Abstract
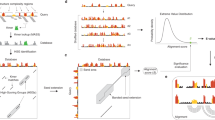
The classical RNA secondary structure model considers A·U and G·C Watson–Crick as well as G·U wobble base pairs. Here we substitute it for a new one, in which sets of nucleotide cyclic motifs define RNA structures. This model allows us to unify all base pairing energetic contributions in an effective scoring function to tackle the problem of RNA folding. We show how pipelining two computer algorithms based on nucleotide cyclic motifs, MC-Fold and MC-Sym, reproduces a series of experimentally determined RNA three-dimensional structures from the sequence. This demonstrates how crucial the consideration of all base-pairing interactions is in filling the gap between sequence and structure. We use the pipeline to define rules of precursor microRNA folding in double helices, despite the presence of a number of presumed mismatches and bulges, and to propose a new model of the human immunodeficiency virus-1 -1 frame-shifting element.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
The RNA World 3rd edn (eds Gesteland, R. F., Cech, T. R. & Atkins, J. F.) (CSHL, Cold Spring Harbor, 2006)
Griffiths-Jones, S. et al. Rfam: annotating non-coding RNAs in complete genomes. Nucleic Acids Res. 33, D121–D124 (2005)
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007)
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000)
Benson, D. A. et al. GenBank. Nucleic Acids Res. 35, D21–D25 (2007)
Shapiro, B. A. et al. Bridging the gap in RNA structure prediction. Curr. Opin. Struct. Biol. 17, 157–165 (2007)
Mathews, D. H. & Turner, D. H. Prediction of RNA secondary structure by free energy minimization. Curr. Opin. Struct. Biol. 16, 270–278 (2006)
Gutell, R. R., Lee, J. C. & Cannone, J. J. The accuracy of ribosomal RNA comparative structure models. Curr. Opin. Struct. Biol. 12, 301–310 (2002)
Mathews, D. H. Revolutions in RNA secondary structure prediction. J. Mol. Biol. 359, 526–532 (2006)
Mathews, D. H. et al. Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure. Proc. Natl Acad. Sci. USA 101, 7287–7292 (2004)
Major, F. et al. The combination of symbolic and numerical computation for three-dimensional modeling of RNA. Science 253, 1255–1260 (1991)
Lescoute, A. et al. Recurrent structural RNA motifs, isostericity matrices and sequence alignments. Nucleic Acids Res. 33, 2395–2409 (2005)
Dima, R. I., Hyeon, C. & Thirumalai, D. Extracting stacking interaction parameters for RNA from the data set of native structures. J. Mol. Biol. 347, 53–69 (2005)
Do, C. B., Woods, D. A. & Batzoglou, S. CONTRAfold: RNA secondary structure prediction without physics-based models. Bioinformatics 22, e90–e98 (2006)
Das, R. & Baker, D. Automated de novo prediction of native-like RNA tertiary structures. Proc. Natl Acad. Sci. USA (2007)
Lemieux, S. & Major, F. Automated extraction and classification of RNA tertiary structure cyclic motifs. Nucleic Acids Res. 34, 2340–2346 (2006)
Kabsch, H. A discussion of the solution for the best rotation to relate two sets of vectors. Acta Crystallogr. A 34, 827–828 (1978)
Williamson, J. R. Induced fit in RNA-protein recognition. Nature Struct. Biol. 7, 834–837 (2000)
Shankar, N. et al. The NMR structure of an internal loop from 23S ribosomal RNA differs from its structure in crystals of 50s ribosomal subunits. Biochemistry 45, 11776–11789 (2006)
Kondo, J., Urzhumtsev, A. & Westhof, E. Two conformational states in the crystal structure of the Homo sapiens cytoplasmic ribosomal decoding A site. Nucleic Acids Res. 34, 676–685 (2006)
Pley, H. W., Flaherty, K. M. & Mckay, D. B. Three-dimensional structure of a hammerhead ribozyme. Nature 372, 68–74 (1994)
Lee, B. M. et al. Induced fit and “lock and key” recognition of 5S RNA by zinc fingers of transcription factor IIIA. J. Mol. Biol. 357, 275–291 (2006)
Giedroc, D. P., Theimer, C. A. & Nixon, P. L. Structure, stability and function of RNA pseudoknots involved in stimulating ribosomal frameshifting. J. Mol. Biol. 298, 167–185 (2000)
Griffiths-Jones, S. et al. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 34, D140–D144 (2006)
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha–DGCR8 complex. Cell 125, 887–901 (2006)
Macrae, I. J. et al. Structural basis for double-stranded RNA processing by Dicer. Science 311, 195–198 (2006)
Leontis, N. B., Stombaugh, J. & Westhof, E. The non-Watson–Crick base pairs and their associated isostericity matrices. Nucleic Acids Res. 30, 3497–3531 (2002)
Merino, E. J. et al. RNA structure analysis at single nucleotide resolution by selective 2'-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005)
Perret, V. et al. Conformation in solution of yeast tRNAAsp transcripts deprived of modified nucleotides. Biochimie 72, 735–743 (1990)
Brunel, C. et al. Three-dimensional model of Escherichia coli ribosomal 5S RNA as deduced from structure probing in solution and computer modeling. J. Mol. Biol. 221, 293–308 (1991)
Leontis, N. B. & Moore, P. B. NMR evidence for dynamic secondary structure in helices II and III of the RNA of Escherichia coli. Biochemistry 25, 3916–3925 (1986)
Hentze, M. W. & Kuhn, L. C. Molecular control of vertebrate iron metabolism: mRNA-based regulatory circuits operated by iron, nitric oxide, and oxidative stress. Proc. Natl Acad. Sci. USA 93, 8175–8182 (1996)
Jaffrey, S. R. et al. The interaction between the iron-responsive element binding protein and its cognate RNA is highly dependent upon both RNA sequence and structure. Nucleic Acids Res. 21, 4627–4631 (1993)
Sierzputowska-Gracz, H., Mckenzie, R. A. & Theil, E. C. The importance of a single G in the hairpin loop of the iron responsive element (IRE) in ferritin mRNA for structure: an NMR spectroscopy study. Nucleic Acids Res. 23, 146–153 (1995)
Griffiths-Jones, S. et al. Rfam: an RNA family database. Nucleic Acids Res. 31, 439–441 (2003)
Leipuviene, R. & Theil, E. C. The family of iron responsive RNA structures regulated by changes in cellular iron and oxygen. Cell. Mol. Life Sci. (in the press)
Clery, A. et al. An improved definition of the RNA-binding specificity of SECIS-binding protein 2, an essential component of the selenocysteine incorporation machinery. Nucleic Acids Res. 35, 1868–1884 (2007)
Jacks, T. et al. Characterization of ribosomal frameshifting in HIV-1 gag-pol expression. Nature 331, 280–283 (1988)
Gaudin, C. et al. Structure of the RNA signal essential for translational frameshifting in HIV-1. J. Mol. Biol. 349, 1024–1035 (2005)
Staple, D. W. & Butcher, S. E. Solution structure and thermodynamic investigation of the HIV-1 frameshift inducing element. J. Mol. Biol. 349, 1011–1023 (2005)
Acknowledgements
We thank P. Thibault for updating MC-Sym and P. Gendron for helping us with the Condor and web services. We thank D. D’Amours, M.-F. Gaumont-Leclerc and V. Lisi for making suggestions to improve the manuscript. We thank D. H. Mathews and E. Westhof for discussions about MC-Fold. This project was supported by grants from the Canadian Institutes of Health Research (CIHR) and from the Natural Sciences and Engineering Research Council (NSERC) of Canada. M.P. holds Ph.D. scholarships from the NSERC and the Fonds Québécois de la Recherche sur la Nature et les Technologies. F.M. is a member of the Centre Robert-Cedergren of the Université de Montréal.
Author Contributions Both authors were involved in every aspect of the research. M.P. programmed MC-Fold and MC-Cons.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
The file contains Supplementary Methods, Supplementary Discussion, Supplementary Tables S1-S3, Supplementary Figures S1-S15 with Legends and additional references. (PDF 1907 kb)
Rights and permissions
About this article
Cite this article
Parisien, M., Major, F. The MC-Fold and MC-Sym pipeline infers RNA structure from sequence data. Nature 452, 51–55 (2008). https://doi.org/10.1038/nature06684
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06684
This article is cited by
-
Targeted systematic evolution of an RNA platform neutralizing DNMT1 function and controlling DNA methylation
Nature Communications (2023)
-
trRosettaRNA: automated prediction of RNA 3D structure with transformer network
Nature Communications (2023)
-
RNA Folding Based on 5 Beads Model and Multiscale Simulation
Interdisciplinary Sciences: Computational Life Sciences (2023)
-
Designing strategies of small-molecule compounds for modulating non-coding RNAs in cancer therapy
Journal of Hematology & Oncology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.