Abstract
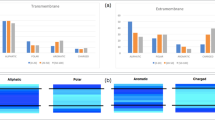
Understanding the energetics of molecular interactions is fundamental to all of the central quests of structural biology including structure prediction and design, mapping evolutionary pathways, learning how mutations cause disease, drug design, and relating structure to function. Hydrogen-bonding is widely regarded as an important force in a membrane environment because of the low dielectric constant of membranes and a lack of competition from water1,2,3,4,5,6. Indeed, polar residue substitutions are the most common disease-causing mutations in membrane proteins6,7. Because of limited structural information and technical challenges, however, there have been few quantitative tests of hydrogen-bond strength in the context of large membrane proteins. Here we show, by using a double-mutant cycle analysis, that the average contribution of eight interhelical side-chain hydrogen-bonding interactions throughout bacteriorhodopsin is only 0.6 kcal mol-1. In agreement with these experiments, we find that 4% of polar atoms in the non-polar core regions of membrane proteins have no hydrogen-bond partner and the lengths of buried hydrogen bonds in soluble proteins and membrane protein transmembrane regions are statistically identical. Our results indicate that most hydrogen-bond interactions in membrane proteins are only modestly stabilizing. Weak hydrogen-bonding should be reflected in considerations of membrane protein folding, dynamics, design, evolution and function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
White, S. H. How hydrogen bonds shape membrane protein structure. Adv. Protein Chem. 72, 157–172 (2005)
Popot, J. L. & Engelman, D. M. Helical membrane protein folding, stability, and evolution. Annu. Rev. Biochem. 69, 881–922 (2000)
Zhou, F. X., Merianos, H. J., Brunger, A. T. & Engelman, D. M. Polar residues drive association of polyleucine transmembrane helices. Proc. Natl Acad. Sci. USA 98, 2250–2255 (2001)
Gratkowski, H., Lear, J. D. & DeGrado, W. F. Polar side chains drive the association of model transmembrane peptides. Proc. Natl Acad. Sci. USA 98, 880–885 (2001)
Adamian, L. & Liang, J. Interhelical hydrogen bonds and spatial motifs in membrane proteins: polar clamps and serine zippers. Proteins 47, 209–218 (2002)
Partridge, A. W., Therien, A. G. & Deber, C. M. Polar mutations in membrane proteins as a biophysical basis for disease. Biopolymers 66, 350–358 (2002)
Partridge, A. W., Therien, A. G. & Deber, C. M. Missense mutations in transmembrane domains of proteins: phenotypic propensity of polar residues for human disease. Proteins 54, 648–656 (2004)
Call, M. E. et al. The structure of the ζζ transmembrane dimer reveals features essential for its assembly with the T cell receptor. Cell 127, 355–368 (2006)
Faham, S. et al. Side-chain contributions to membrane protein structure and stability. J. Mol. Biol. 335, 297–305 (2004)
Duong, M. T., Jaszewski, T. M., Fleming, K. G. & MacKenzie, K. R. Changes in apparent free energy of helix–helix dimerization in a biological membrane due to point mutations. J. Mol. Biol. 371, 422–434 (2007)
Myers, J. K. & Pace, C. N. Hydrogen bonding stabilizes globular proteins. Biophys. J. 71, 2033–2039 (1996)
Busenlehner, L. S. & Armstrong, R. N. Insights into enzyme structure and dynamics elucidated by amide H/D exchange mass spectrometry. Arch. Biochem. Biophys. 433, 34–46 (2005)
Busenlehner, L. S. et al. Stress sensor triggers conformational response of the integral membrane protein microsomal glutathione transferase 1. Biochemistry 43, 11145–11152 (2004)
Molday, R. S., Englander, S. W. & Kallen, R. G. Primary structure effects on peptide group hydrogen exchange. Biochemistry 11, 150–158 (1972)
O’Neil, J. D. & Sykes, B. D. NMR studies of the influence of dodecyl sulfate on the amide hydrogen exchange kinetics of a micelle-solubilized hydrophobic tripeptide. Biochemistry 28, 699–707 (1989)
Hong, H., Szabo, G. & Tamm, L. K. Electrostatic couplings in OmpA ion-channel gating suggest a mechanism for pore opening. Nature Chem. Biol. 2, 627–635 (2006)
Fersht, A. R., Matouschek, A. & Serrano, L. The folding of an enzyme. I. Theory of protein engineering analysis of stability and pathway of protein folding. J. Mol. Biol. 224, 771–782 (1992)
Fleming, P. J. & Rose, G. D. Do all backbone polar groups in proteins form hydrogen bonds? Protein Sci. 14, 1911–1917 (2005)
Adamian, L., Nanda, V., DeGrado, W. F. & Liang, J. Empirical lipid propensities of amino acid residues in multispan alpha helical membrane proteins. Proteins 59, 496–509 (2005)
Rees, D., DeAntonio, L. & Eisenberg, D. Hydrophobic organization of membrane proteins. Science 245, 510–513 (1989)
Stanley, A. M. & Fleming, K. G. The role of a hydrogen bonding network in the transmembrane β-barrel OMPLA. J. Mol. Biol. 370, 912–924 (2007)
Fleming, K. G. & Engelman, D. M. Specificity in transmembrane helix–helix interactions can define a hierarchy of stability for sequence variants. Proc. Natl Acad. Sci. USA 98, 14340–14344 (2001)
Senes, A., Engel, D. E. & DeGrado, W. F. Folding of helical membrane proteins: the role of polar, GxxxG-like and proline motifs. Curr. Opin. Struct. Biol. 14, 465–479 (2004)
Lear, J. D., Gratkowski, H., Adamian, L., Liang, J. & DeGrado, W. F. Position-dependence of stabilizing polar interactions of asparagine in transmembrane helical bundles. Biochemistry 42, 6400–6407 (2003)
Rosenbaum, D. M. et al. GPCR engineering yields high-resolution structural insights into β2-adrenergic receptor function. Science 318, 1266–1273 (2007)
Shan, S. O. & Herschlag, D. Hydrogen bonding in enzymatic catalysis: analysis of energetic contributions. Methods Enzymol. 308, 246–276 (1999)
Faham, S. & Bowie, J. U. Bicelle crystallization: a new method for crystallizing membrane proteins yields a monomeric bacteriorhodopsin structure. J. Mol. Biol. 316, 1–6 (2002)
Chamberlain, A. K., Lee, Y., Kim, S. & Bowie, J. U. Snorkeling preferences foster an amino acid composition bias in transmembrane helices. J. Mol. Biol. 339, 471–479 (2004)
McDonald, I. K. & Thornton, J. M. Satisfying hydrogen bonding potential in proteins. J. Mol. Biol. 238, 777–793 (1994)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998)
Hamuro, Y. et al. Dynamics of cAPK type IIβ activation revealed by enhanced amide H/2H exchange mass spectrometry (DXMS). J. Mol. Biol. 327, 1065–1076 (2003)
Zhang, Z. & Marshall, A. G. A universal algorithm for fast and automated charge state deconvolution of electrospray mass-to-charge ratio spectra. J. Am. Soc. Mass Spectrom. 9, 225–233 (1998)
Zhang, Z., Li, W., Logan, T. M., Li, M. & Marshall, A. G. Human recombinant [C22A] FK506-binding protein amide hydrogen exchange rates from mass spectrometry match and extend those from NMR. Protein Sci. 6, 2203–2217 (1997)
Luecke, H., Schobert, B., Richter, H. T., Cartailler, J. P. & Lanyi, J. K. Structure of bacteriorhodopsin at 1.55 Å resolution. J. Mol. Biol. 291, 899–911 (1999)
Jormakka, M., Tornroth, S., Byrne, B. & Iwata, S. Molecular basis of proton motive force generation: structure of formate dehydrogenase-N. Science 295, 1863–1868 (2002)
Khademi, S. et al. Mechanism of ammonia transport by Amt/MEP/Rh: structure of AmtB at 1.35 Å. Science 305, 1587–1594 (2004)
Yamashita, A., Singh, S. K., Kawate, T., Jin, Y. & Gouaux, E. Crystal structure of a bacterial homologue of Na+/Cl–-dependent neurotransmitter transporters. Nature 437, 215–223 (2005)
Andrade, S. L., Dickmanns, A., Ficner, R. & Einsle, O. Crystal structure of the archaeal ammonium transporter Amt-1 from Archaeoglobus fulgidus . Proc. Natl Acad. Sci. USA 102, 14994–14999 (2005)
Lees, A. M., Deconinck, A. E., Campbell, B. D. & Lees, R. S. Atherin: a newly identified, lesion-specific, LDL-binding protein in human atherosclerosis. Atherosclerosis 182, 219–230 (2005)
Acknowledgements
We thank the staff at the beamlines 8.2.1 and 8.2.2 at the Advanced Light Source; F. Pettit for advice on statistics; M. Philips for assisting with circular dichroism experiments; S. Bassilian for assisting with LC–MS analysis of intact bacteriorhodopsin; Y. Ihm for the identification of transmembrane regions; Z. Zhang for providing deuterium-exchange data reduction software MAGTRAN and LAPLACE; N. L. Kelleher for providing the acid-labile detergent; and M. Chamberlain, H. Cheng, E. Gendel, H. Hong, Y. Ihm, A. D. Meruelo, T. Mitchell and R. Stafford for critically reading the manuscript. This work was supported by National Institutes of Health grant RO1 GM063919 (J.U.B.) and by the National Institutes of Health National Cancer Institute Innovative Molecular Analysis Technologies Program (V.L.W.).
Author Contributions N.H.J. and J.U.B. designed the research and prepared the manuscript. N.H.J. performed the vast majority of the experiments and structure analyses. A.M. and D.Y. assisted with mutagenesis and protein purification. A.M. crystallized the T90A/D115A mutant. S.F. collected and processed some diffraction data and helped with structure determination and refinement. J.P.W. assisted site-directed mutagenesis verification by LC–MS analysis of intact bacteriorhodopsin, provided technical advice on mass spectrometry, and helped develop the H/D exchange method. V.L.W. assisted in H/D exchange data analysis, including the provision of specialized software and hardware, and provided help with H/D exchange methods.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
The file contains Supplementary Methods, Supplementary Tables 1-2, Supplementary Figures 1-3 with legends, and additional references. (PDF 478 kb)
Rights and permissions
About this article
Cite this article
Joh, N., Min, A., Faham, S. et al. Modest stabilization by most hydrogen-bonded side-chain interactions in membrane proteins. Nature 453, 1266–1270 (2008). https://doi.org/10.1038/nature06977
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06977
This article is cited by
-
Crystalline hydrogen bonding of water molecules confined in a metal-organic framework
Communications Chemistry (2022)
-
Aion is a bistable anion-conducting channelrhodopsin that provides temporally extended and reversible neuronal silencing
Communications Biology (2022)
-
Computational Simulation of Holin S105 in Membrane Bilayer and Its Dimerization Through a Helix-Turn-Helix Motif
The Journal of Membrane Biology (2021)
-
The roles and applications of chaotropes and kosmotropes in industrial fermentation processes
World Journal of Microbiology and Biotechnology (2020)
-
MerMAIDs: a family of metagenomically discovered marine anion-conducting and intensely desensitizing channelrhodopsins
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.