Abstract
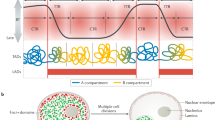
Genome stability requires one, and only one, DNA duplication at each S phase. The mechanisms preventing origin firing on newly replicated DNA are well documented1, but much less is known about the mechanisms controlling the spacing of initiation events2,3, namely the completion of DNA replication. Here we show that origin use in Chinese hamster cells depends on both the movement of the replication forks and the organization of chromatin loops. We found that slowing the replication speed triggers the recruitment of latent origins within minutes, allowing the completion of S phase in a timely fashion. When slowly replicating cells are shifted to conditions of fast fork progression, although the decrease in the overall number of active origins occurs within 2 h, the cells still have to go through a complete cell cycle before the efficiency specific to each origin is restored. We observed a strict correlation between replication speed during a given S phase and the size of chromatin loops in the next G1 phase. Furthermore, we found that origins located at or near sites of anchorage of chromatin loops in G1 are activated preferentially in the following S phase. These data suggest a mechanism of origin programming in which replication speed determines the spacing of anchorage regions of chromatin loops, that, in turn, controls the choice of initiation sites.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Blow, J. J. & Dutta, A. Preventing re-replication of chromosomal DNA. Nature Rev. Mol. Cell Biol. 6, 476–486 (2005)
Gilbert, D. M. Replication origin plasticity, Taylor-made: inhibition vs recruitment of origins under conditions of replication stress. Chromosoma 116, 341–347 (2007)
Anglana, M., Apiou, F., Bensimon, A. & Debatisse, M. Dynamics of DNA replication in mammalian somatic cells: nucleotide pool modulates origin choice and interorigin spacing. Cell 114, 385–394 (2003)
Debatisse, M., Berry, M. & Buttin, G. Stepwise isolation and properties of unstable Chinese hamster cell variants that overproduce adenylate deaminase. Mol. Cell. Biol. 2, 1346–1353 (1982)
Toledo, F. et al. oriGNAI3: a narrow zone of preferential replication initiation in mammalian cells identified by 2D gel and competitive PCR replicon mapping techniques. Nucleic Acids Res. 26, 2313–2321 (1998)
Toledo, F., Lachages, A. M., Mayau, V. & Debatisse, M. Initiation of DNA replication at the Chinese hamster origin oriGNAI3 relies on local sequences and/or chromatin structures, but not on transcription of the nearby GNAI3 gene. Nucleic Acids Res. 27, 1600–1608 (1999)
Svetlova, E. Y., Razin, S. V. & Debatisse, M. Mammalian recombination hot spot in a DNA loop anchorage region: A model for the study of common fragile sites. J. Cell. Biochem. 81, 170–178 (2001)
DePamphilis, M. L. et al. Regulating the licensing of DNA replication origins in metazoa. Curr. Opin. Cell Biol. 18, 231–239 (2006)
Berezney, R., Dubey, D. D. & Huberman, J. A. Heterogeneity of eukaryotic replicons, replicon clusters, and replication foci. Chromosoma 108, 471–484 (2000)
Pasero, P., Bensimon, A. & Schwob, E. Single-molecule analysis reveals clustering and epigenetic regulation of replication origins at the yeast rDNA locus. Genes Dev. 16, 2479–2484 (2002)
Lebofsky, R., Heilig, R., Sonnleitner, M., Weissenbach, J. & Bensimon, A. DNA replication origin interference increases the spacing between initiation events in human cells. Mol. Biol. Cell 17, 5337–5345 (2006)
Buongiorno-Nardelli, M., Micheli, G., Carri, M. T. & Marilley, M. A relationship between replicon size and supercoiled loop domains in the eukaryotic genome. Nature 298, 100–102 (1982)
Hancock, R. Internal organisation of the nucleus: assembly of compartments by macromolecular crowding and the nuclear matrix model. Biol. Cell 96, 595–601 (2004)
Anachkova, B., Djeliova, V. & Russev, G. Nuclear matrix support of DNA replication. J. Cell. Biochem. 96, 951–961 (2005)
Jenke, A. C. et al. Nuclear scaffold/matrix attached region modules linked to a transcription unit are sufficient for replication and maintenance of a mammalian episome. Proc. Natl Acad. Sci. USA 101, 11322–11327 (2004)
Fernandez, M. A. et al. Matrix attachment regions and transcription units in a polygenic mammalian locus overlapping two isochores. J. Cell. Biochem. 67, 541–551 (1997)
Razin, S. V. The nuclear matrix and chromosomal DNA loops: is their any correlation between partitioning of the genome into loops and functional domains? Cell. Mol. Biol. Lett. 6, 59–69 (2001)
Pardoll, D. M., Vogelstein, B. & Coffey, D. S. A fixed site of DNA replication in eucaryotic cells. Cell 19, 527–536 (1980)
Kitamura, E., Blow, J. J. & Tanaka, T. U. Live-cell imaging reveals replication of individual replicons in eukaryotic replication factories. Cell 125, 1297–1308 (2006)
Lawlis, S. J., Keezer, S. M., Wu, J. R. & Gilbert, D. M. Chromosome architecture can dictate site-specific initiation of DNA replication in Xenopus egg extracts. J. Cell Biol. 135, 1207–1218 (1996)
Lemaitre, J. M., Danis, E., Pasero, P., Vassetzky, Y. & Mechali, M. Mitotic remodeling of the replicon and chromosome structure. Cell 123, 787–801 (2005)
Dietzel, S. & Belmont, A. S. Reproducible but dynamic positioning of DNA in chromosomes during mitosis. Nature Cell Biol. 3, 767–770 (2001)
Wu, J. R. & Gilbert, D. M. A distinct G1 step required to specify the Chinese hamster DHFR replication origin. Science 271, 1270–1272 (1996)
Li, F., Chen, J., Solessio, E. & Gilbert, D. M. Spatial distribution and specification of mammalian replication origins during G1 phase. J. Cell Biol. 161, 257–266 (2003)
El Achkar, E., Gerbault-Seureau, M., Muleris, M., Dutrillaux, B. & Debatisse, M. Premature condensation induces breaks at the interface of early and late replicating chromosome bands bearing common fragile sites. Proc. Natl Acad. Sci. USA 102, 18069–18074 (2005)
Gilbert, D. M., Miyazawa, H. & DePamphilis, M. L. Site-specific initiation of DNA replication in Xenopus egg extract requires nuclear structure. Mol. Cell. Biol. 15, 2942–2954 (1995)
Merrick, C. J., Jackson, D. & Diffley, J. F. Visualization of altered replication dynamics after DNA damage in human cells. J. Biol. Chem. 279, 20067–20075 (2004)
Acknowledgements
We thank E. Blackburn, R. Rothstein and F. Toledo for discussions and critical reading of the manuscript, and Genomic Vision for making available the DNA combing technology. S.C. is supported by a grant from the ARC (Association pour la Recherche sur le Cancer), and S.G. and N.A. are supported by a grant from the Ministère de la Recherche. The M.D. team is supported by La Ligue Nationale contre le Cancer and the Agence Nationale de la Recherche (ANR).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-8 with Legends. The Supplementary Figure 1 includes a model for the control of origin usage in mammalian somatic cells. (PDF 6942 kb)
Rights and permissions
About this article
Cite this article
Courbet, S., Gay, S., Arnoult, N. et al. Replication fork movement sets chromatin loop size and origin choice in mammalian cells. Nature 455, 557–560 (2008). https://doi.org/10.1038/nature07233
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07233
This article is cited by
-
Detection and characterization of constitutive replication origins defined by DNA polymerase epsilon
BMC Biology (2023)
-
Transcription-coupled structural dynamics of topologically associating domains regulate replication origin efficiency
Genome Biology (2021)
-
Autophagy buffers Ras-induced genotoxic stress enabling malignant transformation in keratinocytes primed by human papillomavirus
Cell Death & Disease (2021)
-
Sonic hedgehog accelerates DNA replication to cause replication stress promoting cancer initiation in medulloblastoma
Nature Cancer (2020)
-
Chromatin conformation regulates the coordination between DNA replication and transcription
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.