Abstract
Where do animal behaviours come from and are they controlled by genes? This is the fundamental question posed by the field of neurogenetics. Pioneering work from the 1960s in Seymour Benzer’s laboratory demonstrated for the first time that Drosophila melanogaster fruitflies could be mutated to obtain animals with insomnia, learning disabilities and homosexual courtship behaviours.
Similar content being viewed by others
Main
Forty years ago, the field of Drosophila neurogenetics (defined in Box 1) was born in Seymour Benzer’s laboratory at Caltech1. In the mid-1960s, Benzer made an abrupt and orthogonal turn from his early ground-breaking work defining the fine structure of the gene in bacteriophage2 to the heretical idea that single genes could control behaviour in complex animals. A paper appearing in the September 1967 issue of Proc. Natl Acad. Sci. presented the first evidence that mutant flies defective in phototaxis behaviour, or locomotor responses to light, could be identified1. The premise of neurogenetics—widely disbelieved at the time—was that complex behaviours such as the ability to learn and remember, the internal biological rhythms of the body, and courtship and sexuality could all be under genetic control.
Drosophila melanogaster as an experimental organism has contributed much to contemporary neurobiology. The first cloning of a structural gene for a potassium channel was achieved by Benzer’s trainees, Yuh Nung Jan and Lily Jan, when they isolated the gene corresponding to the shaker (sh) mutant3. The founding member of the now enormous transient receptor potential (trp) ion channel family had its origins as a fly mutant defective in light-evoked retinal electrophysiology4. Vertebrate and invertebrate TRP channels have since turned up in biological processes as diverse as the sensation of odours, tastes, pungent compounds such as wasabi, capsaicin and menthol, cold, heat, touch and hearing, among others (reviewed in ref. 5). Beginning in the late 1960s, William Pak amassed a large collection of mutants defective in visual signal transduction, such as the neither inactivation nor afterpotential (nina) mutants6. This genetic dissection of phototransduction in Drosophila enabled later molecular analysis of the molecules underlying visual signal transduction in the laboratories of Pak, Gerald Rubin, Charles Zuker and others (reviewed in ref. 7).
The cloning of sh and trp are excellent examples of the power of neurogenetics. Both arose from genetic screens designed to test the hypothesis that studying shaky flies or flies with altered retinal physiology would lead to interesting insights into neural function. The tools of this discipline are simple and require only a suitable behavioural paradigm (three are shown in Fig. 1), a means to make flies with mutations in single genes, and standard molecular genetic techniques to progress from a mutant phenotype to a genotype.
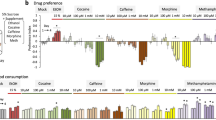
a. The olfactory T-maze is used for Pavlovian olfactory conditioning40,64. Flies are trained to associate odour A (orange) with electric shock (left). During testing, these flies avoid odour A (right). The assay is carried out with reciprocal training, such that only one half of the paradigm is depicted here. Flies are depicted as small black dots. b, Circadian-activity monitors measure locomotor activity of individual flies using an infrared beam (red dotted line)10. An external computer tracks the number of times the fly breaks the beam, allowing continual monitoring of fly locomotor activity over a period of weeks. c, The courtship wheel permits the observation of up to 10 fly couples in which the male engages in such stereotyped sexual activity as the following: genital licking, wing vibration to produce a species-specific song, and copulation. Graphic in a is adapted from figure 1 of ref. 65 with permission from Elsevier (copyright 2004).
This Review article will discuss how the revolution started by Benzer and his students in 1967 has spread to many fields of neurobiological investigation in Drosophila, from whence it jumped to mice, zebrafish and other species, including humans. Here I will focus specifically on three original discoveries in Drosophila neurogenetics and behaviour—biological rhythms, sexual courtship and chemoreception—and how these have blossomed in the last 40 yr.
The genetics of circadian rhythms in the fly
Flies, like all other animals and plants on earth, have a daily routine in synchrony with the rhythms of the Sun and Earth. Like humans, flies tend to wake up around dawn, enjoy a siesta in the afternoon, and are largely inactive after nightfall8. The biological rhythm in locomotor activity recurs on a roughly 24 h cycle, hence it is termed a circadian (circa diem—around a day) rhythm. This modulation of locomotor behaviour is driven by external environmental rhythms, but can also persist in flies raised for generations in the dark9.
Ronald Konopka in Benzer’s laboratory provided the first evidence that the biological clock was under genetic control and could be broken by mutagenesis10. In an elegant and simple screen for flies with altered hatching and locomotor rhythms, using activity-monitoring devices such as those in Fig. 1b, Konopka and Benzer isolated three different mutant alleles of the same gene, called period (per). per0 flies are insomniac, pers flies live a short day, and perl flies a long day10. An amazing parallel with humans was uncovered with the recent identification of mutations in a human period homologue (PER2) as the genetic culprit behind the pers -like phenotype in familial advanced sleep-phase syndrome11
How a single gene could both be necessary for the clock but also set its running speed remained a mystery until the age of molecular cloning and the isolation of other clock genes. The groups of Michael Rosbash and Jeffrey Hall, and the group of Michael Young cloned per in 1984 (refs 12, 13). Both per messenger RNA and PER protein were subsequently shown to cycle with a circadian rhythm, and show a rhythmic nuclear accumulation, prompting a model in which PER acts as a feedback suppressor to control the clock14. per turned out to be just the tip of an enormous iceberg of clock genes, clock accessory genes, and clock-controlled genes. The present model for the Drosophila clock includes a host of core clock components that include a positive transcriptional feedback loop (Clock (Clk), cycle (cyc) and vrille (vri)), a negative transcriptional feedback loop (per and timeless (tim)), and factors that modulate the light-regulated accumulation and output of the core clock genes (double-time (dbt; also known as discs overgrown or dco), shaggy (sgg), cryptochrome (cry), Pigment-dispersing factor (Pdf) and others; reviewed in ref. 8). Some recent surprises in the clock field include a somewhat mysterious cytoplasmic timing mechanism that regulates the delay in nuclear accumulation of period and timeless15, as well as the discovery that the clock protein has chromatin-remodelling activity16.
Homologues of many of the core clock genes have been identified in vertebrates, further validating the fly as a model for circadian biology. Microarray studies by Young17 and others have identified several hundred genes under circadian control, the analysis of which promises to provide an integrated view of how the physiology of the entire organism is synchronized to the daily rhythms of the planet.
fruitless and its power to shape sexual behaviour
Copulation in Drosophila is preceded by an intricate series of sexually dimorphic pre-copulatory courtship behaviours between the male and female fly18 (Fig. 1c). Benzer’s trainees Hall and Yoshiki Hotta (Box 2) used genetic mosaic analysis to define portions of the central nervous system required for male courtship behaviour19,20 and genes that governed heterosexual behaviour21. One gene named fruitless (fru), identified in 1963 by K. S. Gill and cloned over 30 yr later by Daisuke Yamamoto22—and separately by the group effort of Hall, Bruce Baker and Barbara Taylor23—is now known to be a master regulator of sexuality in the fly23,24. The transcription factor encoded by the fru gene is expressed in a subset of central, peripheral, sensory and motor neurons in the adult fly, which are likely to comprise a circuit controlling sexually dimorphic behaviour25,26,27. Mutant fru males show homosexual courtship behaviour in which large groups form chains of males courting each other. In a remarkable experiment, Barry Dickson showed recently that male courtship behaviour directed at females can be induced in chromosomally female flies simply by expressing the male-specific isoform of fru in the female brain24. Recent work in the mouse from Catherine Dulac’s group suggests a similar underlying latency in the female mouse to exhibit male behaviours on manipulation of a single gene28. A major goal in this field is to define the molecular targets of fru and define the neural circuits that drive both male and female sexual behaviours.
Olfactory communication in the fly
Fruitflies are strongly attracted to the smell of vinegar, yeast, rotting fruit and to each other. The genetic basis of this chemosensory behaviour was first studied by Obaid Siddiqi, a Benzer trainee (Box 2). Mutants defective in the olfactory T-maze, using the device in Fig. 1a but omitting electric shock, as well as other olfactory behaviour paradigms were collected throughout the 1980s by Siddiqi29, and later by John Carlson30, and others. One of the Carlson mutants, acj6, proved to be a key transcription factor necessary for the regulation of a subset of odorant receptor genes31. The availability of the genome sequence of Drosophila melanogaster opened this system to rapid molecular analysis by Carlson, Dean Smith, Liqun Luo, Dickson, Richard Axel and a number of former Axel trainees, resulting in the complete description of the sequence and expression of all 62 odorant receptors and 68 taste receptors (reviewed in ref. 32), the complete map of connectivity of primary olfactory centres33,34, an initial view of how primary olfactory information is mapped in the higher brain35,36, and a comprehensive survey of ligand tuning of a majority of the odorant receptors37, including those tuned to pheromones38,39. A major effort in this growing field is to understand the underlying central mechanisms by which a fly discriminates among all the odours it is able to detect and how the circuitry underlying pheromone perception leads to stereotyped behaviours.
A myriad of other complex behaviours
Beyond these brief examples from the original neurogenetic studies to emerge from the Benzer laboratory, many other behaviours have been productively dissected with genetic and behavioural tools in Drosophila. The seminal work of William Quinn, William Harris and Benzer (Box 2) demonstrating that flies can be conditioned to avoid an odour paired with shock (Fig. 1a)40, was followed by the identification of a series of mutant flies that either could not learn this task or rapidly forgot it (reviewed in ref. 41). Subsequent genetic analysis by Quinn, Ronald Davis, Tim Tully and others produced the provocative finding that many learning and memory defective mutations in the fly affect the cyclic AMP pathway (reviewed in ref. 41), the same signalling pathway implicated in conditioned behaviours in Aplysia and the mouse42,43. Subsequent genetic screens for learning mutants by Tully and others, in one case combined with microarray analysis, produced a host of other candidate memory genes, including several involved in local control of mRNA translation44.
The cloning of the dunce (dnc) gene45 and its enrichment in a part of the fly brain called the mushroom body46 allowed the field to move from the genetic to the cellular level. Davis, Martin Heisenberg and others carried out a series of genetic and ablation studies strongly implicating this olfactory processing centre in the fly as the seat of memory47,48. Current work in the field is zeroing in on how fly-brain microcircuitry processes paired odour and shock input49, how the circuitry is modulated by conditioning50, and how the processes of learning and retrieval of memories are compartmentalized51. Neurogenetics has also enabled scientists to localize memory to smaller and smaller areas of the fly brain. A particularly elegant recent example comes from Heisenberg, Li Liu and co-workers, who localized circuits that learn certain visual features to two groups of neurons in a structure called the fan-shaped body52. Although the small size of Drosophila central-brain neurons has hindered electrophysiological access, recent work from Rachel Wilson and Gilles Laurent suggests that this barrier is not insurmountable53, and a more detailed functional analysis of memory at the level of single mushroom-bodies seems likely.
Ulrike Heberlein has turned the fly into a genetic model for alcohol intoxication, demonstrating that flies exhibit progressive and eerily human-like responses to acute alcohol exposure: first, they become hyperexcitable, then they lose coordination, and finally they pass out54. Some of the same cAMP pathway genes required for learning and memory affect a fly’s sensitivity to alcohol55.
Both Edward Kravitz, who studies aggression in lobster, and Ralph Greenspan have recently turned their attention to fly aggression. Flies, both male and female, exhibit aggressive behaviours, with males fighting other males in the presence of a female and females jousting with females over food resources56. Kravitz has shown that fighting style differs between the sexes and is controlled by fru (ref. 57). Multi-generational selection for aggressive or docile strains has been achieved by Greenspan, and such strains have been analysed by whole-genome microarray to identify a large number of genes, the expression of which is modulated differentially in aggressive strains58. These genes will provide avenues for future investigation into the genetic basis of aggression.
A behavioural paradigm recently pioneered by Roland Strauss is that of gap-crossing (Fig. 2). Flies are presented with gaps of varying widths, from narrow and easy to cross to unbreachable chasms, and make sophisticated estimations of which gaps can be reasonably crossed59.This goal-directed climbing behaviour is useful to dissect motor planning and coordination, and to identify the circuits in the fly brain that estimate distance, but could, in principle, also lead to mutants with altered appetite for risk. It is conceivable that both risk-averse flies, capable of crossing a gap but choosing not to, and reckless flies, those choosing to cross impossibly wide gaps, could be identified through genetic screens.
Photo by S. Pick, kindly provided by R. Strauss. Reprinted from ref. 59 with permission from Elsevier (copyright 2005).
Unlike the classic eusocial insects such as ants and bees, flies are not typically known for their group dynamics. This view has been changing somewhat on closer behavioural investigation, which has revealed some surprising evidence of social interactions in Drosophila. For instance, Joel Levine and Hall have shown that circadian rhythms can be phase-shifted by the odour of flies living in another time zone or flies of another genotype60. Hubert Amrein showed that normal circadian locomotor activity of a male is drastically affected by the presence of a female61. These experiments hint at as yet unknown volatile chemical substances produced by other flies and detected by the olfactory system, and suggest that social interactions shape group interactions in the fly. In fact there is a growing trend to monitor Drosophila behaviour in more natural and enriched contextual environments that mimic those they might encounter in the real world. For instance, Levine has been observing fly social interactions in groups in the presence of food (Fig. 3). Free from the constraints of courting a single female in an austere Plexiglas chamber, as is the norm for observing courtship behaviour (Fig. 1c), Drosophila males in group situations seem to engage in complex group sex that combines foreplay, copulation and feeding behaviours (Fig. 3). It will be interesting to study the regulation and modulation of such group social behaviours and the importance of context in regulating them.
a, Three males courting a single virgin female near a wedge of food. b, Male with forelegs raised high copulates with a female, while another male, on his back, touches and licks her abdomen. This occurs on top of a wedge of food. Note that the female’s right foreleg stretches out across the surface of the food as does the left foreleg of the male beneath her. Such sexual behaviour is affected by the presence or absence of food in the assay. Gustatory receptors on the tarsi, the part of the foreleg in contact with the sweet food, are in a good position to sample food and may play a mechanistic role in this sexual interaction. Photo by N. Stepek and J.-C. Billeter, kindly provided by J. D. Levine, Univ. of Toronto, Mississauga.
A related example of a behaviour that emerges in groups is the innate avoidance that flies show for an empty tube previously occupied by flies that experienced stress. Avoidance by naive flies of tubes previously occupied by shaken flies was first noticed by Benzer in 1967 (ref. 1), and subsequently investigated by Greg Suh, working with the Benzer laboratory and the groups of David Anderson and Axel, as an innate olfactory avoidance of a Drosophila stress odor, dSO (ref. 62). This robust behaviour, resulting from the recognition and avoidance of the smell of a fellow fly in trouble, will be useful in future studies of the circuitry of anxiety, stress and innate fear.
Concluding remarks
Significant advances in our understanding of the biological clock, sensory systems, learning and memory, sexual courtship and many other behaviours have been made through neurogenetic research in Drosophila. With these successes behind us, some adventurous Drosophila neurogeneticists are moving beyond these original neurogenetics questions, which may in hindsight represent the low-hanging fruit—robust behaviours amenable to investigation in laboratory-based behavioural paradigms. It now seems possible to approach in the fly more complex behaviours and even emotions, the neurobiological basis of which are not well understood at the genetic or functional level in any animal: sociality, common sense, altruism, empathy, frustration, motivation, hatred, jealousy, peer pressure, and so on. The only a priori limitation to studying any of these traits is the belief that flies can show such emotions and the design of a plausible behavioural paradigm to measure them.
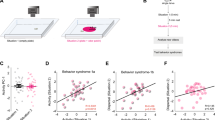
This Progress article accompanies the release of complete genomes of eleven additional Drosophila species (D. ananassae, D. erecta, D. grimshawi, D. mojavensis, D. pseudoobscura, D. simulans, D. virilis, D. yakuba, D. persimilis, D. sechellia, D. willistoni), with vastly different ecologies and lifestyles to Drosophila melanogaster. What will be the impact of these additional Drosophila genomes on neurogenetics and behaviour research? Such information may begin to provide clues to differences in pheromonal communication and species recognition among these flies, some of which occupy overlapping ecological niches and need to pay careful attention to which species they are courting. A second area of interest will be in food preference and how this might be influenced by the evolution of smell and taste receptors. Are there functional differences in chemosensory reception of a fly with omnivorous taste as compared to a fly species with more specialized tastes? Hints that such phenomena are both existent and genetically tractable come from recent work in Bill Hansson’s group, which found that the Seychelles island species D. sechellia has an olfactory system specialized to sense its preferred food, the Noni fruit63.
The little vinegar fly Drosophila melanogaster, along with its sister species, promises to reveal many more surprises about how the nervous system produces complex behaviours in the next 40 yr.
References
Benzer, S. Behavioral mutants of Drosophila isolated by countercurrent distribution. Proc. Natl Acad. Sci. USA 58, 1112–1119 (1967)
Benzer, S. Fine Structure of a genetic region in bacteriophage. Proc. Natl Acad. Sci. USA 41, 344–354 (1955)
Papazian, D. M., Schwarz, T. L., Tempel, B. L., Jan, Y. N. & Jan, L. Y. Cloning of genomic and complementary DNA from Shaker, a putative potassium channel gene from Drosophila . Science 237, 749–753 (1987)
Cosens, D. J. & Manning, A. Abnormal electroretinogram from a Drosophila mutant. Nature 224, 285–287 (1969)
Montell, C. The TRP superfamily of cation channels. Sci. STKE 2005, re3 (2005)
Pak, W. L., Grossfield, J. & Arnold, K. S. Mutants of the visual pathway of Drosophila melanogaster . Nature 227, 518–520 (1970)
Smith, D. P., Stamnes, M. A. & Zuker, C. S. Signal transduction in the visual system of Drosophila . Annu. Rev. Cell Biol. 7, 161–190 (1991)
Hardin, P. E. The circadian timekeeping system of Drosophila . Curr. Biol. 15, R714–R722 (2005)
Sehgal, A., Price, J. & Young, M. W. Ontogeny of a biological clock in Drosophila melanogaster . Proc. Natl Acad. Sci. USA 89, 1423–1427 (1992)
Konopka, R. J. & Benzer, S. Clock mutants of Drosophila melanogaster . Proc. Natl Acad. Sci. USA 68, 2112–2116 (1971)
Toh, K. L. et al. An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291, 1040–1043 (2001)
Bargiello, T. A., Jackson, F. R. & Young, M. W. Restoration of circadian behavioural rhythms by gene transfer in Drosophila . Nature 312, 752–754 (1984)
Zehring, W. A. et al. P-element transformation with period locus DNA restores rhythmicity to mutant, arrhythmic Drosophila melanogaster . Cell 39, 369–376 (1984)
Hardin, P. E., Hall, J. C. & Rosbash, M. Feedback of the Drosophila period gene product on circadian cycling of its messenger RNA levels. Nature 343, 536–540 (1990)
Meyer, P., Saez, L. & Young, M. W. PER–TIM interactions in living Drosophila cells: an interval timer for the circadian clock. Science 311, 226–229 (2006)
Doi, M., Hirayama, J. & Sassone-Corsi, P. Circadian regulator CLOCK is a histone acetyltransferase. Cell 125, 497–508 (2006)
Claridge-Chang, A. et al. Circadian regulation of gene expression systems in the Drosophila head. Neuron 32, 657–671 (2001)
Greenspan, R. J. & Ferveur, J. F. Courtship in Drosophila . Annu. Rev. Genet. 34, 205–232 (2000)
Hotta, Y. & Benzer, S. Courtship in Drosophila mosaics: sex-specific foci for sequential action patterns. Proc. Natl Acad. Sci. USA 73, 4154–4158 (1976)
Hall, J. C. Control of male reproductive behavior by the central nervous system of Drosophila: dissection of a courtship pathway by genetic mosaics. Genetics 92, 437–457 (1979)
Hall, J. C. Courtship among males due to a male-sterile mutation in Drosophila melanogaster . Behav. Genet. 8, 124–141 (1978)
Ito, H. et al. Sexual orientation in Drosophila is altered by the satori mutation in the sex-determination gene fruitless that encodes a zinc finger protein with a BTB domain. Proc. Natl Acad. Sci. USA 93, 9687–9692 (1996)
Ryner, L. C. et al. Control of male sexual behavior and sexual orientation in Drosophila by the fruitless gene. Cell 87, 1079–1089 (1996)
Demir, E. & Dickson, B. J. fruitless splicing specifies male courtship behavior in Drosophila . Cell 121, 785–794 (2005)
Kimura, K., Ote, M., Tazawa, T. & Yamamoto, D. Fruitless specifies sexually dimorphic neural circuitry in the Drosophila brain. Nature 438, 229–233 (2005)
Manoli, D. S. et al. Male-specific fruitless specifies the neural substrates of Drosophila courtship behaviour. Nature 436, 395–400 (2005)
Stockinger, P., Kvitsiani, D., Rotkopf, S., Tirian, L. & Dickson, B. J. Neural circuitry that governs Drosophila male courtship behavior. Cell 121, 795–807 (2005)
Kimchi, T., Xu, J. & Dulac, C. A functional circuit underlying male sexual behaviour in the female mouse brain. Nature 448, 1009–1014 (2007)
Ayyub, C., Paranjape, J., Rodrigues, V. & Siddiqi, O. Genetics of olfactory behavior in Drosophila melanogaster . J. Neurogenet. 6, 243–262 (1990)
Carlson, J. Olfaction in Drosophila: genetic and molecular analysis. Trends Neurosci. 14, 520–524 (1991)
Clyne, P. J. et al. The odor specificities of a subset of olfactory receptor neurons are governed by Acj6, a POU-domain transcription factor. Neuron 22, 339–347 (1999)
Vosshall, L. B. & Stocker, R. F. Molecular architecture of smell and taste in Drosophila . Annu. Rev. Neurosci. 30, 505–533 (2007)
Couto, A., Alenius, M. & Dickson, B. J. Molecular, anatomical, and functional organization of the Drosophila olfactory system. Curr. Biol. 15, 1535–1547 (2005)
Fishilevich, E. & Vosshall, L. B. Genetic and functional subdivision of the Drosophila antennal lobe. Curr. Biol. 15, 1548–1553 (2005)
Lin, H. H., Lai, J. S., Chin, A. L., Chen, Y. C. & Chiang, A. S. A map of olfactory representation in the Drosophila mushroom body. Cell 128, 1205–1217 (2007)
Jefferis, G. S. et al. Comprehensive maps of Drosophila higher olfactory centers: spatially segregated fruit and pheromone representation. Cell 128, 1187–1203 (2007)
Hallem, E. A. & Carlson, J. R. Coding of odors by a receptor repertoire. Cell 125, 143–160 (2006)
van der Goes van Naters. W & Carlson, J. R. Receptors and neurons for fly odors in Drosophila . Curr. Biol. 17, 606–612 (2007)
Kurtovic, A., Widmer, A. & Dickson, B. J. A single class of olfactory neurons mediates behavioural responses to a Drosophila sex pheromone. Nature 446, 542–546 (2007)
Quinn, W. G., Harris, W. A. & Benzer, S. Conditioned behavior in Drosophila melanogaster . Proc. Natl Acad. Sci. USA 71, 708–712 (1974)
Davis, R. L. Olfactory memory formation in Drosophila: from molecular to systems neuroscience. Annu. Rev. Neurosci. 28, 275–302 (2005)
Bourtchuladze, R. et al. Deficient long-term memory in mice with a targeted mutation of the cAMP-responsive element-binding protein. Cell 79, 59–68 (1994)
Schacher, S., Castellucci, V. F. & Kandel, E. R. cAMP evokes long-term facilitation in Aplysia sensory neurons that requires new protein synthesis. Science 240, 1667–1669 (1988)
Dubnau, J. et al. The staufen/pumilio pathway is involved in Drosophila long-term memory. Curr. Biol. 13, 286–296 (2003)
Byers, D., Davis, R. L. & Kiger, J. A. Defect in cyclic AMP phosphodiesterase due to the dunce mutation of learning in Drosophila melanogaster . Nature 289, 79–81 (1981)
Nighorn, A., Healy, M. J. & Davis, R. L. The cyclic AMP phosphodiesterase encoded by the Drosophila dunce gene is concentrated in the mushroom body neuropil. Neuron 6, 455–467 (1991)
Heisenberg, M., Borst, A., Wagner, S. & Byers, D. Drosophila mushroom body mutants are deficient in olfactory learning. J. Neurogenet. 2, 1–30 (1985)
Han, P. L., Levin, L. R., Reed, R. R. & Davis, R. L. Preferential expression of the Drosophila rutabaga gene in mushroom bodies, neural centers for learning in insects. Neuron 9, 619–627 (1992)
Yu, D., Keene, A. C., Srivatsan, A., Waddell, S. & Davis, R. L. Drosophila DPM neurons form a delayed and branch-specific memory trace after olfactory classical conditioning. Cell 123, 945–957 (2005)
Yu, D., Ponomarev, A. & Davis, R. L. Altered representation of the spatial code for odors after olfactory classical conditioning; memory trace formation by synaptic recruitment. Neuron 42, 437–449 (2004)
Krashes, M. J., Keene, A. C., Leung, B., Armstrong, J. D. & Waddell, S. Sequential use of mushroom body neuron subsets during Drosophila odor memory processing. Neuron 53, 103–115 (2007)
Liu, G. et al. Distinct memory traces for two visual features in the Drosophila brain. Nature 439, 551–556 (2006)
Wilson, R. I., Turner, G. C. & Laurent, G. Transformation of olfactory representations in the Drosophila antennal lobe. Science 303, 366–370 (2004)
Wolf, F. W., Rodan, A. R., Tsai, L. T. & Heberlein, U. High-resolution analysis of ethanol-induced locomotor stimulation in Drosophila . J. Neurosci. 22, 11035–11044 (2002)
Moore, M. S. et al. Ethanol intoxication in Drosophila: Genetic and pharmacological evidence for regulation by the cAMP signaling pathway. Cell 93, 997–1007 (1998)
Chen, S., Lee, A. Y., Bowens, N. M., Huber, R. & Kravitz, E. A. Fighting fruit flies: a model system for the study of aggression. Proc. Natl Acad. Sci. USA 99, 5664–5668 (2002)
Vrontou, E., Nilsen, S. P., Demir, E., Kravitz, E. A. & Dickson, B. J. fruitless regulates aggression and dominance in Drosophila . Nature Neurosci. 9, 1469–1471 (2006)
Dierick, H. A. & Greenspan, R. J. Molecular analysis of flies selected for aggressive behavior. Nature Genet. 38, 1023–1031 (2006)
Pick, S. & Strauss, R. Goal-driven behavioral adaptations in gap-climbing Drosophila . Curr. Biol. 15, 1473–1478 (2005)
Levine, J. D., Funes, P., Dowse, H. B. & Hall, J. C. Resetting the circadian clock by social experience in Drosophila melanogaster . Science 298, 2010–2012 (2002)
Fujii, S., Krishnan, P., Hardin, P. & Amrein, H. Nocturnal male sex drive in Drosophila . Curr. Biol. 17, 244–251 (2007)
Suh, G. S. et al. A single population of olfactory sensory neurons mediates an innate avoidance behaviour in Drosophila . Nature 431, 854–859 (2004)
Dekker, T., Ibba, I., Siju, K. P., Stensmyr, M. C. & Hansson, B. S. Olfactory shifts parallel superspecialism for toxic fruit in Drosophila melanogaster sibling, D. sechellia . Curr. Biol. 16, 101–109 (2006)
Tully, T. & Quinn, W. G. Classical conditioning and retention in normal and mutant Drosophila melanogaster . J. Comp. Physiol. 157, 263–277 (1985)
Gerber, B., Tanimoto, H. & Heisenberg, M. Classical conditioning and retention in normal and mutant Drosophila melanogaster . Curr. Opin. Neurobiol. 14, 737–744 (2004)
Weiner, J. Time, Love, Memory: A Great Biologist and His Quest for the Origins of Behavior (Alfred A. Knopf, Inc., New York, 1999)
Acknowledgements
Work in the author’s laboratory is supported by the NIH, FNIH and the Gates Foundation, and the Irma T. Hirschl Trust. I apologise to the many excellent Drosophila neurogeneticists whose work was not discussed here due to space constraints.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Rights and permissions
About this article
Cite this article
Vosshall, L. Into the mind of a fly. Nature 450, 193–197 (2007). https://doi.org/10.1038/nature06335
Issue Date:
DOI: https://doi.org/10.1038/nature06335
This article is cited by
-
The Divider Assay is a high-throughput pipeline for aggression analysis in Drosophila
Communications Biology (2021)
-
A coupled core-mantle evolution: review and future prospects
Progress in Earth and Planetary Science (2020)
-
The making of a pest: Insights from the evolution of chemosensory receptor families in a pestiferous and invasive fly, Drosophila suzukii
BMC Genomics (2016)
-
Volatile codes: Correlation of olfactory signals and reception in Drosophila-yeast chemical communication
Scientific Reports (2015)
-
How deeply does your mutant sleep? Probing arousal to better understand sleep defects in Drosophila
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.