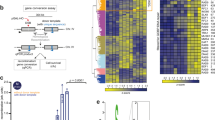
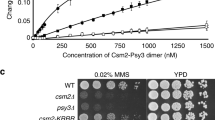
Abstract
DNA double-strand breaks (DSBs) are repaired by two principal mechanisms: non-homologous end-joining (NHEJ) and homologous recombination (HR)1. HR is the most accurate DSB repair mechanism but is generally restricted to the S and G2 phases of the cell cycle, when DNA has been replicated and a sister chromatid is available as a repair template2,3,4,5. By contrast, NHEJ operates throughout the cell cycle but assumes most importance in G1 (refs 4, 6). The choice between repair pathways is governed by cyclin-dependent protein kinases (CDKs)2,3,5,7, with a major site of control being at the level of DSB resection, an event that is necessary for HR but not NHEJ, and which takes place most effectively in S and G2 (refs 2, 5). Here we establish that cell-cycle control of DSB resection in Saccharomyces cerevisiae results from the phosphorylation by CDK of an evolutionarily conserved motif in the Sae2 protein. We show that mutating Ser 267 of Sae2 to a non-phosphorylatable residue causes phenotypes comparable to those of a sae2Δ null mutant, including hypersensitivity to camptothecin, defective sporulation, reduced hairpin-induced recombination, severely impaired DNA-end processing and faulty assembly and disassembly of HR factors. Furthermore, a Sae2 mutation that mimics constitutive Ser 267 phosphorylation complements these phenotypes and overcomes the necessity of CDK activity for DSB resection. The Sae2 mutations also cause cell-cycle-stage specific hypersensitivity to DNA damage and affect the balance between HR and NHEJ. These findings therefore provide a mechanistic basis for cell-cycle control of DSB repair and highlight the importance of regulating DSB resection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shrivastav, M., De Haro, L. P. & Nickoloff, J. A. Regulation of DNA double-strand break repair pathway choice. Cell Res. 18, 134–147 (2008)
Aylon, Y., Liefshitz, B. & Kupiec, M. The CDK regulates repair of double-strand breaks by homologous recombination during the cell cycle. EMBO J. 23, 4868–4875 (2004)
Caspari, T., Murray, J. M. & Carr, A. M. Cdc2-cyclin B kinase activity links Crb2 and Rqh1-topoisomerase III. Genes Dev. 16, 1195–1208 (2002)
Hinz, J. M., Yamada, N. A., Salazar, E. P., Tebbs, R. S. & Thompson, L. H. Influence of double-strand-break repair pathways on radiosensitivity throughout the cell cycle in CHO cells. DNA Repair (Amst.) 4, 782–792 (2005)
Ira, G. et al. DNA end resection, homologous recombination and DNA damage checkpoint activation require CDK1. Nature 431, 1011–1017 (2004)
Karathanasis, E. & Wilson, T. E. Enhancement of Saccharomyces cerevisiae end-joining efficiency by cell growth stage but not by impairment of recombination. Genetics 161, 1015–1027 (2002)
Esashi, F. et al. CDK-dependent phosphorylation of BRCA2 as a regulatory mechanism for recombinational repair. Nature 434, 598–604 (2005)
Aylon, Y. & Kupiec, M. DSB repair: the yeast paradigm. DNA Repair (Amst.) 3, 797–815 (2004)
Clerici, M., Mantiero, D., Lucchini, G. & Longhese, M. P. The Saccharomyces cerevisiae Sae2 protein promotes resection and bridging of double strand break ends. J. Biol. Chem. 280, 38631–38638 (2005)
McKee, A. H. & Kleckner, N. A general method for identifying recessive diploid-specific mutations in Saccharomyces cerevisiae, its application to the isolation of mutants blocked at intermediate stages of meiotic prophase and characterization of a new gene SAE2 . Genetics 146, 797–816 (1997)
Neale, M. J., Pan, J. & Keeney, S. Endonucleolytic processing of covalent protein-linked DNA double-strand breaks. Nature 436, 1053–1057 (2005)
Prinz, S., Amon, A. & Klein, F. Isolation of COM1, a new gene required to complete meiotic double-strand break-induced recombination in Saccharomyces cerevisiae . Genetics 146, 781–795 (1997)
Lengsfeld, B. M., Rattray, A. J., Bhaskara, V., Ghirlando, R. & Paull, T. T. Sae2 is an endonuclease that processes hairpin DNA cooperatively with the Mre11/Rad50/Xrs2 complex. Mol. Cell 28, 638–651 (2007)
Baroni, E., Viscardi, V., Cartagena-Lirola, H., Lucchini, G. & Longhese, M. P. The functions of budding yeast Sae2 in the DNA damage response require Mec1- and Tel1-dependent phosphorylation. Mol. Cell. Biol. 24, 4151–4165 (2004)
Mendenhall, M. D. & Hodge, A. E. Regulation of Cdc28 cyclin-dependent protein kinase activity during the cell cycle of the yeast Saccharomyces cerevisiae . Microbiol. Mol. Biol. Rev. 62, 1191–1243 (1998)
Penkner, A. et al. A conserved function for a Caenorhabditis elegans Com1/Sae2/CtIP protein homolog in meiotic recombination. EMBO J. 26, 5071–5082 (2007)
Sartori, A. A. et al. Human CtIP promotes DNA end resection. Nature 450, 509–514 (2007)
Uanschou, C. et al. A novel plant gene essential for meiosis is related to the human CtIP and the yeast COM1/SAE2 gene. EMBO J. 26, 5061–5070 (2007)
Pommier, Y. Topoisomerase I inhibitors: camptothecins and beyond. Nature Rev. Cancer 6, 789–802 (2006)
Chen, J., Saha, P., Kornbluth, S., Dynlacht, B. D. & Dutta, A. Cyclin-binding motifs are essential for the function of p21CIP1 . Mol. Cell. Biol. 16, 4673–4682 (1996)
Lobachev, K. S., Gordenin, D. A. & Resnick, M. A. The Mre11 complex is required for repair of hairpin-capped double-strand breaks and prevention of chromosome rearrangements. Cell 108, 183–193 (2002)
Sugawara, N. & Haber, J. E. Repair of DNA double strand breaks: in vivo biochemistry. Methods Enzymol. 408, 416–429 (2006)
Bishop, A. C. et al. A chemical switch for inhibitor-sensitive alleles of any protein kinase. Nature 407, 395–401 (2000)
Lazzaro, F. et al. Histone methyltransferase Dot1 and Rad9 inhibit single-stranded DNA accumulation at DSBs and uncapped telomeres. EMBO J. 27, 1502–1512 (2008)
Lisby, M., Barlow, J. H., Burgess, R. C. & Rothstein, R. Choreography of the DNA damage response: spatiotemporal relationships among checkpoint and repair proteins. Cell 118, 699–713 (2004)
Rattray, A. J., McGill, C. B., Shafer, B. K. & Strathern, J. N. Fidelity of mitotic double-strand-break repair in Saccharomyces cerevisiae: a role for SAE2/COM1 . Genetics 158, 109–122 (2001)
Cortes-Ledesma, F. & Aguilera, A. Double-strand breaks arising by replication through a nick are repaired by cohesin-dependent sister-chromatid exchange. EMBO Rep. 7, 919–926 (2006)
Boulton, S. J. & Jackson, S. P. Saccharomyces cerevisiae Ku70 potentiates illegitimate DNA double-strand break repair and serves as a barrier to error-prone DNA repair pathways. EMBO J. 15, 5093–5103 (1996)
Lee, K. & Lee, S. E. Saccharomyces cerevisiae Sae2- and Tel1-dependent single-strand DNA formation at DNA break promotes microhomology-mediated end joining. Genetics 176, 2003–2014 (2007)
Limbo, O. et al. Ctp1 is a cell-cycle-regulated protein that functions with Mre11 complex to control double-strand break repair by homologous recombination. Mol. Cell 28, 134–146 (2007)
Puig, O. et al. The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods 24, 218–229 (2001)
Acknowledgements
We thank M. P. Longhese, R. Rothstein, K. Lobachev, M. Lichten and M. Foiani for providing strains, and R. Driscoll, S. Gravel, K. Dry and K. Miller for helpful discussions and comments on the manuscript. P.H. is the recipient of a Long-Term EMBO Fellowship. A.A.S. is supported by a Swiss National Foundation Grant. The S.P.J. laboratory is supported by grants from Cancer Research UK and the European Community (Integrated Project DNA repair, grant LSHG-CT-2005-512113). The A.A. laboratory is supported by grants from the Spanish Ministry of Science and Education (BFU2006-05260 and CDS2007-0015) and Junta de Andalucia (CVI624).
Author Contributions A.A.S. identified the homology between Sae2 and CtIP, cloned SAE2 into pGEX-4T1 and made the original sae2-S267A and sae2-S267E mutations. All the experiments shown were performed by P.H. and were conceived by P.H. and S.P.J., except those on SCR analyses that were performed by F.C.-L. and conceived by F.C.-L. and A.A. P.H. and S.P.J. wrote the paper. All authors discussed and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
The file contains Supplementary Table 1 and Supplementary Figures 1-4 with Legends. (PDF 3649 kb)
Rights and permissions
About this article
Cite this article
Huertas, P., Cortés-Ledesma, F., Sartori, A. et al. CDK targets Sae2 to control DNA-end resection and homologous recombination. Nature 455, 689–692 (2008). https://doi.org/10.1038/nature07215
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07215
This article is cited by
-
Cyclers’ kinases in cell division: from molecules to cancer therapy
Cell Death & Differentiation (2023)
-
Cyclins and cyclin-dependent kinases: from biology to tumorigenesis and therapeutic opportunities
Journal of Cancer Research and Clinical Oncology (2023)
-
Mre11-Rad50 oligomerization promotes DNA double-strand break repair
Nature Communications (2022)
-
METTL16 antagonizes MRE11-mediated DNA end resection and confers synthetic lethality to PARP inhibition in pancreatic ductal adenocarcinoma
Nature Cancer (2022)
-
RNF19A-mediated ubiquitination of BARD1 prevents BRCA1/BARD1-dependent homologous recombination
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.