt, P. Veiby, M. Manfredi and B. Hibner for support and guidance.
Author Contributions T.A.S., P.G.S., M.A.M., A.J.B. and J.M.G. participated in the planning, initiation, data generation and analysis of biological experiments and preparation of the manuscript. C.A.C., E.J.O., S.V., G.S.W., S.C., S.P. and S.P.L. participated in the planning, initiation, design and execution of chemical synthesis. S.A., D.P.C., A.D., H.M., M.R., J.L.G., R.E.G. and T.Talreja performed chemical synthesis; U.N. performed in vitro cell culture experiments; T.Traore and J.Z. performed in vivo anti-tumour activity and pharmacodynamic experiments; J.Y., J.J.G. and M.P.T. performed pharmacodynamic analysis experiments; A.R.L. evaluated compound potencies in cell-based assays; K.E.B. and A.M. performed immunohistochemical experiments; M.D.S. performed crystallography studies. L.R.D., J.E.B., C.F.C., M.R. and J.B.B. provided oversight of the project, reviewed and edited the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
[Competing Interests: All authors were employees and stockholders of Millennium Pharmaceuticals Inc. during the time the studies were completed.]
Additional information
MLN4924 will be provided once a standard Materials Transfer Agreement has been executed.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-6 with Legends, Supplementary Tables 1-2, Supplementary Methods and a Supplementary Discussion and Supplementary References. (PDF 523 kb)
Rights and permissions
About this article
Cite this article
Soucy, T., Smith, P., Milhollen, M. et al. An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature 458, 732–736 (2009). https://doi.org/10.1038/nature07884
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature07884
This article is cited by
-
Integrated analysis reveals the regulatory mechanism of the neddylation inhibitor MLN4924 on the metabolic dysregulation in rabbit granulosa cells
BMC Genomics (2024)
-
Interplay between proteasome inhibitors and NF-κB pathway in leukemia and lymphoma: a comprehensive review on challenges ahead of proteasome inhibitors
Cell Communication and Signaling (2024)
-
Deciphering the role of neddylation in tumor microenvironment modulation: common outcome of multiple signaling pathways
Biomarker Research (2024)
-
Targeting NEDD8 suppresses surgical stress-facilitated metastasis of colon cancer via restraining regulatory T cells
Cell Death & Disease (2024)
-
DNA-encoded library-enabled discovery of proximity-inducing small molecules
Nature Chemical Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.