Abstract
Huntington’s disease (HD) is a dominantly inherited neurodegenerative disorder caused by expansion of CAG triplet repeats in the huntingtin (HTT) gene (also called HD) and characterized by accumulation of aggregated fragments of polyglutamine-expanded HTT protein in affected neurons1,2. Abnormal enrichment of HD inclusion bodies with ubiquitin, a diagnostic characteristic of HD and many other neurodegenerative disorders including Alzheimer’s and Parkinson’s diseases3,4, has suggested that dysfunction in ubiquitin metabolism may contribute to the pathogenesis of these diseases5,6. Because modification of proteins with polyubiquitin chains regulates many essential cellular processes including protein degradation, cell cycle, transcription, DNA repair and membrane trafficking7, disrupted ubiquitin signalling is likely to have broad consequences for neuronal function and survival. Although ubiquitin-dependent protein degradation is impaired in cell-culture models of HD8,9,10,11 and of other neurodegenerative diseases12,13, it has not been possible to evaluate the function of the ubiquitin–proteasome system (UPS) in HD patients or in animal models of the disease, and a functional role for UPS impairment in neurodegenerative disease pathogenesis remains controversial14,15,16. Here we exploit a mass-spectrometry-based method to quantify polyubiquitin chains17 and demonstrate that the abundance of these chains is a faithful endogenous biomarker of UPS function. Lys 48-linked polyubiquitin chains accumulate early in pathogenesis in brains from the R6/2 transgenic mouse model of HD, from a knock-in model of HD and from human HD patients, establishing that UPS dysfunction is a consistent feature of HD pathology. Lys 63- and Lys 11-linked polyubiquitin chains, which are not typically associated with proteasomal targeting, also accumulate in the R6/2 mouse brain. Thus, HD is linked to global changes in the ubiquitin system to a much greater extent than previously recognized.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
MacDonald, M. E. et al. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 72, 971–983 (1993)
Tobin, A. J. & Signer, E. R. Huntington’s disease: the challenge for cell biologists. Trends Cell Biol. 10, 531–536 (2000)
Lowe, J. et al. Ubiquitin is a common factor in intermediate filament inclusion bodies of diverse type in man, including those of Parkinson’s disease, Pick’s disease, and Alzheimer’s disease, as well as Rosenthal fibres in cerebellar astrocytomas, cytoplasmic bodies in muscle, and mallory bodies in alcoholic liver disease. J. Pathol. 155, 9–15 (1988)
Mayer, R. J., Lowe, J., Lennox, G., Doherty, F. & Landon, M. Intermediate filaments and ubiquitin: a new thread in the understanding of chronic neurodegenerative diseases. Prog. Clin. Biol. Res. 317, 809–818 (1989)
DiFiglia, M. et al. Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science 277, 1990–1993 (1997)
Ross, C. A. & Pickart, C. M. The ubiquitin–proteasome pathway in Parkinson’s disease and other neurodegenerative diseases. Trends Cell Biol. 14, 703–711 (2004)
Pickart, C. M. & Fushman, D. Polyubiquitin chains: polymeric protein signals. Curr. Opin. Chem. Biol. 8, 610–616 (2004)
Bence, N. F., Sampat, R. M. & Kopito, R. R. Impairment of the ubiquitin–proteasome system by protein aggregation. Science 292, 1552–1555 (2001)
Bennett, E. J., Bence, N. F., Jayakumar, R. & Kopito, R. R. Global impairment of the ubiquitin–proteasome system by nuclear or cytoplasmic protein aggregates precedes inclusion body formation. Mol. Cell 17, 351–365 (2005)
Jana, N. R., Zemskov, E. A., Wang, G. & Nukina, N. Altered proteasomal function due to the expression of polyglutamine-expanded truncated N-terminal huntingtin induces apoptosis by caspase activation through mitochondrial cytochrome c release. Hum. Mol. Genet. 10, 1049–1059 (2001)
Verhoef, L. G., Lindsten, K., Masucci, M. G. & Dantuma, N. P. Aggregate formation inhibits proteasomal degradation of polyglutamine proteins. Hum. Mol. Genet. 11, 2689–2700 (2002)
Petrucelli, L. et al. Parkin protects against the toxicity associated with mutant α-synuclein: proteasome dysfunction selectively affects catecholaminergic neurons. Neuron 36, 1007–1019 (2002)
Urushitani, M., Kurisu, J., Tsukita, K. & Takahashi, R. Proteasomal inhibition by misfolded mutant superoxide dismutase 1 induces selective motor neuron death in familial amyotrophic lateral sclerosis. J. Neurochem. 83, 1030–1042 (2002)
Wang, H. L. et al. Polyglutamine-expanded ataxin-7 decreases nuclear translocation of NF-κB p65 and impairs NF-κB activity by inhibiting proteasome activity of cerebellar neurons. Cell Signal. 19, 573–581 (2007)
Bowman, A. B., Yoo, S. Y., Dantuma, N. P. & Zoghbi, H. Y. Neuronal dysfunction in a polyglutamine disease model occurs in the absence of ubiquitin–proteasome system impairment and inversely correlates with the degree of nuclear inclusion formation. Hum. Mol. Genet. 14, 679–691 (2005)
Kabashi, E., Agar, J. N., Taylor, D. M., Minotti, S. & Durham, H. D. Focal dysfunction of the proteasome: a pathogenic factor in a mouse model of amyotrophic lateral sclerosis. J. Neurochem. 89, 1325–1335 (2004)
Kirkpatrick, D. S. et al. Quantitative analysis of in vitro ubiquitinated cyclin B1 reveals complex chain topology. Nature Cell Biol. 8, 700–710 (2006)
Raasi, S., Varadan, R., Fushman, D. & Pickart, C. M. Diverse polyubiquitin interaction properties of ubiquitin-associated domains. Nature Struct. Mol. Biol. 12, 708–714 (2005)
Barr, J. R. et al. Isotope dilution–mass spectrometric quantification of specific proteins: model application with apolipoprotein A-I. Clin. Chem. 42, 1676–1682 (1996)
Dass, C., Fridland, G. H., Tinsley, P. W., Killmar, J. T. & Desiderio, D. M. Characterization of β-endorphin in human pituitary by fast atom bombardment mass spectrometry of trypsin-generated fragments. Int. J. Pept. Protein Res. 34, 81–87 (1989)
Baker, R. T. et al. Using deubiquitylating enzymes as research tools. Methods Enzymol. 398, 540–554 (2005)
Lindsten, K., Menendez-Benito, V., Masucci, M. G. & Dantuma, N. P. A transgenic mouse model of the ubiquitin/proteasome system. Nature Biotechnol. 21, 897–902 (2003)
Mangiarini, L. et al. Exon 1 of the HD gene with an expanded CAG repeat is sufficient to cause a progressive neurological phenotype in transgenic mice. Cell 87, 493–506 (1996)
Lin, C. H. et al. Neurological abnormalities in a knock-in mouse model of Huntington’s disease. Hum. Mol. Genet. 10, 137–144 (2001)
Woodman, B. et al. The HdhQ150/Q150 knock-in mouse model of HD and the R6/2 exon 1 model develop comparable and widespread molecular phenotypes. Brain Res. Bull. 72, 83–97 (2007)
Hanna, J. et al. Deubiquitinating enzyme ubp6 functions noncatalytically to delay proteasomal degradation. Cell 127, 99–111 (2006)
Kisselev, A. F. & Goldberg, A. L. Monitoring activity and inhibition of 26S proteasomes with fluorogenic peptide substrates. Methods Enzymol. 398, 364–378 (2005)
Ryu, K. Y., Baker, R. T. & Kopito, R. R. Ubiquitin-specific protease 2 as a tool for quantification of total ubiquitin levels in biological specimens. Anal. Biochem. 353, 153–155 (2006)
Acknowledgements
We are grateful to P. Howley, R. Baker and N. Nukina for reagents. We thank D. Kirkpatrick and N. Hathaway for suggestions, and J.-P. Vonsattel and the New York Brain Bank for the human brain tissue. This work was supported by a predoctoral training grant from NIGMS (E.J.B.), a small business innovation research grant from NINDS (H.S.), grants from the Huntington’s Disease Society of America Coalition for the Cure, Hereditary Disease Foundation and High Q Foundation (G.P.B. and R.R.K.), and a grant from the Wellcome Trust (G.P.B.).
Author Contributions E.J.B., T.A.S., C.H.B., H.S. and R.R.K. devised the overall proteomic approach. E.J.B. performed all of the biochemical analyses, the pull-down assays and, together with T.A.S., obtained and analysed all the mass spectrometry data. All of the mouse breeding and dissection was performed by B.W. and G.P.B. T.S.Z. performed all experiments in Supplementary Fig. 2 and the analysis of the HdhQ150/Q150 knock-in mice. K.-Y.R. contributed the real-time RT–PCR data in Supplementary Fig. 3 and performed the ubiquitin ELISA on R6/2 and control mice. E.J.B. and R.R.K. wrote the manuscript. All authors discussed the results and contributed to the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
H.S., T.A.S. and C.H.B. are employees of PPD Biomarker Discovery Inc.
Supplementary information
Supplementary Information
This file contains Supplementary Discussion, Supplementary Figures S1-S7 with Legends and Supplementary Table S1. (PDF 377 kb)
Rights and permissions
About this article
Cite this article
Bennett, E., Shaler, T., Woodman, B. et al. Global changes to the ubiquitin system in Huntington's disease. Nature 448, 704–708 (2007). https://doi.org/10.1038/nature06022
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06022
This article is cited by
-
A clinical study and future prospects for bioactive compounds and semi-synthetic molecules in the therapies for Huntington's disease
Molecular Neurobiology (2024)
-
Deubiquitinating enzymes (DUBs): decipher underlying basis of neurodegenerative diseases
Molecular Psychiatry (2022)
-
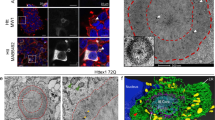
Nuclear and cytoplasmic huntingtin inclusions exhibit distinct biochemical composition, interactome and ultrastructural properties
Nature Communications (2021)
-
Withaferin A Induces Heat Shock Response and Ameliorates Disease Progression in a Mouse Model of Huntington’s Disease
Molecular Neurobiology (2021)
-
Inhibition of p38 Mitogen–Activated Protein Kinase Ameliorates HAP40 Depletion–Induced Toxicity and Proteasomal Defect in Huntington’s Disease Model
Molecular Neurobiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.