Abstract
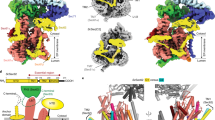
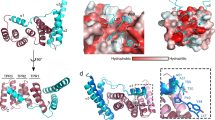
Most proteins are secreted from bacteria by the interaction of the cytoplasmic SecA ATPase with a membrane channel, formed by the heterotrimeric SecY complex. Here we report the crystal structure of SecA bound to the SecY complex, with a maximum resolution of 4.5 ångström (Å), obtained for components from Thermotoga maritima. One copy of SecA in an intermediate state of ATP hydrolysis is bound to one molecule of the SecY complex. Both partners undergo important conformational changes on interaction. The polypeptide-cross-linking domain of SecA makes a large conformational change that could capture the translocation substrate in a ‘clamp’. Polypeptide movement through the SecY channel could be achieved by the motion of a ‘two-helix finger’ of SecA inside the cytoplasmic funnel of SecY, and by the coordinated tightening and widening of SecA’s clamp above the SecY pore. SecA binding generates a ‘window’ at the lateral gate of the SecY channel and it displaces the plug domain, preparing the channel for signal sequence binding and channel opening.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rapoport, T. A. Protein translocation across the eukaryotic endoplasmic reticulum and bacterial plasma membranes. Nature 450, 663–669 (2007)
Neumann-Haefelin, C., Schafer, U., Muller, M. & Koch, H. G. SRP-dependent co-translational targeting and SecA-dependent translocation analyzed as individual steps in the export of a bacterial protein. EMBO J. 19, 6419–6426 (2000)
Qi, H. Y. & Bernstein, H. D. SecA is required for the insertion of inner membrane proteins targeted by the Escherichia coli signal recognition particle. J. Biol. Chem. 274, 8993–8997 (1999)
Duong, F. & Wickner, W. Sec-dependent membrane protein biogenesis: SecYEG, preprotein hydrophobicity and translocation kinetics control the stop-transfer function. EMBO J. 17, 696–705 (1998)
van den Berg, B. et al. X-ray structure of a protein-conducting channel. Nature 427, 36–44 (2004)
Osborne, A. R. & Rapoport, T. A. Protein translocation is mediated by oligomers of the SecY complex with one SecY copy forming the channel. Cell 129, 97–110 (2007)
Duong, F. Binding, activation and dissociation of the dimeric SecA ATPase at the dimeric SecYEG translocase. EMBO J. 22, 4375–4384 (2003)
Harris, C. R. & Silhavy, T. J. Mapping an interface of SecY (PrlA) and SecE (PrlG) by using synthetic phenotypes and in vivo cross-linking. J. Bacteriol. 181, 3438–3444 (1999)
Tam, P. C., Maillard, A. P., Chan, K. K. & Duong, F. Investigating the SecY plug movement at the SecYEG translocation channel. EMBO J. 24, 3380–3388 (2005)
Plath, K., Mothes, W., Wilkinson, B. M., Stirling, C. J. & Rapoport, T. A. Signal sequence recognition in posttranslational protein transport across the yeast ER membrane. Cell 94, 795–807 (1998)
Brundage, L., Hendrick, J. P., Schiebel, E., Driessen, A. J. M. & Wickner, W. The purified E. coli integral membrane protein SecY/E is sufficient for reconstitution of SecA-dependent precursor proteintranslocation. Cell 62, 649–657 (1990)
Akimaru, J., Matsuyama, S. I., Tokuda, H. & Mizushima, S. Reconstitution of a protein translocation system containing purified SecY, SecE, and SecA from Escherichia coli . Proc. Natl Acad. Sci. USA 88, 6545–6549 (1991)
Economou, A. & Wickner, W. SecA promotes preprotein translocation by undergoing ATP-driven cycles of membrane insertion and deinsertion. Cell 78, 835–843 (1994)
Hunt, J. F. et al. Nucleotide control of interdomain interactions in the conformational reaction cycle of SecA. Science 297, 2018–2026 (2002)
Or, E., Navon, A. & Rapoport, T. Dissociation of the dimeric SecA ATPase during protein translocation across the bacterial membrane. EMBO J. 21, 4470–4479 (2002)
Or, E., Boyd, D., Gon, S., Beckwith, J. & Rapoport, T. The bacterial ATPase SecA functions as a monomer in protein translocation. J. Biol. Chem. 280, 9097–9105 (2005)
Alami, M., Dalal, K., Lelj-Garolla, B., Sligar, S. G. & Duong, F. Nanodiscs unravel the interaction between the SecYEG channel and its cytosolic partner SecA. EMBO J. 26, 1995–2004 (2007)
Jilaveanu, L. B., Zito, C. R. & Oliver, D. Dimeric SecA is essential for protein translocation. Proc. Natl Acad. Sci. USA 102, 7511–7516 (2005)
de Keyzer, J. et al. Covalently dimerized SecA is functional in protein translocation. J. Biol. Chem. 280, 35255–35260 (2005)
Mitra, K., Frank, J. & Driessen, A. Co- and post-translational translocation through the protein-conducting channel: analogous mechanisms at work? Nature Struct. Mol. Biol. 13, 957–964 (2006)
Osborne, A. R., Clemons, W. M. & Rapoport, T. A. A large conformational change of the translocation ATPase SecA. Proc. Natl Acad. Sci. USA 101, 10937–10942 (2004)
Zimmer, J., Li, W. & Rapoport, T. A. A novel dimer interface and conformational changes revealed by an X-ray structure of B. subtilis SecA. J. Mol. Biol. 364, 259–265 (2006)
Shiba, K., Ito, K., Yura, T. & Cerretti, D. P. A defined mutation in the protein export gene within the spc ribosomal protein operon of Escherichia coli: isolation and characterization of a new temperature-sensitive secY mutant. EMBO J. 3, 631–635 (1984)
Mori, H. & Ito, K. An essential amino acid residue in the protein translocation channel revealed by targeted random mutagenesis of SecY. Proc. Natl Acad. Sci. USA 98, 5128–5133 (2001)
Chiba, K., Mori, H. & Ito, K. Roles of the C-terminal end of SecY in protein translocation and viability of Escherichia coli . J. Bacteriol. 184, 2243–2250 (2002)
Mori, H., Shimizu, Y. & Ito, K. Superactive SecY variants that fulfill the essential translocation function with a reduced cellular quantity. J. Biol. Chem. 277, 48550–48557 (2002)
de Vrije, T., de Swart, R., Dowhan, W., Tommassen, J. & de Kruijff, B. Phosphatidylglycerol is involved in protein translocation across Escherichia coli inner membranes. Nature 334, 173–175 (1988)
Lill, R., Dowhan, W. & Wickner, W. The ATPase activity of SecA is regulated by acidic phospholipids, SecY, and the leader and mature domains of precursor proteins. Cell 60, 271–280 (1990)
Velankar, S. S., Soultanas, P., Dillingham, M. S., Subramanya, H. S. & Wigley, D. B. Crystal structures of complexes of PcrA DNA helicase with a DNA substrate indicate an inchworm mechanism. Cell 97, 75–84 (1999)
Nishiyama, K., Suzuki, T. & Tokuda, H. Inversion of the membrane topology of SecG coupled with SecA-dependent preprotein translocation. Cell 85, 71–81 (1996)
Satoh, Y., Matsumoto, G., Mori, H. & Ito, K. Nearest neighbor analysis of the SecYEG complex. 1. Identification of a SecY–SecG interface. Biochemistry 42, 7434–7441 (2003)
Breyton, C., Haase, W., Rapoport, T. A., Kuhlbrandt, W. & Collinson, I. Three-dimensional structure of the bacterial protein-translocation complex SecYEG. Nature 418, 662–665 (2002)
Bostina, M., Mohsin, B., Kuhlbrandt, W. & Collinson, I. Atomic model of the E. coli membrane-bound protein translocation complex SecYEG. J. Mol. Biol. 352, 1035–1043 (2005)
Cooper, D. B., Smith, V. F., Crane, J. M., Roth, H. C., Lilly, A. A. & Randall, L. L. SecA, the motor of the secretion machine, binds diverse partners on one interactive surface. J. Mol. Biol. 382, 74–87 (2008)
Hartl, F. U., Lecker, S., Schiebel, E., Hendrick, J. P. & Wickner, W. The binding cascade of SecB to SecA to SecY/E mediates preprotein targeting to the E. coli plasma membrane. Cell 63, 269–279 (1990)
Gelis, I. et al. Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell 131, 756–769 (2007)
Prinz, W. A., Spiess, C., Ehrmann, M., Schierle, C. & Beckwith, J. Targeting of signal sequenceless proteins for export in Escherichia coli with altered protein translocase. EMBO J. 15, 5209–5217 (1996)
Zhou, J. & Xu, Z. Structural determinants of SecB recognition by SecA in bacterial protein translocation. Nature Struct. Biol. 10, 942–947 (2003)
Heinrich, S. U., Mothes, W., Brunner, J. & Rapoport, T. A. The Sec61p complex mediates the integration of a membrane protein by allowing lipid partitioning of the transmembrane domain. Cell 102, 233–244 (2000)
Erlandson, K. J. et al. A role for the two-helix finger of the SecA ATPase in protein translocation. Nature doi: 10.1038/nature07439 (this issue)
Jarosik, G. P. & Oliver, D. B. Isolation and analysis of dominant SecA mutations in Escherichia coli . J. Bacteriol. 173, 860–868 (1991)
Karamanou, S. et al. A molecular switch in SecA protein couples ATP hydrolysis to protein translocation. Mol. Microbiol. 34, 1133–1145 (1999)
Wang, J. et al. Crystal structures of the HslVU peptidase–ATPase complex reveal an ATP-dependent proteolysis mechanism. Structure 9, 177–184 (2001)
Siddiqui, S. M., Sauer, R. T. & Baker, T. A. Role of the processing pore of the ClpX AAA+ ATPase in the recognition and engagement of specific protein substrates. Genes Dev. 18, 369–374 (2004)
Hinnerwisch, J., Fenton, W. A., Furtak, K. J., Farr, G. W. & Horwich, A. L. Loops in the central channel of ClpA chaperone mediate protein binding, unfolding, and translocation. Cell 121, 1029–1041 (2005)
DeLaBarre, B., Christianson, J. C., Kopito, R. R. & Brunger, A. T. Central pore residues mediate the p97/VCP activity required for ERAD. Mol. Cell 22, 451–462 (2006)
Martin, A., Baker, T. A. & Sauer, R. T. Diverse pore loops of the AAA+ ClpX machine mediate unassisted and adaptor-dependent recognition of ssrA-tagged substrates. Mol. Cell 29, 441–450 (2008)
Mori, H. & Ito, K. The long α-helix of SecA is important for the ATPase coupling of translocation. J. Biol. Chem. 281, 36249–36256 (2006)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Storoni, L. C., McCoy, A. J. & Read, R. J. Likelihood-enhanced fast rotation functions. Acta Crystallogr. D 60, 432–438 (2004)
McCoy, A. J., Grosse-Kunstleve, R. W., Storoni, L. C. & Read, R. J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D 61, 458–464 (2005)
Brunger, A. T. Version 1.2 of the Crystallography and NMR system. Nature Protoc. 2, 2728–2733 (2007)
Collaborative Computational Project, 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Cowtan, K. D. & Main, P. Phase combination and cross validation in iterated density-modification calculations. Acta Crystallogr. D 52, 43–48 (1996)
Cowtan, K. D. & Zhang, K. Y. Density modification for macromolecular phase improvement. Prog. Biophys. Mol. Biol. 72, 245–270 (1999)
Terwilliger, T. & Berendzen, J. Automated MAD and MIR structure determination. Acta Crystallogr. D 55, 849–861 (1999)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Read, R. J. Improved Fourier coefficients for maps using phases from partial structures with errors. Acta Crystallogr. A 42, 140–149 (1986)
DeLaBarre, B. & Brunger, A. T. Considerations for the refinement of low-resolution crystal structures. Acta Crystallogr. D 62, 923–932 (2006)
Chen, B. et al. Determining the structure of an unliganded and fully glycosylated SIV gp120 envelope glycoprotein. Structure 13, 197–211 (2005)
DeLano, W. L. The PyMOL Molecular Graphics System. <http://www.pymol.org> (2002)
Glaser, F. et al. ConSurf: identification of functional regions in proteins by surface-mapping of phylogenetic information. Bioinformatics 19, 163–164 (2003)
Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, W299–302 (2005)
Acknowledgements
We thank B. van den Berg and P. Bendapudi for initial experiments with the B. subtilis SecA and T. maritima SecY complex, G. Skiniotis for electron microscopy analysis, W. Li for his help with data processing, the staff at Advanced Photon Source beamlines ID-19 and 24ID and at Brookhaven National Laboratory beamline X29, and the SBGrid consortium at Harvard Medical School. We thank A. Brunger and S. Harrison for comments, and A. Brunger, B. van den Berg, W. Li, A. Osborne and S. Schulman for critical reading of the manuscript. The work was supported by a National Institutes of Health grant. T.A.R. is an HHMI investigator. Y.N. is supported by the Damon Runyon Cancer Research Foundation (DRG-#1953-07).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S12 with Legends and a Supplementary Discussion (PDF 5672 kb)
Rights and permissions
About this article
Cite this article
Zimmer, J., Nam, Y. & Rapoport, T. Structure of a complex of the ATPase SecA and the protein-translocation channel. Nature 455, 936–943 (2008). https://doi.org/10.1038/nature07335
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature07335
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.