Abstract
Papaya, a fruit crop cultivated in tropical and subtropical regions, is known for its nutritional benefits and medicinal applications. Here we report a 3× draft genome sequence of ‘SunUp’ papaya, the first commercial virus-resistant transgenic fruit tree1 to be sequenced. The papaya genome is three times the size of the Arabidopsis genome, but contains fewer genes, including significantly fewer disease-resistance gene analogues. Comparison of the five sequenced genomes suggests a minimal angiosperm gene set of 13,311. A lack of recent genome duplication, atypical of other angiosperm genomes sequenced so far2,3,4,5, may account for the smaller papaya gene number in most functional groups. Nonetheless, striking amplifications in gene number within particular functional groups suggest roles in the evolution of tree-like habit, deposition and remobilization of starch reserves, attraction of seed dispersal agents, and adaptation to tropical daylengths. Transgenesis at three locations is closely associated with chloroplast insertions into the nuclear genome, and with topoisomerase I recognition sites. Papaya offers numerous advantages as a system for fruit-tree functional genomics, and this draft genome sequence provides the foundation for revealing the basis of Carica’s distinguishing morpho-physiological, medicinal and nutritional properties.
Similar content being viewed by others
Main
Papaya is an exceptionally promising system for the exploration of tropical-tree genomes and fruit-tree genomics. It has a relatively small genome of 372 megabases (Mb)6, diploid inheritance with nine pairs of chromosomes, a well-established transformation system7, a short generation time (9–15 months), continuous flowering throughout the year and a primitive sex-chromosome system8. It is a member of the Brassicales, sharing a common ancestor with Arabidopsis about 72 million years ago9. Papaya is ranked first on nutritional scores among 38 common fruits, based on the percentage of the United States Recommended Daily Allowance for vitamin A, vitamin C, potassium, folate, niacin, thiamine, riboflavin, iron and calcium, plus fibre. Consumption of its fruit is recommended for preventing vitamin A deficiency, a cause of childhood blindness in tropical and subtropical developing countries. The fruit, stems, leaves and roots of papaya are used in a wide range of medical applications, including production of papain, a valuable proteolytic enzyme.
A total of 2.8 million whole-genome shotgun (WGS) sequencing reads were generated from a female plant of transgenic cultivar SunUp, which was developed through transformation of Sunset that had undergone more than 25 generations of inbreeding10. The estimated residual heterozygosity of SunUp is 0.06% (Supplementary Note 1). After excluding low-quality and organellar reads, 1.6 million high-quality reads were assembled into contigs containing 271 Mb and scaffolds spanning 370 Mb including embedded gaps (Supplementary Tables 1 and 2). Of 16,362 unigenes derived from expressed sequence tags (ESTs), 15,064 (92.1%) matched this assembly. Paired-end reads from 34,065 bacterial artificial chromosome (BAC) clones provided alignment to an fingerprinted contig (FPC)-based physical map (Supplementary Note 2). Among 706 BAC end and WGS sequence-derived simple sequence repeats on the genetic map, 652 (92.4%) could be used to anchor 167 Mb of contigs or 235 Mb of scaffolds, to the 12 papaya linkage groups in the current genetic map (Supplementary Fig. 1).
Papaya chromosomes at the pachytene stage of meiosis are generally stained lightly by 4′,6-diamidino-2-phenylindole (DAPI), revealing that the papaya genome is largely euchromatic. However, highly condensed heterochromatin knobs were observed on most chromosomes (Supplementary Fig. 2), concentrated in the centromeric and pericentromeric regions. The lengths of the pachytene bivalents that are heavily stained only account for approximately 17% of the genome. However, these cytologically distinct and highly condensed heterochromatic regions could represent 30–35% of the genomic DNA11. A large portion of the heterochromatic DNA was probably not covered by the WGS sequence. The 271 Mb of contig sequence should represent about 75% of the papaya genome and more than 90% of the euchromatic regions, which is similar to the 92.1% of the EST and 92.4% of genetic markers covered by the assembled genome and the theoretical 95% coverage by 3× WGS sequence12.
Gene annotation was carried out using the TIGR Eukaryotic Annotation Pipeline. The assembled genome was masked based on similarity to known repeat elements in RepBase and the TIGR Plant Repeat Database, plus a de novo papaya repeat database (see Methods). Ab initio gene predictions were combined with spliced alignments of proteins and transcripts to produce a reference gene set of 28,629 gene models (Supplementary Table 3). A total of 21,784 (76.1%) of the predicted papaya genes with average length of 1,057 base pairs (bp) have similarity to proteins in the non-redundant database from the National Center for Biotechnology Information, with 9,760 (44.8%) of these supported by papaya unigenes. Among 6,845 genes with average length 309 bp that had no hits to the non-redundant proteins, only 515 (7.5%) were supported by papaya unigenes, implying that the number of predicted papaya-specific genes was inflated. If the 515 genes with unigene support represent 44.8% of the total, then 1,150 predicted papaya-specific genes may be real, and the number of predicted genes in the assembled papaya genome would be 22,934. Considering the assembled genome covers 92.1% of the unigenes and 92.4% of the mapped genetic markers, the number of predicted genes in the papaya genome could be 7.9% higher, or 24,746, about 11–20% less than Arabidopsis (based on either the 27,873 protein coding and RNA genes, or including the 3,241 novel genes)2,13, 34% less than rice3, 46% less than poplar4 and 19% less than grape5 (Table 1).
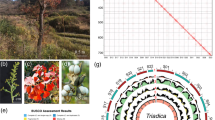
Comparison of the papaya genome with that of Arabidopsis sheds new light on angiosperm evolutionary history in several ways. Considering only the 200 longest papaya scaffolds, we found 121 co-linear blocks. The papaya blocks range in size from 1.36 Mb containing 181 genes to 0.16 Mb containing 19 genes (a statistical, rather than a biological, lower limit); the corresponding Arabidopsis regions range from 0.69 Mb containing 163 genes to 60 kilobases (kb) containing 18 genes. Across the 121 papaya segments for which co-linearity can be detected, 26 show primary correspondence (that is, excluding the effects of ancient triplication detailed below) to only one Arabidopsis segment, 41 to two, 21 to three, 30 to four, and only 3 to more than four.
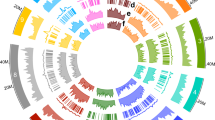
The fact that many papaya segments show co-linearity with two to four Arabidopsis segments (Fig. 1, and Supplementary Figs 3 and 4) is most parsimoniously explained if either one or two genome duplications have affected the Arabidopsis lineage since its divergence from papaya. Although it was suspected that the most recent Arabidopsis genome duplication, α14, might affect only a subset of the Brassicales15, previous phylogenetic dating of these events15 had suggested that the more ancient β-duplication occurred early in the eudicot radiation, well before the Arabidopsis–Carica divergence. This incongruity is under investigation.
‘Vv chr16r’ is an unordered ultracontig that has been assigned to grape chromosome 16. Triangles represent individual genes with transcriptional orientations. Several Arabidopsis regions belong to previously identified duplication segments (α3, α11, α20, β6, γ7, shown to the right)23. The whole syntenic alignment supports four distinct whole-genome duplication events: α, β within the Arabidopsis lineage, an independent duplication in poplar, and γ which is shared by all four eudicot genomes. Co-linear regions can be grouped into three γ sub-genomes based on Camin–Sokal parsimony criteria.
In contrast, individual Arabidopsis genome segments correspond to only one papaya segment, indicating that no genome duplication has occurred in the papaya lineage since its divergence from Arabidopsis about 72 million years ago5. The lack of relatively recent papaya genome doubling is further supported by an L-shaped distribution of intra-EST correspondence for papaya (not shown). However, multiple genome/subgenome alignments (see Supplementary Methods) reveal evidence in papaya of the ancient ‘γ’ genome duplication shared with Arabidopsis and poplar that is postulated to have occurred near the origin of angiosperms14. Indeed, both papaya (with no subsequent duplication) and poplar (with a relatively low rate of duplicate gene loss) suggest that γ was not a duplication but a triplication (Fig. 1), with triplicated patterns evident for about 25% of the 247 Mb comprising the 200 largest papaya scaffolds. This is most probably an underestimate that will increase as papaya contiguity is improved. Triplication in papaya and poplar corresponds closely to the triplication suggested by an independent analysis of the grape genome5.
A few hundred papaya chromosomal segments were aligned using BLASTZ to their one to four syntenic regions in Arabidopsis, and the results examined visually using the Genome Evolution (GEvo) viewer16. The orthologous region of grape was also included5, making the alignment a six-way comparison. One example is given in Supplementary Fig. 5: a 500 kb segment of papaya, its four 60 kb syntenic, orthologous Arabidopsis segments and the 400 kb orthologous segment of grape.
For the homologous Arabidopsis segments that are discernibly co-linear (by MC-SCANNER) to the 200 longest papaya scaffolds, 34.8% of Arabidopsis genes in any one segment correspond to a papaya gene, whereas only 24.8% of papaya genes in any one segment correspond to an Arabidopsis gene. Moreover, the Arabidopsis homologous segments contain fewer genes, on average only about 57.9% of the number in their papaya counterparts.
Papaya provides a useful outgroup necessary to detect subfunctionalization. Supplementary Fig. 6 is a GEvo screenshot of a blastn alignment illustrating subfunctionalization of conserved non-coding sequences (CNSs)17 upstream of two syntenic, duplicate Arabidopsis genes and their single papaya orthologous gene. The α-duplicated genomes within Arabidopsis are perfect for CNS discovery18.
Comparative analysis of the papaya and Arabidopsis 5′ untranslated regions showed that only 14% of orthologous promoter pairs exhibit significantly higher levels of sequence identity than random comparisons (Supplementary Figs 7 and 8). Although some highly conserved promoters show substantial conservation across much of their length, sequence similarity for most orthologous papaya promoters is indistinguishable from background.
Global analysis of all inferred protein models from papaya, Arabidopsis, poplar, grape and rice clusters the 208,901 non-redundant protein sequences into 39,706 similarity groups, or ‘tribes’19, 11,851 of which contain two or more genes (see Supplementary Methods). Tribes with multiple genes in a species typically correspond to families or subfamilies of genes; however, tribes may also contain just one gene (‘singleton tribes’). In papaya, 25,312 gene models were classified into 12,958 tribes, 5,669 of which were specific to papaya (Supplementary Table 4). Of the papaya-specific tribes, 5,314 were singleton tribes. EST support was markedly lower for genes in papaya-specific tribes (below 14%) than in tribes that included genes from at least one other taxon (72.4%).
To investigate the smaller number of genes in papaya, we compared tribe membership from each of the five sequenced angiosperm species (Supplementary Table 5). Among the 6,726 tribes that contain genes from both Arabidopsis and papaya, 3,595 contain equal numbers of genes from both species. However, tribes with more Arabidopsis genes outnumber those with more papaya genes by more than 2:1 (2,153:979). The trend of smaller number of papaya genes is widespread across tribes of all sizes and major functional categories (Supplementary Table 6 and Supplementary Fig. 9).
We then examined membership in the 815 tribes with members identified as being likely transcription factors in the Arabidopsis transcription factor database (http://arabidopsis.med.ohio-state.edu/AtTFDB/). This set includes 2,897 genes in Arabidopsis and 2,438 in papaya (a ratio of 1.19:1). The details of tribe membership are illustrated for 25 exemplar families and superfamilies (Fig. 2), where most transcription-factor tribes have fewer genes in papaya than Arabidopsis. Some transcription-factor tribes had more genes in papaya, specifically RWP-RK, MADS-box, Scarecrow, TCP and Jumonji gene families. Interestingly, the difference in MADS protein family size appears to be due to expanded numbers for half of the 36 MADS tribes. The other 18 MADS tribes had fewer papaya genes, including 14 that were not found in papaya.
Most transcription factors are represented by fewer genes in papaya than Arabidopsis. Transcription-factor names are given, with values after the names corresponding to: number of tribes with genes assigned to transcription factor group, number of tribes with smaller counts in papaya than Arabidopsis, number of tribes with equal counts in papaya and Arabidopsis, number of tribes with larger counts in papaya, and number of tribes with zero members in papaya. Supporting data are provided in Supplementary Table 8.
Assuming that a generalized angiosperm could potentially require only the types and minimal numbers of genes that are shared among divergent plant species, we examined each of the tribes shared among the five angiosperms with sequenced genomes. The number of genes required in a minimal flowering plant is based on the observed minimum number of genes across each of the shared tribes (Table 2). When the smallest observed number is taken for each evolutionarily conserved tribe, a minimal angiosperm genome of 13,311 genes is estimated. Papaya has the smallest number of genes for more tribes than any other sequenced taxon (4,515, or 76% of 5,925 shared tribes), reinforcing the notion that papaya has fewer genes than any angiosperm sequenced so far.
Only 55 nucleotide-binding site (NBS)-containing R genes were identified in papaya; about 28% of the 200 NBS genes in Arabidopsis20 and less than 10% of the 600 NBS genes in rice21. Resistance proteins also have a carboxy-terminal leucine-rich repeat (LRR) domain. These NBS-containing R-gene families can be subdivided into three classes: NBS–LRR, toll interleukin receptor (TIR)–NBS–LRR, and coiled-coil (CC)–NBS–LRR on the basis of their amino-terminal region. Papaya NBS–LRR outnumbered both TIR–NBS–LRR and CC–NBS–LRR genes, in contrast to both poplar (with more CC–NBS–LRR genes4) and Arabidopsis (with more TIR–NBS–LRR). More than 50% of the NBS-type R genes were clustered in about eight scaffolds, indicating that resistance gene evolution may involve duplication and divergence of linked gene families.
Homologues for genes involved in cellulose biosynthesis are present in papaya and Arabidopsis, with more cellulose synthase genes in poplar, perhaps associated with wood formation. Papaya has at least 32 putative β-glucosyl transferase (GT1) genes compared with 121 in Arabidopsis identified using sequence alignment. A total of 38 and 40 cellulose synthase-related genes (GT2) were identified in papaya using the 48 poplar and 31 Arabidopsis genes as queries, respectively. These genes include 11 cellulose synthase (CesA) genes, the same number as in Arabidopsis but 7 fewer than in poplar. Putative cellulose orientation genes (COBRA) were more abundant in Arabidopsis (12) than in papaya (8).
Papaya also has a similar complement though fewer genes for cell-wall synthesis than Arabidopsis. Papaya and Arabidopsis, respectively, have 6 and 12 callose synthase genes (GT2); 15 and 15 xyloglucan α-1,2-fucosyl transferases (GT37); 5 and 7 β-glucuronic acid transferases in familes GT43 and GT47; and 27 and 42 in GT8 that includes galacturonosyl transferases, associated with pectin synthesis.
The cell wall of plants is capable of both plastic and elastic extension, and controls the rate and direction of cell expansion22. Despite fewer whole-genome duplications, papaya has a similar number of putative expansin A genes (24) as Arabidopsis (26) and poplar (27), and more expansin B genes (10) than Arabidopsis (6) and poplar (3).
In contrast to expansion-related genes, papaya has on average about 25% fewer cell-wall degradation genes than Arabidopsis, in some cases far fewer. For example, papaya and Arabidopsis, respectively, have 4 and 12 endoxylanase-like genes in glycoside hydrolase family 10 (GH10); 29 and 67 pectin methyl esterases (carbohydrate esterase family 8); 28 and 69 polygalacturonases (GH28); 15 and 49 xyloglucan endotransglycosylase/hydrolases (GH16); 18 and 25 β-1,4-endoglucanases (GH9); 42 and 91 β-1,3-glucanases (GH17); and 15 and 27 pectin lyases (PL1).
A semi-woody giant herb that accumulates lignin in the cell wall at an intermediate level between Arabidopsis and poplar, papaya generally has intermediate numbers of lignin synthetic genes, fewer than poplar but more than Arabidopsis despite fewer opportunities for duplication in papaya. Poplar, papaya and Arabidopsis have 37, 30 and 18 candidate genes for the lignin synthesis pathway, respectively4,23, with papaya having an intermediate number of genes for the PAL, C4H, 4CL and HCT gene families, and only one COMT and two C3H genes. In contrast, poplar has three C3H genes, which are presumed to convert p-coumaroyl quinic acid to caffeoyl shikimic acid, whereas there are two in papaya and one in Arabidopsis. Papaya, Arabidopsis and poplar each have two genes in the family CCoAOMT, which are presumed to convert caffeic acid to ferulic acid4. Compared with these other plants, papaya has the fewest genes in the CCR gene family (1 gene) and the most in the F5H (4 genes) and CAD gene families (18 genes), which all mediate later steps of the lignin biosynthesis pathway.
More starch-associated genes in papaya, a perennial, may be due to a greater need for storage in leaves, stem and developing fruit than in Arabidopsis, an ephemeral that stores oil in the seed. Papaya and Arabidopsis, respectively, have 13 and 6 putative starch synthase (GT5) genes; 8 and 3 starch branching genes; 6 and 3 isoamylases (GH13); and 12 and 9 β-amylases (GH14). Early unloading of fruit sugar in papaya is probably symplastic24, with five genes for sucrose synthase/sucrose phosphate synthase (GT4); seven are reported for Arabidopsis. Five acid invertase (GH32) sequences were found in papaya whereas 11 have been reported in Arabidopsis. Papaya has at least seven putative neutral invertase (GH32) genes; Arabidopsis has six. Wall-associated kinases (WAK) are thought to be involved in the regulation of vacuolar invertases, with 17 in Arabidopsis and only 10 in papaya. Arabidopsis and papaya have 14 and 7 hexose transporters, respectively. The greater number of genes for sugar accumulation in Arabidopsis may reflect recent genome duplications.
Papaya has undergone particularly striking amplification of genes involved in volatile development. Papaya and Arabidopsis, respectively, have 18 and 8 genes for cinnamyl alcohol dehydrogenase; 2 and 1 genes for cinnamate-4-hydroxylase; 9 and 3 genes for phenylalanine ammonia lyase; and 24 and 3 limonene cyclase genes.
Papaya ripening is climacteric, with the rise in ethylene production occurring at the same time as the respiratory increase25. Papaya and Arabidopsis, respectively, have similar numbers of genes involved in ethylene synthesis, with four each for S-adenosyl methionine synthase (SAM synthase); 8 and 13 for aminocyclopropane carboxylic acid (ACC) synthase (ACS); 8 and 12 for ACC oxidase (ACO); and 42 and 64 for ethylene-responsive binding factors (AP2/ERF).
Because papaya grows in tropical climates where daily light/dark cycles do not change much over the year, we can ask if more or fewer light/circadian genes are required to synchronize with the environment. In fact, there are fewer light/clock genes in the papaya genome (49% and 34% of poplar and Arabidopsis, respectively; Supplementary Table 7). However, among the core circadian clock genes, the pseudo-response regulators (PRRs; Supplementary Fig. 10) have expanded in poplar compared with Arabidopsis, and the papaya PRR7 cluster has seemingly duplicated with the recent poplar salicoid-specific genome duplication4 (Supplementary Fig. 11). Against the backdrop of fewer overall genes, the parallel expansion of the PRRs is consistent with circadian timing being important in papaya.
The PAS–FBOX–KELCH genes control light signalling and flowering time; however, the only papaya orthologue (ZTL) lacks an obvious KELCH domain compared with Arabidopsis and poplar, which have five and one KELCH domains, respectively (Supplementary Fig. 10). In fact, the papaya genome contains fewer KELCH domains (37 compared with 130 and 74 in Arabidopsis and poplar, respectively). In contrast, there are three constitutive photomorphogenic 1 (COP1) paralogues in the papaya genome compared with only one in Arabidopsis (Supplementary Tables 7 and 8). A similar expansion has been noted in moss (Physcomitrella patens), which has nine COP1 paralogues that are hypothesized to aid in tolerance to ultraviolet light (Supplementary Fig. 12)26. Both KELCH domains and the WD-40 of the COP1 family form β-propellers and play a role in light-mediated ubiquitination. There is not a general expansion of WD-40 genes in papaya (173 compared with 227 in Arabidopsis). Perhaps papaya has developed an alternative way of integrating light or timing information specific to day-neutral plants, such as a strict adherence to the diel light/dark cycle that is better served by the COP-mediated system.
Sex determination in papaya is controlled by a pair of primitive sex chromosomes, with a small male-specific region of the Y chromosome (MSY)8. The physical map of the MSY is currently estimated by chromosome walking to span about 8 Mb (ref. 27). Two scaffolds in the current female-genome sequence align to the X chromosome physical map based on BAC end sequences, spanning 4.5 Mb and including 254 predicted protein-encoding genes, of which 75 (29.5%) have EST support (Supplementary Table 9 and Supplementary Fig. 13). If adjusted for the percentage of unigene validation for other genes (48.0%), the estimated number of genes in the X-specific region would be 156. The average gene density would be one gene per 19.5 kb, lower than the estimated genome average of one gene per 14.3 kb. By contrast, among seven completely sequenced MSY BACs totalling 1.2 Mb, a total of four expressed genes were found on two of the BACs14,28. The somewhat lower-than-average gene density in the X-specific scaffolds is accompanied by more repetitive DNA (58.3%) than the genome-wide average, perhaps because this region is near the centromere28. Re-analysis of the repetitive DNA content of the MSY BACs, to include the new papaya-specific repeat families identified herein, increased the average repeat sequence to 85.6%, with 54.1% Gypsy and 1.9% Copia retro-elements (Supplementary Table 10). This compares with an earlier estimate of 17.9% using the Arabidopsis repeat database alone28.
The SunUp genome has presented an opportunity to analyse transgene insertion sites critically. Southern blot analysis was key in the initial identification of transgenic insertion fragments and was performed with probes spanning the entire 19,567-bp transformation vector used for bombardment (Supplementary Fig. 14). Among the identified inserts were the functional coat-protein transgene conferring resistance to papaya ringspot virus, which was found in an intact 9,789-bp fragment of the transformation plasmid, and a 1,533-bp fragment composed of a truncated, non-functional tetA gene and flanking vector backbone sequence. The structures of the coat-protein transgene and tetA region insertion sites were determined from cloned sequences. Southern analysis also confirmed a 290-bp non-functional fragment of the nptII gene originally identified by WGS sequence analysis (Supplementary Fig. 15). Five of the six flanking sequences of the three insertions are nuclear DNA copies of papaya chloroplast DNA fragments. The integration of the transgenes into chloroplast DNA-like sequences may be related to the observation that transgenes produced either by Agrobacterium-mediated or biolistic transformation are often inserted in AT-rich DNA29, as is the chloroplast DNA of papaya and other land plants. Four of the six insert junctions have sequences that match topoisomerase I recognition sites, which are associated with breakpoints in genomic DNA transgene insertion sites and transgene rearrangements29. The presence of these inserts was confirmed by high-throughput MUMmer30 analysis for each region of the transformation vector. Evidence for the presence of other transgene inserts is not conclusive (Supplementary Note 3).
Its lower overall gene number notwithstanding, striking variations in gene number within particular functional groups, superimposed on the average approximate 20% reduction in papaya gene number relative to Arabidopsis, may be related to key features of papaya morphological evolution. Despite a closer evolutionary relationship to Arabidopsis, papaya shares with poplar an increased number of genes associated with cell expansion, consistent with larger plant size; and lignin biosynthesis, consistent with the convergent evolution of tree-like habit. Amplification of starch-synthesis genes in papaya relative to Arabidopsis is consistent with a greater need for storage in leaves, stem and developing fruit of this perennial. Tremendous amplification in papaya of genes related to volatile development implies strong natural selection for enhanced attractants that may be key to fruit (seed) dispersal by animals and which may also have attracted the attention of aboriginal peoples. This also foreshadows what we might expect to discover in the genomes of other fragrant-fruited trees, as well as plants with striking fragrance of leaves (herbs), flowers or other organs.
Arguably, the sequencing of the genome of SunUp papaya makes it the best-characterized commercial transgenic crop. Because papaya ringspot virus is widespread in nearly all papaya-growing regions, SunUp could serve as a transgenic germplasm source that could be used to breed suitable cultivars resistant to the virus in various parts of the world. The characterization of the precise transgenic modifications in SunUp papaya should also serve to lower regulatory barriers currently in place in some countries.
Methods Summary
Gene annotation
Papaya unigenes from complementary DNA were aligned to the unmasked genome assembly, which was then used in training ab initio gene prediction software. Spliced alignments of proteins from the plant division of GenBank, and transcripts from related angiosperms, were generated. Gene predictions were combined with spliced alignments of proteins and transcripts to produce a reference gene set. Detailed descriptions are given in Methods.
Online Methods
Genome assembly
The Genome sequence was assembled by Arachne31. WGS reads and BAC end reads were trimmed by LUCY and screened for organellar sequences32. Two approaches were applied to screening and removing reads of presumably organellar origin to alleviate the load in assembling highly repetitive regions by WGS assembly software. The first approach was an iterative process, in which reads were assembled, contigs matching with organellar genomes identified, constituent reads removed, and the process repeated by two or three more rounds. This approach produced the read sets for the released assemblies Stripped3 and Stripped4. The second approach was to remove plasmid clones and BAC clones of presumably organellar origin by identifying clones with both end reads matching entirely with organellar genomes, with physical map information an amendment to the identification of BAC clones. Two rounds of iterative screening based on pairing information of assembled and unplaced reads were added to the second approach to generate the read set for the released Papaya1.0 assembly.
The sequence error rates were estimated by aligning assembled shotgun sequences with two finished BACs (GenBank accession numbers EF661023 and EF661026). The error rate of the assembly at 3× coverage or deeper (74.2% of assembled sequences) was less than 0.01% based on average quality values of 20 or greater in trimmed sequence. The error rate at 2× coverage (16.3%) was 0.37%. The error rate at 1× coverage (9.5%) was approximately 0.75%, because these sequences are at the ends of the contigs (and sequence reads) where the sequence quality declined.
Genome annotation
Gene annotation was conducted following the TIGR Eukaryotic Annotation Pipeline. Repeat sequences were identified in the assembled genome and masked by RepeatMasker, RepeatScout and TransposonPSI, based on known repeat elements in RepBase databases and TIGR Plant Repeat Databases, and the papaya novel repeat database constructed in this study33,34. Program to Assemble Spliced Alignments (PASA)35 was used to generate spliced alignments of papaya unigenes to the unmasked assembly, which was then used in training ab initio gene prediction software Augustus, GlimmerHMM and SNAP36,37,38. Ab initio gene prediction software Fgenesh, Genscan and TWINSCAN were trained on Arabidopsis39,40,41. Spliced alignments of proteins from the plant division of GenBank and transcripts from related angiosperms (Arabidopsis thaliana, Glycine max, Gossypium hirsutum, Medicago truncatula, Nicotiana tabacum, Oryza sativa, Zea mays) were generated by the Analysis and Annotation Tool (AAT)42. Spliced alignment of proteins from the Pfam database were generated using GeneWise43,44. Gene predictions generated by Augustus, Fgenesh, Genscan, GlimmerHMM, SNAP and TWINSCAN were combined with spliced alignments of proteins and transcripts to produce a reference gene set using the evidence-based combiner EVidenceModeler (EVM)45. Protein domains were predicted using InterProScan against protein databases (PRINTS, Pfam, ProDom, PROSITE, SMART)46,47,48,49,50.
Construction of papaya repeat database
We used a combination of homology-based and de novo methods to identify signatures of transposable elements in the papaya genome. We used RepeatMasker (http://www.repeatmasker.org) in combination with a custom-built library of plant repeat elements for our initial classification of transposable elements. The customized library was generated by combining plant repeats from Repbase and plant repeat databases from TIGR (ftp://ftp.tigr.org/pub/data/TIGR_Plant_Repeats)33. Repeat elements identified as ribosomal RNA sequences in the TIGR databases match a large fraction of the papaya genome (about 3%). Ribosomal RNAs were identified separately, and therefore were excluded from our repeat library, leaving a database of 76,924 repeat sequences that were used to search the papaya genome.
Homology-based methods are limited to finding elements that have not diverged too greatly from known repeats. Because databases of known transposable elements are necessarily incomplete, we used additional de novo methods to search for repeat elements in papaya contigs. For this, we applied two recently developed repeat-finding tools, PILER and RepeatScout to the complete set of contigs from the papaya genome34,51. PILER was able to find 428 repeat families whereas RepeatScout found 6,596 repeat sequences.
The repeat families obtained from PILER and RepeatScout were annotated using a combination of manual curation (786 repeat families) and automated analysis. For the automated annotation, the combined data set from PILER and RepeatScout was made non-redundant (using CD-HIT at the 90% similarity level), leaving behind 6,240 repeat families52. As a post-processing step, we selected only those families that had at least ten good (E value < 1 × 1020) BLAST matches to papaya contigs. The resulting data set contained 2,198 repeat families in the papaya genome. BLAST searches against non-redundant and PTREP (http://wheat.pw.usda.gov/ITMI/Repeats) were then used to identify repeat families matching genes associated with transposons and retrotransposons. This procedure discovered an additional 103 repeat families that could be annotated as being retrotransposons. The combined database of 889 annotated papaya-specific transposable-element sequences was used in addition to the database of known repeats to annotate the papaya genome. The remaining, unannotated repeat families (1,455 sequences with no matches to known genes) were then used to estimate the additional repeat content of the genome.
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Data deposits
The papaya WGS sequence is deposited at DNA Data Bank of Japan/European Molecular Biology Laboratory/GenBank under accession number ABIM00000000. The version described in this paper is the first version, ABIM01000000. The GenBank accession numbers of the papaya ESTs are EX227656–EX303501. This paper is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence, and is freely available to all readers at www.nature.com/nature.
References
Gonsalves, D. Control of papaya ringspot virus in papaya: a case study. Annu. Rev. Phytopathol. 36, 415–437 (1998)
The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana . Nature 408, 796–815 (2000)
International Rice Genome Sequencing Project. The map-based sequence of the rice genome. Nature 436, 793–800 (2005)
Tuskan, G. A. et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313, 1596–1604 (2006)
Jaillon, C. O. et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449, 463–467 (2007)
Arumuganathan, K. & Earle, E. D. Nuclear DNA content of some important plant species. Plant Mol. Biol. Rep. 9, 208–218 (1991)
Fitch, M. M. M., Manshardt, R. M., Gonsalves, D., Slightom, J. L. & Sanford, J. C. Virus resistant papaya plants derived from tissues bombarded with the coat protein gene of papaya ringspot virus. Bio/technology 10, 1466–1472 (1992)
Liu, Z. et al. A primitive Y chromosome in papaya marks incipient sex chromosome evolution. Nature 427, 348–352 (2004)
Wikström, N., Savolainen, V. & Chase, M. W. Evolution of the angiosperms: calibrating the family tree. Proc. R. Soc. Lond. B 268, 2211–2220 (2001)
Storey, W. B. Papaya. in Outlines of Perennial Crop Breeding in the Tropics (eds Ferwerda, F. P. and Wit, F.) 389–408 (H. Veenman & Zonen, Wageningen, 1969)
Li, L. et al. Genome-wide transcription analyses in rice using tiling microarrays. Nature Genet. 38, 124–129 (2006)
Lander, E. S. & Waterman, M. S. Genomic mapping by fingerprinting random clones: a mathematical analysis. Genomics 2, 231–239 (1988)
Hanada, K., Zhang, X., Borevitz, J. O., Li, W.-H. & Shiu, S.-H. A large number of novel coding small open reading frames in the intergenic regions of the Arabidopsis thaliana genome are transcribed and/or under purifying selection. Genome Res. 17, 632–640 (2007)
Bowers, J. E., Chapman, B. A., Rong, J. & Paterson, A. H. Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature 422, 433–438 (2003)
Schranz, M. E. & Mitchell-Olds, T. Independent ancient polyploidy events in the sister families Brassicaceae and Cleomaceae. Plant Cell 18, 1152–1165 (2006)
Lyons, E. & Freeling, M. How to usefully compare homologous plant genes and chromosomes as DNA sequence. Plant J. 53, 661–673 (2008)
Inada, D. C. et al. Conserved noncoding sequences in the grasses. Genome Res. 13, 2030–2041 (2003)
Thomas, B. C., Rapaka, L., Lyons, E., Pedersen, B. & Freeling, M. Arabidopsis intragenomic conserved noncoding sequence. Proc. Natl Acad. Sci. USA 104, 3348–3353 (2007)
Wall, P. K. et al. PlantTribes: a gene and gene family resource for comparative genomics in plants. Nucleic Acids Res. 36, D970–D976 (2008)
Meyers, B. C., Morgante, M. & Michelmore, R. W. TIR-X and TIR-NBS proteins: two new families related to disease resistance TIR-NBS-LRR proteins encoded in Arabidopsis and other plant genomes. Plant J. 32, 77–92 (2002)
Zhou, T. et al. Genome-wide identification of NBS genes in japonica rice reveals significant expansion of divergent non-TIR NBS-LRR genes. Mol. Genet. Genomics 271, 402–415 (2004)
Fry, S. C. Primary cell wall metabolism: tracking the careers of wall polymers in living plant cells. New Phytol. 161, 641–675 (2004)
Ehlting, J. et al. Global transcript profiling of primary stems from Arabidopsis thaliana identifies candidate genes for missing links in lignin biosynthesis and transcriptional regulators of fiber differentiation. Plant J. 42, 618–640 (2005)
Zhou, L. L. & Paull, R. E. Sucrose metabolism during papaya (Carica papaya) fruit growth and ripening. J. Am. Soc. Hortic. Sci. 126, 351–357 (2001)
Paull, R. E. & Chen, N. J. Postharvest variation in cell wall-degrading enzymes of papaya (Carica papaya L.) during fruit ripening. Plant Physiol. 72, 382–385 (1983)
Richardt, S., Lang, D., Reski, R., Frank, W. & Rensing, S. A. PlanTAPDB, a phylogeny-based resource of plant transcription-associated proteins. Plant Physiol. 143, 1452–1466 (2007)
Yu, Q. et al. Low X/Y divergence of four pairs of papaya sex-liked genes. Plant J. 53, 124–132 (2008)
Yu, Q. et al. Chromosomal location and gene paucity of the male specific region on papaya Y chromosome. Mol. Genet. Genomics 278, 177–185 (2007)
Sawasaki, T., Takahashi, M., Goshima, N. & Morikawa, H. Structures of transgene loci in transgenic Arabidopsis plants obtained by particle bombardment: junction regions can bind to nuclear matrices. Gene 218, 27–35 (1998)
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004)
Jaffe, D. B. et al. Whole-genome sequence assembly for mammalian genomes: Arachne 2. Genome Res. 13, 91–96 (2003)
Chou, H. H. & Holmes, M. H. DNA sequence quality trimming and vector removal. Bioinformatics 17, 1093–1104 (2001)
Smit, A. F. A., Hubley, R. & Green, P. RepeatMasker (Release Open-3.1.3, 2006)
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21 (suppl.). i351–i358 (2005)
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666 (2003)
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19 (suppl.). ii215–ii225 (2003)
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 20, 2878–2879 (2004)
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004)
Salamov, A. A. & Solovyev, V. V. Ab initio gene finding in Drosophila genomic DNA. Genome Res. 10, 516–522 (2000)
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94 (1997)
Korf, I., Flicek, P., Duan, D. & Brent, M. R. Integrating genomic homology into gene structure prediction. Bioinformatics 17 (suppl. 1). S140–S148 (2001)
Huang, X., Adams, M. D., Zhou, H. & Kerlavage, A. R. A tool for analyzing and annotating genomic sequences. Genomics 46, 37–45 (1997)
Finn, R. D. et al. Pfam: clans, web tools and services. Nucleic Acids Res. 34 (Database issue). D247–D251 (2006)
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004)
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, R7.1–R7.19 (2008)
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Res. 33, W116–W120 (2005)
Attwood, T. K. et al. PRINTS and its automatic supplement, prePRINTs. Nucleic Acids Res. 31, 400–402 (2003)
Bru, C. et al. The ProDom database of protein domain families: more emphasis on 3D. Nucleic Acids Res. 33 (Database issue). D212–D215 (2005)
Hulo, N. et al. The PROSITE database. Nucleic Acids Res. 34 (Database issue). D227–D230 (2006)
Letunic, I. et al. SMART 5: domains in the context of genomes and networks. Nucleic Acids Res. 34 (Database issue). D257–D260 (2006)
Edgar, R. C. & Myers, E. W. PILER: Identification and classification of genomic repeats. Bioinformatics 21 (suppl.). i152–i158 (2005)
Li, W. & Godzik, A. CD-HIT: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, i1658–i1659 (2006)
Acknowledgements
We thank X. Wan, J. Saito and A. Young at the University of Hawaii for technical assistance; C. Detter at the DOE Joint Genome Institute; F. MacKenzie, O. Veatch and T. Uhm at the Hawaii Agriculture Research Center; L. Li, W. Teng, Y. Wu, Y. Yang, C. Zhou, N. Wang, P. Wang and D. Fei at the Tianjin Biochip Corporation, Tianjin Economic-Technological Development Area, Tianjin; and R. Herdes, L. Diebold, R. Kim, A. Hernandez, S. Ali and L. Bynum at the University of Illinois at Urbana-Champaign. This papaya genome-sequencing project was given support by the University of Hawaii and the US Department of Defense grant number W81XWH0520013 to M.A., the Maui High Performance Computing Center to M.A., the Hawaii Agriculture Research Center to R.M. and Q.Y., and Nankai University, China, to L.W. Other support to the papaya genome project included the United States Department of Agriculture T-STAR program; a United States Department of Agriculture–Agricultural Research Service cooperative agreement (CA 58-3020-8-134) with the Hawaii Agriculture Research Center; the University of Illinois; the National Science Foundation Plant Genome Research Program; and Tianjin Municipal Special Fund for Science and Technology Innovation Grant 05FZZDSH00800. We thank P. Englert, former chancellor of the University of Hawaii, for initial infrastructure support of the research.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary information
The file contains Supplementary Notes, Supplementary Methods, Supplementary Tables 1-12 and Supplementary Figures 1-17 with Legends. (PDF 1040 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Ming, R., Hou, S., Feng, Y. et al. The draft genome of the transgenic tropical fruit tree papaya (Carica papaya Linnaeus). Nature 452, 991–996 (2008). https://doi.org/10.1038/nature06856
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06856
This article is cited by
-
Carica papaya L. sex chromosome review and physical mapping of the serk 2, svp-like and mdar 4 sequences
Scientific Reports (2024)
-
Genetics and evolutionary insights from Solanaceae genome sequences
Plant Systematics and Evolution (2024)
-
Pan-genome of Citrullus genus highlights the extent of presence/absence variation during domestication and selection
BMC Genomics (2023)
-
Development of a Mesophyll Protoplast-Based System for Gene Editing of Papaya
In Vitro Cellular & Developmental Biology - Plant (2023)
-
Developing a robust in vivo hairy root system for assessing transgene expression and genome editing efficiency in papaya
Plant Cell, Tissue and Organ Culture (PCTOC) (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.