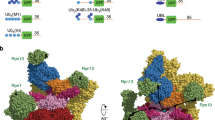
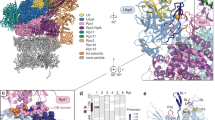
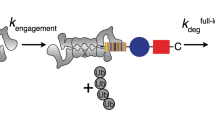
Abstract
Proteasomal receptors that recognize ubiquitin chains attached to substrates are key mediators of selective protein degradation in eukaryotes. Here we report the identification of a new ubiquitin receptor, Rpn13/ARM1, a known component of the proteasome. Rpn13 binds ubiquitin through a conserved amino-terminal region termed the pleckstrin-like receptor for ubiquitin (Pru) domain, which binds K48-linked diubiquitin with an affinity of approximately 90 nM. Like proteasomal ubiquitin receptor Rpn10/S5a, Rpn13 also binds ubiquitin-like (UBL) domains of UBL-ubiquitin-associated (UBA) proteins. In yeast, a synthetic phenotype results when specific mutations of the ubiquitin binding sites of Rpn10 and Rpn13 are combined, indicating functional linkage between these ubiquitin receptors. Because Rpn13 is also the proteasomal receptor for Uch37, a deubiquitinating enzyme, our findings suggest a coupling of chain recognition and disassembly at the proteasome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Voges, D., Zwickl, P. & Baumeister, W. The 26S proteasome: a molecular machine designed for controlled proteolysis. Annu. Rev. Biochem. 68, 1015–1068 (1999)
Deveraux, Q., Ustrell, V., Pickart, C. & Rechsteiner, M. A 26 S protease subunit that binds ubiquitin conjugates. J. Biol. Chem. 269, 7059–7061 (1994)
Elsasser, S. & Finley, D. Delivery of ubiquitinated substrates to protein-unfolding machines. Nature Cell Biol. 7, 742–749 (2005)
Madura, K. Rad23 and Rpn10: perennial wallflowers join the melee. Trends Biochem. Sci. 29, 637–640 (2004)
Elsasser, S., Chandler-Militello, D., Muller, B., Hanna, J. & Finley, D. Rad23 and Rpn10 serve as alternative ubiquitin receptors for the proteasome. J. Biol. Chem. 279, 26817–26822 (2004)
Verma, R., Oania, R., Graumann, J. & Deshaies, R. J. Multiubiquitin chain receptors define a layer of substrate selectivity in the ubiquitin-proteasome system. Cell 118, 99–110 (2004)
Kleijnen, M. F. et al. The hPLIC proteins may provide a link between the ubiquitination machinery and the proteasome. Mol. Cell 6, 409–419 (2000)
Chen, L. & Madura, K. Rad23 promotes the targeting of proteolytic substrates to the proteasome. Mol. Cell. Biol. 22, 4902–4913 (2002)
Kaplun, L. et al. The DNA damage-inducible UbL–UbA protein Ddi1 participates in Mec1-mediated degradation of Ho endonuclease. Mol. Cell. Biol. 25, 5355–5362 (2005)
Bertolaet, B. L. et al. UBA domains of DNA damage-inducible proteins interact with ubiquitin. Nature Struct. Biol. 8, 417–422 (2001)
Wilkinson, C. R. et al. Proteins containing the UBA domain are able to bind to multi-ubiquitin chains. Nature Cell Biol. 3, 939–943 (2001)
Wang, Q., Goh, A. M., Howley, P. M. & Walters, K. J. Ubiquitin recognition by the DNA repair protein hHR23a. Biochemistry 42, 13529–13535 (2003)
Hiyama, H. et al. Interaction of hHR23 with S5a. The ubiquitin-like domain of hHR23 mediates interaction with S5a subunit of 26 S proteasome. J. Biol. Chem. 274, 28019–28025 (1999)
Elsasser, S. et al. Proteasome subunit Rpn1 binds ubiquitin-like protein domains. Nature Cell Biol. 4, 725–730 (2002)
Walters, K. J., Kleijnen, M. F., Goh, A. M., Wagner, G. & Howley, P. M. Structural studies of the interaction between ubiquitin family proteins and proteasome subunit S5a. Biochemistry 41, 1767–1777 (2002)
Verma, R. et al. Proteasomal proteomics: identification of nucleotide-sensitive proteasome-interacting proteins by mass spectrometric analysis of affinity-purified proteasomes. Mol. Biol. Cell 11, 3425–3439 (2000)
Sone, T., Saeki, Y., Toh-e, A. & Yokosawa, H. Sem1p is a novel subunit of the 26 S proteasome from Saccharomyces cerevisiae . J. Biol. Chem. 279, 28807–28816 (2004)
Hamazaki, J. et al. A novel proteasome interacting protein recruits the deubiquitinating enzyme UCH37 to 26S proteasomes. EMBO J. 25, 4524–4536 (2006)
Jorgensen, J. P. et al. Adrm1, a putative cell adhesion regulating protein, is a novel proteasome-associated factor. J. Mol. Biol. 360, 1043–1052 (2006)
Qiu, X. B. et al. hRpn13/ADRM1/GP110 is a novel proteasome subunit that binds the deubiquitinating enzyme, UCH37. EMBO J. 25, 5742–5753 (2006)
Yao, T. et al. Proteasome recruitment and activation of the Uch37 deubiquitinating enzyme by Adrm1. Nature Cell Biol. 8, 994–1002 (2006)
Ito, T. et al. A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proc. Natl Acad. Sci. USA 98, 4569–4574 (2001)
Gandhi, T. K. et al. Analysis of the human protein interactome and comparison with yeast, worm and fly interaction datasets. Nature Genet. 38, 285–293 (2006)
Lam, Y. A., Xu, W., DeMartino, G. N. & Cohen, R. E. Editing of ubiquitin conjugates by an isopeptidase in the 26S proteasome. Nature 385, 737–740 (1997)
Bienko, M. et al. Ubiquitin-binding domains in Y-family polymerases regulate translesion synthesis. Science 310, 1821–1824 (2005)
Seong, K. M., Baek, J. H., Yu, M. H. & Kim, J. Rpn13p and Rpn14p are involved in the recognition of ubiquitinated Gcn4p by the 26S proteasome. FEBS Lett. 581, 2567–2573 (2007)
Schreiner, P. et al. Ubiquitin docking at the proteasome through a novel pleckstrin-homology domain interaction. Nature 10.1038/nature06924 (this issue)
Raasi, S., Orlov, I., Fleming, K. G. & Pickart, C. M. Binding of polyubiquitin chains to ubiquitin-associated (UBA) domains of HHR23A. J. Mol. Biol. 341, 1367–1379 (2004)
Glickman, M. H. et al. A subcomplex of the proteasome regulatory particle required for ubiquitin-conjugate degradation and related to the COP9-signalosome and eIF3. Cell 94, 615–623 (1998)
Johnson, E. S., Bartel, B., Seufert, W. & Varshavsky, A. Ubiquitin as a degradation signal. EMBO J. 11, 497–505 (1992)
Schmidt, M., Hanna, J., Elsasser, S. & Finley, D. Proteasome-associated proteins: regulation of a proteolytic machine. Biol. Chem. 386, 725–737 (2005)
Saeki, Y., Saitoh, A., Toh-e, A. & Yokosawa, H. Ubiquitin-like proteins and Rpn10 play cooperative roles in ubiquitin-dependent proteolysis. Biochem. Biophys. Res. Commun. 293, 986–992 (2002)
Raasi, S. & Pickart, C. M. Rad23 ubiquitin-associated domains (UBA) inhibit 26 S proteasome-catalyzed proteolysis by sequestering lysine 48-linked polyubiquitin chains. J. Biol. Chem. 278, 8951–8959 (2003)
Guterman, A. & Glickman, M. H. Complementary roles for Rpn11 and Ubp6 in deubiquitination and proteolysis by the proteasome. J. Biol. Chem. 279, 1729–1738 (2004)
Boutet, S. C., Disatnik, M. H., Chan, L. S., Iori, K. & Rando, T. A. Regulation of pax3 by proteasomal degradation of monoubiquitinated protein in skeletal muscle progenitors. Cell 130, 349–362 (2007)
Soubeyran, P., Kowanetz, K., Szymkiewicz, I., Langdon, W. Y. & Dikic, I. Cbl-CIN85-endophilin complex mediates ligand-induced downregulation of EGF receptors. Nature 416, 183–187 (2002)
Leggett, D. S. et al. Multiple associated proteins regulate proteasome structure and function. Mol. Cell 10, 495–507 (2002)
Acknowledgements
We thank members of our laboratories, as well as D. Hoeller, G. Dittmar, J. Lipscomb and M. Schmidt, for discussions, comments and reading of the manuscript. We also thank the University of Minnesota’s NMR facility, Minnesota Supercomputing Institutes’s Basic Sciences Computing Laboratory and E. Arriaga for allowing us to use his spectrofluorometer. We thank G. Zapart for the initial Y2H ubiquitin screening, and M. Groll and P. Schneider for allowing us to use the mRpn13–ubiquitin coordinates to generate Fig. 5c. This work was supported by grants from Deutsche Forschungsgemeinschaft (DI 931/3-1), the Cluster of Excellence ‘Macromolecular Complexes’ of the Goethe University Frankfurt (EXC115) (I.D.) and the National Institutes of Health (CA097004 to K.J.W.; GM043601 to D.F.; GM008700-CBITG to L.R.).
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary information
The file contains Supplementary Methods and Supplementary Figures 1-7 with Legends. (PDF 767 kb)
Rights and permissions
About this article
Cite this article
Husnjak, K., Elsasser, S., Zhang, N. et al. Proteasome subunit Rpn13 is a novel ubiquitin receptor. Nature 453, 481–488 (2008). https://doi.org/10.1038/nature06926
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06926
This article is cited by
-
1H, 15N, 13C resonance assignments for proteasome shuttle factor hHR23a
Biomolecular NMR Assignments (2023)
-
Role of Ubiquitin–Proteasome and Autophagy-Lysosome Pathways in α-Synuclein Aggregate Clearance
Molecular Neurobiology (2022)
-
Bioengineered models of Parkinson’s disease using patient-derived dopaminergic neurons exhibit distinct biological profiles in a 3D microenvironment
Cellular and Molecular Life Sciences (2022)
-
Kinectin1 depletion promotes EGFR degradation via the ubiquitin-proteosome system in cutaneous squamous cell carcinoma
Cell Death & Disease (2021)
-
The role of TXNL1 in disease: treatment strategies for cancer and diseases with oxidative stress
Molecular Biology Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.