Abstract
Bacterial flagella contain a specialized secretion apparatus that functions to deliver the protein subunits that form the filament and other structures to outside the membrane1. This apparatus is related to the injectisome used by many gram-negative pathogens and symbionts to transfer effector proteins into host cells; in both systems this export mechanism is termed ‘type III’ secretion2,3. The flagellar secretion apparatus comprises a membrane-embedded complex of about five proteins, and soluble factors, which include export-dedicated chaperones and an ATPase, FliI, that was thought to provide the energy for export1,4. Here we show that flagellar secretion in Salmonella enterica requires the proton motive force (PMF) and does not require ATP hydrolysis by FliI. The export of several flagellar export substrates was prevented by treatment with the protonophore CCCP, with no accompanying decrease in cellular ATP levels. Weak swarming motility and rare flagella were observed in a mutant deleted for FliI and for the non-flagellar type-III secretion ATPases InvJ and SsaN. These findings show that the flagellar secretion apparatus functions as a proton-driven protein exporter and that ATP hydrolysis is not essential for type III secretion.
Similar content being viewed by others
Main
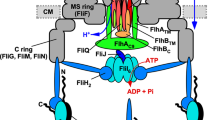
Flagellar assembly begins with structures in the cytoplasmic membrane and proceeds through steps that add the exterior structures in a proximal-to-distal sequence (Fig. 1)1. Assembly of the rod, hook and filament requires the action of the secretion apparatus, which transports the needed subunits into a central channel through the structure that conducts them to their site of incorporation at the tip (Fig. 1). Flagellar export is notably fast: in the early stages of filament growth flagellin is delivered at a rate of several 55 kDa subunits per second5.
Dashed boxes indicate the proteins that function in flagellar secretion, either in the membrane-bound part of the apparatus or in delivery of substrate. Flagellar components that depend on export are indicated in light- (early substrates) or dark- (late substrates) grey; these include the structural proteins that form the rod, hook and filament, the transcriptional regulator FlgM, and the hook-length regulator FliK. OM, outer membrane; PG, peptidoglycan; CM, cytoplasmic membrane.
ATP hydrolysis by FliI was thought to provide the energy for export because mutations that delete or reduce the activity of FliI block flagellar synthesis at the stage of rod assembly1,4,6 (Fig. 1). Homologues of FliI also occur in the type III secretion apparatus of injectisomes and are usually assumed to energize export in those systems as well. Some evidence for a different view has also been reported: it was observed that type III secretion in Yersinia enterocolitica was prevented by the protonophore CCCP7, and it was shown that the secretion ATPase InvC of Salmonella functions to dissociate export substrate from the chaperone8, a role distinct from transport itself. The energy source for type III secretion thus remains uncertain.
To address the energy requirements for type III secretion, we first measured the effect of the uncoupler CCCP on flagellar export in S. enterica, assayed by accumulation of the export substrate FlgM in the medium. FlgM export was prevented by 10 μM or more CCCP (Fig. 2a). Overall cellular energy levels seemed unaffected, because cells grew normally in 10 μM CCCP (growth data not shown) and ATP levels were unchanged (Supplementary Fig. 1). The effect was reversible: FlgM export was largely restored following a 30-min washout of the 10 μM CCCP (Fig. 2b). FlgM inhibits transcription of its own gene, and so reduced FlgM export might partially reflect decreased cellular levels of the protein9. To circumvent this autoinhibitory effect the experiment was repeated with a strain in which flgM was placed under control of a non-native (P araB ) promoter. In this strain, the cytoplasmic level of FlgM remained nearly constant, whereas FlgM secretion was again prevented by 10 μM or more CCCP (Fig. 2c).
a, Secretion in Salmonella strain TH3730 (Tet-inducible flhDC). b, Partial restoration of export following a 30-min wash into CCCP-free buffer. c, Secretion in strain TH10874 (arabinose-inducible flgM). d, Inhibition of FlgM secretion by CCCP in an ATP-synthase defective (ΔatpA) strain (TH11802). e, ATP levels in the ΔatpA mutant at various times following treatment with CCCP. Open circles, no treatment; open square, 10 μM CCCP; and filled circles, 30 μM CCCP. f, Inhibition of secretion of other flagellar substrates (FlgK, FlgL, FliC and FliK) by CCCP.
The maintenance of normal ATP levels in the presence of CCCP was noted previously in experiments with Y. enterocolitica7 and is thought to be due to a regulatory mechanism that inhibits the hydrolytic activity of the ATP synthase when the membrane is de-energized10. Because this protective mechanism may not act instantaneously, cellular ATP levels might undergo a transient drop following CCCP treatment that would escape detection in our measurements. To rule out such an effect, we measured FlgM export in a ΔatpA strain that lacks a major subunit of the ATP synthase. FlgM secretion was again prevented by 10 μM or higher CCCP, and ATP levels were unaffected (Fig. 2d, e).
The flagellum exports more than a dozen substrates, which are classified as early or late according to whether they are secreted during assembly of the hook/basal-body or the filament1 (Fig. 1). To test the generality of the PMF requirement we examined the effect of CCCP on secretion of additional substrates, assayed by their accumulation in the culture medium11. We observed the early substrate FliK and the late-export substrates FlgK, FlgL and FliC in culture supernatants of LB-grown cells. Accumulation of all four substrates was prevented by treatment with 20 μM CCCP (Fig. 2f). The band between FliC and FlgK does not correspond to the size of a known flagellar protein and is likely to be a stable protein secreted in (relatively) low amounts.
CCCP functions as a proton carrier to discharge both the electric potential (Δψ) and concentration (ΔpH) components of the PMF. To examine the contribution of the Δψ component separately, we measured FlgM secretion in cells treated with the K+-ionophore valinomycin. In medium containing 150 mM KCl, FlgM secretion was inhibited by 10 μM valinomycin (Fig. 3a). Thus, the electrical potential component seems essential for export under the conditions of this experiment (extracellular pH = 7.3). Next, the ΔpH component of the gradient was discharged by the weak acid acetate (34 mM), which crosses the membrane in neutral (protonated) form and releases protons inside the cell. At an external pH of 7, FlgM secretion was not affected by treatment with acetate (Fig. 3b), indicating that flagellar export can be supported by Δψ alone. At an external pH of 5, acetate prevented secretion (Fig. 3b), presumably owing to acidification of the cytoplasm and the resulting protonation of one or more functionally important acidic groups. A similar effect was reported with the flagellar motor, which ceased rotating when the cytosolic pH was lowered to 5 (ref. 12).
We characterized export requirements further using an assay based on secretion of a FlgE–Bla (hook/β-lactamase) fusion protein. Cells were deleted for the rod proteins FlgB and FlgC (Fig. 1) to direct the fusion protein into the periplasm, allowing export to be quantified by the MIC (minimum inhibitory concentration) for ampicillin13. The MIC value was reduced by uncoupler, from a value of 25 μg ml-1 in the untreated control to about 4 μg ml-1 in 30 μM CCCP (Table 1, and Supplementary Information).
If energy for flagellar transport comes from the proton gradient, then ATP hydrolysis by FliI may be less important than has been supposed. To examine the FliI requirement more closely we measured FlgE–Bla export in a strain deleted for fliI and the flanking genes fliH and fliJ. FliH is a regulator of FliI14 and FliJ interacts with the FliHI complex and with other export components15,16. The MIC measured for the ΔfliHIJ strain was 12 μg ml-1, reproducibly larger than that of a negative-control strain lacking the MS-ring gene fliF (<3 μg ml-1) or a strain with all the flagellar genes repressed by downregulation of the master regulators flhDC (<3 μg ml-1) (Table 1). Furthermore, the MIC value of the ΔfliHIJ strain was greatly increased (to 800 μg ml-1) by overexpression of FliR, a membrane-associated part of the export apparatus (Table 1, and Supplementary Data). Prompted by this evidence of export in the absence of FliI, we examined motility of ΔfliHIJ cells. The ΔfliHIJ cells migrated in soft agar at about one-tenth the wild-type rate (Fig. 4a), and a fraction of the cells were observed to swim in liquid media. Cells isolated from the centre, edges or intermediate positions in the swarm showed the same phenotype when purified and re-tested (not shown), and so the slow motility is a property of the population and is not due to suppressing mutations. Staining showed flagella on a small fraction (<1%) of the cells (Fig. 4b, left panel). A ΔfliI mutant swarmed more slowly than the ΔfliHIJ strain (Fig. 4a) and also showed reduced export in the MIC assay (Table 1), consistent with the more severe motility defect reported previously for a ΔfliI mutant4,17.
a, Swarming of the ΔfliHIJ deletion strain and a ΔfliHIJ ΔinvC ΔssaN strain with all type-III secretion ATPases deleted. The ΔfliF strain, blocked in the earliest step of flagellar assembly (Fig. 1), is included as a negative control. Plates were incubated at 32 °C. b, Flagella on cells of the ΔfliHIJ mutant (left panel) and the ΔfliHIJ ΔinvC ΔssaN triple-deletion (right panel). Flagella were visualized with FITC-conjugated anti-FliC antibody (green)26. DNA was stained with DAPI (blue), and membranes by FM64 (red).
In addition to the flagella apparatus, members of Salmonella spp. contain two non-flagellar (injectisome) type III secretion systems, with associated ATPases InvC and SsaN8,18. To rule out any involvement of InvC or SsaN in the secretion observed in ΔfliHIJ cells, we repeated the experiments in a ΔinvC ΔssaN ΔfliHIJ strain. The triple-deletion mutant swarmed equally as well as the ΔfliHIJ strain in soft agar (Fig. 4a), and flagella were again seen on a few cells (Fig. 4b, right). Thus, none of the secretion ATPases is required for flagellar export, assembly or function.
Our conclusions are consistent with previous observations of a PMF requirement for flagellar growth from more than 25 years ago19, and extend the earlier findings in showing that export can be energized by PMF alone in the absence of any type III secretion ATPase7. Use of the proton gradient is perhaps not surprising given the speed of type III secretion and the likely advantage of tapping a proximal energy source. Rapid subunit export presumably requires a rapid supply of energy, which might be more easily delivered by a proton current than by ATP hydrolysis. Given that type III secretion is energized by PMF, future studies should focus on the molecular mechanism of proton movement through the apparatus and its coupling to movement of substrate.
Methods Summary
Media, growth conditions, methods for phage-mediated transduction and motility assays were performed as described previously9,20,21. Carbonylcyanide m-chlorophenylhydrozone (CCCP) and valinomycin were from Sigma (analytical grade). Potassium acetate was from J. T. Baker, growth media from Difco, and buffers from Sigma.
FlgM secretion was assayed by accumulation of the protein in the culture medium, using procedures described previously22 with minor modifications described in Methods. Cells at mid-log growth stage were treated with the PMF-discharging agents (CCCP or valinomycin) at the concentrations indicated in the figures, introduced from freshly prepared stocks. Following centrifugation to remove cells, supernatants were filtered and FlgM was detected by immunoblotting, essentially as described23.
ATP was measured using the firefly luciferase assay and sample-processing procedures of ref. 24.
Ampicillin resistance in FlgE–Bla-exporting cells was assayed as described13, with minor modifications described in Methods. Briefly, cells were diluted to D600 = 0.01, cultured for 4.5 h in LB media containing ampicillin in a series of dilutions ranging from 800 μg ml-1 to 1.56 μg ml-1, and the minimum inhibitory concentration (MIC) was taken as the lowest ampicillin concentration giving D600 < 0.05. The periplasmic fractionation and anti-Bla immunoblot detection of FlgE/Bla in the cellular and periplasmic fractions was performed as described by ref. 25.
Swarming motility was assayed in plates containing tryptone broth and 0.28% bacto-agar. Plates were incubated at 32 °C. Flagellar immunostaining used the methods of ref. 26.
Online Methods
FlgM-secretion assay
FlgM secreted into the culture medium was measured as described previously22, with the following modifications. Strain TH3730 (Tc-induced flagellar synthesis) was cultured overnight at 37 °C in LB media, then diluted 100-fold into 3 ml of fresh LB and cultured to anD600 = 0.5. Tetraycline (15 μg ml-1) was added to induce transcription of the flagellar regulatory genes flhDC and growth was continued for one hour at 37 °C before cells were harvested. Strain LT2 was cultured in the same way except without tetracycline induction and harvested at an D600 = 0.8. Cells were collected by centrifugation and washed twice with 3 ml LB containing appropriate concentrations of the PMF-discharging agents (CCCP or valinomycin), introduced from freshly prepared stocks in DMSO. Controls received only DMSO. Following resuspension the cells were incubated at 37 °C for the times indicated in the figures (typically 30 min), then kept on ice until further processing. Strains TH10874 and TH11802 were treated in a similar way, except media included 0.2% L-arabinose to induce transcription of flgM.
To quantify FlgM in the culture medium, cells of a 2-ml sample were pelleted in a tabletop centrifuge and 1.8 ml of the supernatant was passed through a 0.45-μm cellulose acetate filter (Nalgene) to remove any remaining cells. Filtrate (1.6 ml) was filtered through a prewetted 0.45 μm pore-size BA85 nitrocellulose filter (Schleicher & Schuell) for protein binding. Proteins were eluted by addition of 40 μl of 2× SDS sample buffer27 and heating to 65 °C for 30 min. For quantification of intracellular FlgM, the pelleted cells were resuspended in 50 μl 1× SDS sample buffer and boiled for 10 min. Pellet and supernatant fractions were adjusted to equal volumes (corresponding to 20 D units for the cells), 10 μl samples were loaded onto SDS–PAGE gels and FlgM was detected by immunoblotting, essentially as described23.
ATP measurement
ATP was measured using the firefly luciferase assay (time-stable ATP determination kit, Biaffin) and the sample-processing procedures of ref. 24. Following resuspension in the media with or without CCCP, cells were incubated at 37 °C for the times indicated, then 2-ml samples were withdrawn and mixed with 2 ml of ice-cold 6% HClO4, 2.5 mM Na2HPO4, 1 mM KCl. Samples were left on ice for 30 min then centrifuged in the cold to remove denatured protein. Supernatants were brought to pH 7.0 with 2 M KOH, 0.3 M MOPS, then frozen at -20°C, quickly thawed, and immediately centrifuged at 4 °C to remove crystals of KClO4. Samples were kept on ice and used for ATP determination within 2 h. For ATP measurements, 50 μl of supernatant was mixed with an equal volume of luciferase reagents in a 96-well plate and the luminescence was measured at 560 nm using a TopCount NXT Microplate scintillation and luminescence counter (Packard).
FlgE/β-lactamase activity
FlgE/β-lactamase assays were performed essentially as described by in ref. 13, with minor modifications. Briefly, 1-ml cultures in LB were grown overnight at 37 °C. Cultures were diluted 100-fold into 3 ml LB and grown at 37 °C to a D600 = 0.5–0.7, then diluted 50-fold into fresh LB media containing ampicillin in a series of dilutions ranging from 800 μg ml-1 to 1.56 μg ml-1. The D600 of each sample was measured after 4.5 h of further growth at 37 °C, and the minimum inhibitory concentration (MIC) was taken as the lowest ampicillin concentration giving D600 < 0.05. For the experiment that measured the effect of CCCP, the flgE–bla gene was expressed from the ara promoter, induced with arabinose 30 min before addition of CCCP, to minimize FlgE/Bla accumulation in the periplasm before the addition of the uncoupler.
Staining
Flagellar immunostaining was performed as described in ref. 26.
Measurement of FlgE–Bla in periplasm
Spheroplast preparation, periplasmic fractionation and immunoblot detection of FlgE–Bla in the cellular and periplasmic fractions was performed as described in ref. 25.
References
Macnab, R. M. How bacteria assemble flagella. Annu. Rev. Microbiol. 57, 77–100 (2003)
Cornelis, G. R. The type III secretion injectisome. Nature Rev. Microbiol. 4, 811–825 (2006)
Blocker, A., Komoriya, K. & Aizawa, S. Type III secretion systems and bacterial flagella: insights into their function from structural similarities. Proc. Natl Acad. Sci. USA 100, 3027–3030 (2003)
Fan, F. & Macnab, R. M. Enzymatic characterization of FliI. An ATPase involved in flagellar assembly in Salmonella typhimurium . J. Biol. Chem. 271, 31981–31988 (1996)
Iino, T. Assembly of Salmonella flagellin in vitro and in vivo . J. Supramol. Struct. 2, 372–384 (1974)
Vogler, A. P., Homma, M., Irikura, V. M. & Macnab, R. M. Salmonella typhimurium mutants defective in flagellar filament regrowth and sequence similarity of FliI to F0F1, vacuolar, and archaebacterial ATPase subunits. J. Bacteriol. 173, 3564–3572 (1991)
Wilharm, G. et al. Yersinia enterocolitica type III secretion depends on the proton motive force but not on the flagellar motor components MotA and MotB. Infect. Immun. 72, 4004–4009 (2004)
Akeda, Y. & Galan, J. E. Chaperone release and unfolding of substrates in type III secretion. Nature 437, 911–915 (2005)
Gillen, K. L. & Hughes, K. T. Negative regulatory loci coupling flagellin synthesis to flagellar assembly in Salmonella typhimurium . J. Bacteriol. 173, 2301–2310 (1991)
Suzuki, T. et al. F0F1-ATPase/synthase is geared to the synthesis mode by conformational rearrangement of ε subunit in response to proton motive force and ADP/ATP balance. J. Biol. Chem. 278, 46840–46846 (2003)
Komoriya, K. et al. Flagellar proteins and type III-exported virulence factors are the predominant proteins secreted into the culture media of Salmonella typhimurium . Mol. Microbiol. 34, 767–779 (1999)
Minamino, T., Imae, Y., Oosawa, F., Kobayashi, Y. & Oosawa, K. Effect of intracellular pH on rotational speed of bacterial flagellar motors. J. Bacteriol. 185, 1190–1194 (2003)
Lee, H. J. & Hughes, K. T. Posttranscriptional control of the Salmonella enterica flagellar hook protein FlgE. J. Bacteriol. 188, 3308–3316 (2006)
Minamino, T. & MacNab, R. M. FliH, a soluble component of the type III flagellar export apparatus of Salmonella, forms a complex with FliI and inhibits its ATPase activity. Mol. Microbiol. 37, 1494–1503 (2000)
Fraser, G. M., Gonzalez-Pedrajo, B., Tame, J. R. & Macnab, R. M. Interactions of FliJ with the Salmonella type III flagellar export apparatus. J. Bacteriol. 185, 5546–5554 (2003)
Evans, L. D., Stafford, G. P., Ahmed, S., Fraser, G. M. & Hughes, C. An escort mechanism for cycling of export chaperones during flagellum assembly. Proc. Natl Acad. Sci. USA 103, 17474–17479 (2006)
Dreyfus, G., Williams, A. W., Kawagishi, I. & Macnab, R. M. Genetic and biochemical analysis of Salmonella typhimurium FliI, a flagellar protein related to the catalytic subunit of the F0F1 ATPase and to virulence proteins of mammalian and plant pathogens. J. Bacteriol. 175, 3131–3138 (1993)
Hensel, M. et al. Functional analysis of ssaJ and the ssaK/U operon, 13 genes encoding components of the type III secretion apparatus of Salmonella pathogenicity island 2. Mol. Microbiol. 24, 155–167 (1997)
Galperin, M., Dibrov, P. A. & Glagolev, A. N. ΔμH+ is required for flagellar growth in Escherichia coli . FEBS Lett. 143, 319–322 (1982)
Davis, R. W., Botstein, D. & Roth, J. R. Advanced Bacterial Genetics (Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, 1980)
Gillen, K. L. & Hughes, K. T. Molecular characterization of flgM, a gene encoding a negative regulator of flagellin synthesis in Salmonella typhimurium . J. Bacteriol. 173, 6453–6459 (1991)
Karlinsey, J. E., Tsui, H.-C. T., Winkler, M. E. & Hughes, K. T. Flk couples flgM translation to flagellar ring assembly in Salmonella typhimurium . J. Bacteriol. 180, 5384–5397 (1998)
Hughes, K. T., Gillen, K. L., Semon, M. J. & Karlinsey, J. E. Sensing structural intermediates in bacterial flagellar assembly by export of a negative regulator. Science 262, 1277–1280 (1993)
Bakker, E. P. & Mangerich, W. E. The effects of weak acids on potassium uptake by Escherichia coli K-12 inhibition by low cytoplasmic pH. Biochim. Biophys. Acta 730, 379–386 (1983)
Aldridge, P., Karlinsey, J. E., Becker, E., Chevance, F. F. & Hughes, K. T. Flk prevents premature secretion of the anti-sigma factor FlgM into the periplasm. Mol. Microbiol. 60, 630–643 (2006)
Rosu, V. & Hughes, K. T. σ28-dependent transcription in Salmonella enterica is independent of flagellar shearing. J. Bacteriol. 188, 5196–5203 (2006)
Laemmli, U. K. & Favre, M. Maturation of the head of bacteriophage T4. I. DNA packaging events. J. Mol. Biol. 80, 575–599 (1973)
Acknowledgements
We thank S. Williams for his help and permission for using the luminometer, F. Chevance for advice and assistance with strain constructions, M. Sarkar for assistance with MIC assays, and V. Clougherty for flagellar-staining experiments. M. Erhardt gratefully acknowledges scholarship support of the Studienstiftung des deutschen Volkes. This work was supported by Public Service grants (K.T.H. and D.F.B.) from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Figure
This file contains Supplementary Figure S1 with Legend. (PDF 1238 kb)
Rights and permissions
About this article
Cite this article
Paul, K., Erhardt, M., Hirano, T. et al. Energy source of flagellar type III secretion. Nature 451, 489–492 (2008). https://doi.org/10.1038/nature06497
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06497
This article is cited by
-
A small molecule, C24H17ClN4O2S, inhibits the function of the type III secretion system in Salmonella Typhimurium
Journal of Genetic Engineering and Biotechnology (2022)
-
Insights into the structure of Escherichia coli outer membrane as the target for engineering microbial cell factories
Microbial Cell Factories (2021)
-
Reversion of antibiotic resistance in multidrug-resistant pathogens using non-antibiotic pharmaceutical benzydamine
Communications Biology (2021)
-
Substrate-engaged type III secretion system structures reveal gating mechanism for unfolded protein translocation
Nature Communications (2021)
-
The FlgN chaperone activates the Na+-driven engine of the Salmonella flagellar protein export apparatus
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.