Abstract
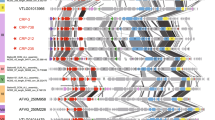
Viruses, and more particularly phages (viruses that infect bacteria), represent one of the most abundant living entities in aquatic and terrestrial environments. The biogeography of phages has only recently been investigated and so far reveals a cosmopolitan distribution of phage genetic material (or genotypes)1,2,3,4. Here we address this cosmopolitan distribution through the analysis of phage communities in modern microbialites, the living representatives of one of the most ancient life forms on Earth. On the basis of a comparative metagenomic analysis of viral communities associated with marine (Highborne Cay, Bahamas) and freshwater (Pozas Azules II and Rio Mesquites, Mexico) microbialites, we show that some phage genotypes are geographically restricted. The high percentage of unknown sequences recovered from the three metagenomes (>97%), the low percentage similarities with sequences from other environmental viral (n = 42) and microbial (n = 36) metagenomes, and the absence of viral genotypes shared among microbialites indicate that viruses are genetically unique in these environments. Identifiable sequences in the Highborne Cay metagenome were dominated by single-stranded DNA microphages that were not detected in any other samples examined, including sea water, fresh water, sediment, terrestrial, extreme, metazoan-associated and marine microbial mats. Finally, a marine signature was present in the phage community of the Pozas Azules II microbialites, even though this environment has not been in contact with the ocean for tens of millions of years. Taken together, these results prove that viruses in modern microbialites display biogeographical variability and suggest that they may be derived from an ancient community.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Data deposits
The microbialite viral metagenomes have been deposited into the ftp server of the SEED public database ftp://ftp.theseed.org/metagenomes under the project accession numbers 4440323.3(Highborne Cay), 4440320.3 (Pozas Azules II) and 4440321.3 (Rio Mesquites). The metagenomes are also publicly accessible in the CAMERA metagenomic database (http://camera.calit2.net) under the project accession numbers HBCStromBahamasVir011105 (Highborne Cay), PAStromCCMexVir072205 (Pozas Azules II), and RMStromCCMexVir072205 (Rio Mesquites). The Vp1 cloned sequences from the Highborne Cay sample have been deposited in GenBank under accession numbers EF679227 to EF679234.
References
Angly, F. E. et al. The marine viromes of four oceanic regions. PLoS Biol. 4, e368 (2006)
Casas, V. et al. Widespread occurrence of phage-encoded exotoxin genes in terrestrial and aquatic environments in Southern California. FEMS Microbiol. Lett. 261, 141–149 (2006)
Breitbart, M., Miyake, J. H. & Rohwer, F. Global distribution of nearly identical phage-encoded DNA sequences. FEMS Microbiol. Lett. 236, 249–256 (2004)
Short, C. M. & Suttle, C. A. Nearly identical bacteriophage structural gene sequences are widely distributed in both marine and freshwater environments. Appl. Environ. Microbiol. 71, 480–486 (2005)
Walter, M. R. Stromatolites. (Elsevier, Amsterdam, 1976)
Allwood, A. C., Walter, M. R., Kamber, B. S., Marshall, C. P. & Burch, I. W. Stromatolite reef from the Early Archaean era of Australia. Nature 441, 714–718 (2006)
Schopf, J. W. Fossil evidence of Archaean life. Phil. Trans. R. Soc. Lond. B 361, 869–885 (2006)
Suttle, C. A. Viruses in the sea. Nature 437, 356–361 (2005)
Cho, J. C. & Tiedje, J. M. Biogeography and degree of endemicity of fluorescent Pseudomonas strains in soil. Appl. Environ. Microbiol. 66, 5448–5456 (2000)
Papke, R. T., Ramsing, N. B., Bateson, M. M. & Ward, D. M. Geographical isolation in hot spring cyanobacteria. Environ. Microbiol. 5, 650–659 (2003)
Whitaker, R. J., Grogan, D. W. & Taylor, J. W. Geographic barriers isolate endemic populations of hyperthermophilic Archaea. Science 301, 976–978 (2003)
Whitaker, R. J. Allopatric origins of microbial species. Phil. Trans. R. Soc. Lond. B 361, 1975–1984 (2006)
Sano, E., Carlson, S., Wegley, L. & Rohwer, F. Movement of viruses between biomes. Appl. Environ. Microbiol. 70, 5842–5846 (2004)
Breitbart, M. & Rohwer, F. Here a virus, there a virus, everywhere the same virus? Trends Microbiol. 13, 278–284 (2005)
Breitbart, M. et al. Genomic analysis of uncultured marine viral communities. Proc. Natl Acad. Sci. USA 99, 14250–14255 (2002)
Rohwer, F. & Edwards, R. A. The phage proteomic tree: a genome-based taxonomy for phage. J. Bacteriol. 184, 4529–4535 (2002)
Fane, B. Microviridae, in Virus Taxonomy: Eighth Report of the International Committee on Taxonomy of Viruses. (eds Fauquet, M. A. M. C., Maniloff, J., Desselberger, U. & Ball, L. A.) 289–299 (Elsevier Academic Press, San Diego, California, 2005)
Souza, V. et al. An endangered oasis of aquatic microbial biodiversity in the Chihuahuan desert. Proc. Natl Acad. Sci. USA 103, 6565–6570 (2006)
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular Cloning: A Laboratory Manual. (Cold Spring Harbor Laboratory Press, New York, 1989)
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005)
Lozupone, C., Hamady, M. & Knight, R. UniFrac - An online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinformatics 7, 371 (2006)
Jensen, J., Bohonak, A. & Kelley, S. Isolation by distance, web service. BMC Genet. 6, 13 (2005)
Reid, R. P., Macintyre, I. G. & Steneck, R. S. A microbialite/algal ridge fringing reef complex, Highborne Cay, Bahamas. Atoll Res. Bull. 465, 1–18 (1999)
Chen, F., Lu, J., Binder, B. J., Liu, Y. & Hodson, R. E. Application of digital image analysis and flow cytometry to enumerate marine viruses stained with SYBR Gold. Appl. Environ. Microbiol. 67, 539–545 (2001)
Altschul, S. F., Gish, W., Miller, W., Meyers, E. W. & Lipman, D. J. Basic Local Alignment Search Tool. J. Mol. Biol. 215, 403–410 (1990)
Angly, F. et al. PHACCS, an online tool for estimating the structure and diversity of uncultured viral communities using metagenomic information. BMC Bioinformatics 6, 41 (2005)
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994)
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001)
Saito, N. & Nei, M. The neighbour-joining method, a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 79, 426–434 (1987)
Perriere, G. & Gouy, M. WWW-Query: An on-line retrieval system for biological sequence banks. Biochimie 78, 364–369 (1996)
Acknowledgements
Logistical field support was provided by the crew of the RV Walton Smith, Highborne Cay management and personnel of the Area de Proteccion de Flora y Fauna of Cuatro Ciénegas. This work was supported by an NSF grant to F.R. Support for B.K.S. and D.L.V. was provided by the NSF. M.B. was supported by a grant from the University of South Florida's Internal New Research Awards Program. V.S. was funded by the CONACYT 2002-C01-0237 project. The authors thank P. Visscher, K. Przekop, L. Rothschild, D. Rogoff, V. Michotey, P. Bonin, S. Norman and E. Bowlin for providing samples of marine microbial mats and M. Schaechter for a critical reading of the manuscript.
Author Contributions C.D. and F.R. designed the project. C.D. analysed most of the bioinformatic results, conducted the molecular biology and wrote the article. S.K. performed the bayesian analysis. S.R. implemented the cross-contig analyses. M.H. extracted viral DNAs. B.R.-B., H.L., F.E.A. and R.A.E. performed bioinformatic analyses. R.V.T. and D.H. helped with the interpretation of the bioinformatic results. V.S., M.B., J.S. and R.P.R. collected the samples. B.K.S., D.L.V., M.F., T.T., L.L., Y.R., L.W. and B.C. provided metagenomic data. F.R. supervised the project and helped with the writing. All authors edited and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-8 and Supplementary Tables 1-5 with Legends (Part 1); the viral capsid sequences reconstructed from the metagenomes (Part 2); and the sequences obtained from the cloning experiment with accession numbers (Part 3). (PDF 1315 kb)
Supplementary Video 1
The file contains Supplementary Video 1 describing the microbialites. (MOV 5481 kb)
Supplementary Video 2
The file contains Supplementary Video 2. The movie provides information on the sampling site in the Cuatro Ciénegas Basin (Mexico). (MOV 5854 kb)
Rights and permissions
About this article
Cite this article
Desnues, C., Rodriguez-Brito, B., Rayhawk, S. et al. Biodiversity and biogeography of phages in modern stromatolites and thrombolites. Nature 452, 340–343 (2008). https://doi.org/10.1038/nature06735
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06735
This article is cited by
-
Viruses participate in the organomineralization of travertines
Scientific Reports (2023)
-
Bacteriophages diversity in India’s major river Ganga: a repository to regulate pathogenic bacteria in the aquatic environment
Environmental Science and Pollution Research (2022)
-
Ethnobotanical biocultural diversity by rural communities in the Cuatrociénegas Valley, Coahuila; Mexico
Journal of Ethnobiology and Ethnomedicine (2021)
-
Diverse and unique viruses discovered in the surface water of the East China Sea
BMC Genomics (2020)
-
Argonaute bypasses cellular obstacles without hindrance during target search
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.