Abstract
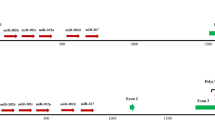
MicroRNAs (miRNAs) are short RNAs that direct messenger RNA degradation or disrupt mRNA translation in a sequence-dependent manner1,2,3,4,5,6,7. For more than a decade, attempts to study the interaction of miRNAs with their targets were confined to the 3′ untranslated regions of mRNAs1, fuelling an underlying assumption that these regions are the principal recipients of miRNA activity. Here we focus on the mouse Nanog, Oct4 (also known as Pou5f1) and Sox2 genes8,9,10,11 and demonstrate the existence of many naturally occurring miRNA targets in their amino acid coding sequence (CDS). Some of the mouse targets analysed do not contain the miRNA seed, whereas others span exon–exon junctions or are not conserved in the human and rhesus genomes. miR-134, miR-296 and miR-470, upregulated on retinoic-acid-induced differentiation of mouse embryonic stem cells, target the CDS of each transcription factor in various combinations, leading to transcriptional and morphological changes characteristic of differentiating mouse embryonic stem cells, and resulting in a new phenotype. Silent mutations at the predicted targets abolish miRNA activity, prevent the downregulation of the corresponding genes and delay the induced phenotype. Our findings demonstrate the abundance of CDS-located miRNA targets, some of which can be species-specific, and support an augmented model whereby animal miRNAs exercise their control on mRNAs through targets that can reside beyond the 3′ untranslated region.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004)
Elbashir, S. M., Lendeckel, W. & Tuschl, T. RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev. 15, 188–200 (2001)
Hammond, S. M. Dicing and slicing: the core machinery of the RNA interference pathway. FEBS Lett. 579, 5822–5829 (2005)
Hannon, G. J. RNA interference. Nature 418, 244–251 (2002)
Lagos-Quintana, M., Rauhut, R., Lendeckel, W. & Tuschl, T. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001)
Pillai, R. S., Bhattacharyya, S. N. & Filipowicz, W. Repression of protein synthesis by miRNAs: how many mechanisms? Trends Cell Biol. 17, 118–126 (2007)
Pillai, R. S. MicroRNA function: multiple mechanisms for a tiny RNA? RNA 11, 1753–1761 (2005)
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nature Genet. 24, 372–376 (2000)
Okumura-Nakanishi, S., Saito, M., Niwa, H. & Ishikawa, F. Oct-3/4 and Sox2 regulate Oct-3/4 gene in embryonic stem cells. J. Biol. Chem. 280, 5307–5317 (2005)
Pan, G. & Thomson, J. A. Nanog and transcriptional networks in embryonic stem cell pluripotency. Cell Res. 17, 42–49 (2007)
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003)
Slack, F. J. et al. The lin-41 RBCC gene acts in the C. elegans heterochronic pathway between the let-7 regulatory RNA and the LIN-29 transcription factor. Mol. Cell 5, 659–669 (2000)
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14 . Cell 75, 843–854 (1993)
Easow, G., Teleman, A. A. & Cohen, S. M. Isolation of microRNA targets by miRNP immunopurification. RNA 13, 1198–1204 (2007)
Lytle, J. R., Yario, T. A. & Steitz, J. A. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5′ UTR as in the 3′ UTR. Proc. Natl Acad. Sci. USA 104, 9667–9672 (2007)
Bentwich, I. Prediction and validation of microRNAs and their targets. FEBS Lett. 579, 5904–5910 (2005)
Rajewsky, N. microRNA target predictions in animals. Nature Genet. 38, S8–S13 (2006)
Miranda, K. C. et al. A pattern-based method for the identification of microRNA binding sites and their corresponding heteroduplexes. Cell 126, 1203–1217 (2006)
Kloosterman, W. P., Wienholds, E., Ketting, R. F. & Plasterk, R. H. Substrate requirements for let-7 function in the developing zebrafish embryo. Nucleic Acids Res. 32, 6284–6291 (2004)
Silva, J., Chambers, I., Pollard, S. & Smith, A. Nanog promotes transfer of pluripotency after cell fusion. Nature 441, 997–1001 (2006)
Loh, Y. H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nature Genet. 38, 431–440 (2006)
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003)
Tay, Y. M. et al. MicroRNA-134 modulates the differentiation of mouse embryonic stem cells, where it causes post-transcriptional attenuation of Nanog and LRH1. Stem Cells 26, 17–29 (2008)
Palmqvist, L. et al. Correlation of murine embryonic stem cell gene expression profiles with functional measures of pluripotency. Stem Cells 23, 663–680 (2005)
Duursma, A. M., Kedde, M., Schrier, M., le Sage, C. & Agami, R. miR-148 targets human DNMT3b protein coding region. RNA 14, 872–877 (2008)
Lal, A. et al. p16INK4atranslation suppressed by miR-24. PLoS ONE 3, e1864 (2008)
Hobert, O. Gene regulation by transcription factors and microRNAs. Science 319, 1785–1786 (2008)
Floyd, S. K. & Bowman, J. L. Gene regulation: ancient microRNA target sequences in plants. Nature 428, 485–486 (2004)
Didiano, D. & Hobert, O. Perfect seed pairing is not a generally reliable predictor for miRNA-target interactions. Nat. Struct. Mol. Biol. 13, 849–851 (2006)
Thomson, A. M. et al. The acute box cis-element in human heavy ferritin mRNA 5′-untranslated region is a unique translation enhancer that binds poly(C)-binding proteins. J. Biol. Chem. 280, 30032–30045 (2005)
Acknowledgements
We thank T. Huynh for assistance with building the rna22 pipeline that was used to make the miRNA target predictions, and H. Zhang and Y. L. Lee for their help with some of the constructs. The work of Y.T., J.Z., A.M.T. and B.L. was supported by the Agency for Science, Technology and Research of Singapore. Y.T. was the recipient of an A*STAR graduate scholarship. B.L. was also partially supported by National Institutes of Health grants DK47636 and AI54973.
Author Contributions Y.T. carried out the experiments and assisted with writing the paper. J.Z. assisted with the experiments. A.M.T. assisted with and supervised the experiments, and assisted with writing the paper. B.L. supervised the experiments and supported the experimental aspects of the research. I.R. designed the research and experiments, carried out the computational analyses and wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Discussion, Supplementary Table 1, Supplementary Figures 1-13 and References (PDF 3522 kb)
Rights and permissions
About this article
Cite this article
Tay, Y., Zhang, J., Thomson, A. et al. MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature 455, 1124–1128 (2008). https://doi.org/10.1038/nature07299
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07299
This article is cited by
-
Non-coding RNAs in disease: from mechanisms to therapeutics
Nature Reviews Genetics (2024)
-
Oligonucleotide therapeutics and their chemical modification strategies for clinical applications
Journal of Pharmaceutical Investigation (2024)
-
Inhibition of miR-101-3p prevents human aortic valve interstitial cell calcification through regulation of CDH11/SOX9 expression
Molecular Medicine (2023)
-
Epigenetic regulation in hematopoiesis and its implications in the targeted therapy of hematologic malignancies
Signal Transduction and Targeted Therapy (2023)
-
Base editing-mediated one-step inactivation of the Dnmt gene family reveals critical roles of DNA methylation during mouse gastrulation
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.