Abstract
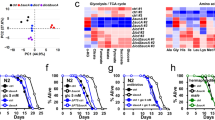
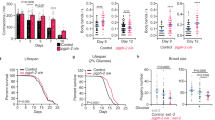
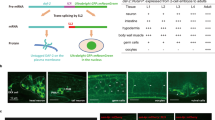
Dietary restriction is the most effective and reproducible intervention to extend lifespan in divergent species1. In mammals, two regimens of dietary restriction, intermittent fasting (IF) and chronic caloric restriction, have proven to extend lifespan and reduce the incidence of age-related disorders2. An important characteristic of IF is that it can increase lifespan even when there is little or no overall decrease in calorie intake2. The molecular mechanisms underlying IF-induced longevity, however, remain largely unknown. Here we establish an IF regimen that effectively extends the lifespan of Caenorhabditis elegans, and show that the low molecular weight GTPase RHEB-1 has a dual role in lifespan regulation; RHEB-1 is required for the IF-induced longevity, whereas inhibition of RHEB-1 mimics the caloric-restriction effects. RHEB-1 exerts its effects in part by the insulin/insulin growth factor (IGF)-like signalling effector DAF-16 in IF. Our analyses demonstrate that most fasting-induced upregulated genes require RHEB-1 function for their induction, and that RHEB-1 and TOR signalling are required for the fasting-induced downregulation of an insulin-like peptide, INS-7. These findings identify the essential role of signalling by RHEB-1 in IF-induced longevity and gene expression changes, and suggest a molecular link between the IF-induced longevity and the insulin/IGF-like signalling pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bordone, L. & Guarente, L. Calorie restriction, SIRT1 and metabolism: understanding longevity. Nature Rev. Mol. Cell Biol. 6, 298–305 (2005)
Anson, R. M., Jones, B. & de Cabod, R. The diet restriction paradigm: a brief review of the effects of every-other-day feeding. Age (Omaha) 27, 17–25 (2005)
Lakowski, B. & Hekimi, S. The genetics of caloric restriction in Caenorhabditis elegans . Proc. Natl Acad. Sci. USA 95, 13091–13096 (1998)
Bishop, N. A. & Guarente, L. Two neurons mediate diet-restriction-induced longevity in C. elegans . Nature 447, 545–549 (2007)
Panowski, S. H., Wolff, S., Aguilaniu, H., Durieux, J. & Dillin, A. PHA-4/Foxa mediates diet-restriction-induced longevity of C. elegans . Nature 447, 550–555 (2007)
Kaeberlein, M. et al. Regulation of yeast replicative life span by TOR and Sch9 in response to nutrients. Science 310, 1193–1196 (2005)
Hansen, M. et al. Lifespan extension by conditions that inhibit translation in Caenorhabditis elegans . Aging Cell 6, 95–110 (2007)
Kapahi, P. et al. Regulation of lifespan in Drosophila by modulation of genes in the TOR signaling pathway. Curr. Biol. 14, 885–890 (2004)
Saucedo, L. J. et al. Rheb promotes cell growth as a component of the insulin/TOR signalling network. Nature Cell Biol. 5, 566–571 (2003)
Long, X. et al. TOR deficiency in C. elegans causes developmental arrest and intestinal atrophy by inhibition of mRNA translation. Curr. Biol. 12, 1448–1461 (2002)
Smith, E. D. et al. Age- and calorie-independent life span extension from dietary restriction by bacterial deprivation in Caenorhabditis elegans . BMC Dev. Biol. 8, 49 (2008)
Pan, K. Z. et al. Inhibition of mRNA translation extends lifespan in Caenorhabditis elegans . Aging Cell 6, 111–119 (2007)
Greer, E. L. et al. An AMPK-FOXO pathway mediates longevity induced by a novel method of dietary restriction in C. elegans . Curr. Biol. 17, 1646–1656 (2007)
Guarente, L. & Kenyon, C. Genetic pathways that regulate ageing in model organisms. Nature 408, 255–262 (2000)
Derry, W. B., Putzke, A. P. & Rothman, J. H. Caenorhabditis elegans p53: role in apoptosis, meiosis, and stress resistance. Science 294, 591–595 (2001)
Hinnebusch, A. G. Translational regulation of GCN4 and the general amino acid control of yeast. Annu. Rev. Microbiol. 59, 407–450 (2005)
Henderson, S. T. & Johnson, T. E. daf-16 integrates developmental and environmental inputs to mediate aging in the nematode Caenorhabditis elegans . Curr. Biol. 11, 1975–1980 (2001)
Murphy, C. T. et al. Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans . Nature 424, 277–283 (2003)
Hansen, M., Hsu, A. L., Dillin, A. & Kenyon, C. New genes tied to endocrine, metabolic, and dietary regulation of lifespan from a Caenorhabditis elegans genomic RNAi screen. PLoS Genet. 1, e17 (2005)
Hsu, A. L., Murphy, C. T. & Kenyon, C. Regulation of aging and age-related disease by DAF-16 and heat-shock factor. Science 300, 1142–1145 (2003)
Rajasekaran, N. S. et al. Human αB-crystallin mutation causes oxido-reductive stress and protein aggregation cardiomyopathy in mice. Cell 130, 427–439 (2007)
Murphy, C. T., Lee, S. J. & Kenyon, C. Tissue entrainment by feedback regulation of insulin gene expression in the endoderm of Caenorhabditis elegans . Proc. Natl Acad. Sci. USA 104, 19046–19050 (2007)
Malone, E. A., Inoue, T. & Thomas, J. H. Genetic analysis of the roles of daf-28 and age-1 in regulating Caenorhabditis elegans dauer formation. Genetics 143, 1193–1205 (1996)
Luong, N. et al. Activated FOXO-mediated insulin resistance is blocked by reduction of TOR activity. Cell Metab. 4, 133–142 (2006)
Lee, G. D. et al. Dietary deprivation extends lifespan in Caenorhabditis elegans . Aging Cell 5, 515–524 (2006)
Kaeberlein, T. L. et al. Lifespan extension in Caenorhabditis elegans by complete removal of food. Aging Cell 5, 487–494 (2006)
Weinkove, D., Halstead, J. R., Gems, D. & Divecha, N. Long-term starvation and ageing induce AGE-1/PI 3-kinase-dependent translocation of DAF-16/FOXO to the cytoplasm. BMC Biol. 4, 1 (2006)
Hansen, M. et al. A role for autophagy in the extension of lifespan by dietary restriction in C. elegans . PLoS Genet. 4, e24 (2008)
Steinkraus, K. A. et al. Dietary restriction suppresses proteotoxicity and enhances longevity by an hsf-1-dependent mechanism in Caenorhabditis elegans . Aging Cell 7, 394–404 (2008)
Fabrizio, P. et al. Sir2 blocks extreme life-span extension. Cell 123, 655–667 (2005)
Brenner, S. The genetics of Caenorhabditis elegans . Genetics 77, 71–94 (1974)
Kamath, R. S., Martinez-Campos, M., Zipperlen, P., Fraser, A. G. & Ahringer, J. Effectiveness of specific RNA-mediated interference through ingested double-stranded RNA in Caenorhabditis elegans . Genome Biol. 2, research0002.1–research0002.10 (2001)
Maeda, I., Kohara, Y., Yamamoto, M. & Sugimoto, A. Large-scale analysis of gene function in Caenorhabditis elegans by high-throughput RNAi. Curr. Biol. 11, 171–176 (2001)
Acknowledgements
We thank members of our laboratory for technical comments and helpful discussion. This work was supported by grants from the Ministry of Education, Culture, Sports, Science and Technology of Japan (to E.N.). Some nematode strains used in this work were provided by the Caenorhabditis Genetics Center, which is funded by the NIH National Center for Research Resources (NCRR).
Author Contributions S.H. conceived the study, designed and performed the experiments, and wrote the manuscript with the help of E.N.; S.H. and T.Y. analysed the microarray data; M.U. conducted DAF-16::GFP localization experiments; E.N. supervised the project. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures1-11 with Legends and Supplementary Tables 1-2. (PDF 2910 kb)
Rights and permissions
About this article
Cite this article
Honjoh, S., Yamamoto, T., Uno, M. et al. Signalling through RHEB-1 mediates intermittent fasting-induced longevity in C. elegans. Nature 457, 726–730 (2009). https://doi.org/10.1038/nature07583
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07583
This article is cited by
-
Dauer juvenile recovery transcriptome of two contrasting EMS mutants of the entomopathogenic nematode Heterorhabditis bacteriophora
World Journal of Microbiology and Biotechnology (2024)
-
Reduced insulin/IGF-1 signalling upregulates two anti-viral immune pathways, decreases viral load and increases survival under viral infection in C. elegans
GeroScience (2024)
-
Intermittent fasting shifts the diurnal transcriptome atlas of transcription factors
Molecular and Cellular Biochemistry (2024)
-
Exploring the precision redox map during fasting-refeeding and satiation in C. elegans
Stress Biology (2023)
-
Dietary regulation in health and disease
Signal Transduction and Targeted Therapy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.