Abstract
Jasmonates are essential phytohormones for plant development and survival. However, the molecular details of their signalling pathway remain largely unknown. The identification more than a decade ago of COI1 as an F-box protein suggested the existence of a repressor of jasmonate responses that is targeted by the SCFCOI1 complex for proteasome degradation in response to jasmonate. Here we report the identification of JASMONATE-INSENSITIVE 3 (JAI3) and a family of related proteins named JAZ (jasmonate ZIM-domain), in Arabidopsis thaliana. Our results demonstrate that JAI3 and other JAZs are direct targets of the SCFCOI1 E3 ubiquitin ligase and jasmonate treatment induces their proteasome degradation. Moreover, JAI3 negatively regulates the key transcriptional activator of jasmonate responses, MYC2. The JAZ family therefore represents the molecular link between the two previously known steps in the jasmonate pathway. Furthermore, we demonstrate the existence of a regulatory feed-back loop involving MYC2 and JAZ proteins, which provides a mechanistic explanation for the pulsed response to jasmonate and the subsequent desensitization of the cell.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Devoto, A. & Turner, J. G. Regulation of jasmonate-mediated plant responses in Arabidopsis. Ann. Bot. (Lond.) 92, 329–337 (2003)
Farmer, E. E., Almeras, E. & Krishnamurthym, V. Jasmonates and related oxylipins in plant responses to pathogenesis and herbivory. Curr. Opin. Plant Biol. 6, 372–378 (2003)
Rojo, E., Solano, R. & Sánchez-Serrano, J. J. Interactions between signaling compounds involved in plant defense. J. Plant Growth Regul. 22, 82–98 (2003)
Lorenzo, O. & Solano, R. Molecular players regulating the jasmonate signalling network. Curr. Opin. Plant Biol. 8, 532–540 (2005)
Flescher, E. Jasmonates-a new family of anti-cancer agents. Anticancer Drugs 16, 911–916 (2005)
Demole, E., Lederer, E. & Mercier, D. Isolement et determination de la structure du jasmonate de methyle, constituant odorant characteristique de l'essence de jasmin. Helv. Chim. Acta XLV, 675–685 (1962)
Liechti, R. & Farmer, E. E. Jasmonate biochemical pathway. Sci. STKE 203, CM18 (2006)
Liechti, R., Gfeller, A. & Farmer, E. E. Jasmonate signaling pathway. Sci. STKE 322, CM2 (2006)
Devoto, A. et al. COI1 links jasmonate signalling and fertility to the SCF ubiquitin-ligase complex in Arabidopsis. Plant J. 32, 457–466 (2002)
Feng, S. et al. The COP9 signalosome interacts physically with SCF COI1 and modulates jasmonate responses. Plant Cell 15, 1083–1094 (2003)
Feys, B., Benedetti, C. E., Penfold, C. N. & Turner, J. G. Arabidopsis mutants selected for resistance to the phytotoxin coronatine are male sterile, insensitive to methyl jasmonate, and resistant to a bacterial pathogen. Plant Cell 6, 751–759 (1994)
Xie, D. X., Feys, B. F., James, S., Nieto-Rostro, M. & Turner, J. G. COI1: an Arabidopsis gene required for jasmonate-regulated defense and fertility. Science 280, 1091–1094 (1998)
Xu, L. et al. The SCF(COI1) ubiquitin-ligase complexes are required for jasmonate response in Arabidopsis. Plant Cell 14, 1919–1935 (2002)
Boter, M., Ruiz-Rivero, O., Abdeen, A. & Prat, S. Conserved MYC transcription factors play a key role in jasmonate signaling both in tomato and Arabidopsis. Genes Dev. 18, 1577–1591 (2004)
Lorenzo, O., Chico, J. M., Sánchez-Serrano, J. J. & Solano, R. JASMONATE-INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell 16, 1938–1950 (2004)
Lorenzo, O., Piqueras, R., Sánchez-Serrano, J. J. & Solano, R. ETHYLENE RESPONSE FACTOR1 integrates signals from ethylene and jasmonate pathways in plant defense. Plant Cell 15, 165–178 (2003)
Nishii, A. et al. Characterization of a novel gene encoding a putative single zinc-finger protein, ZIM, expressed during the reproductive phase in Arabidopsis thaliana. Biosci. Biotechnol. Biochem. 64, 1402–1409 (2000)
Shikata, M. et al. Characterization of Arabidopsis ZIM, a member of a novel plant-specific GATA factor gene family. J. Exp. Bot. 55, 631–639 (2004)
Datta, S., Hettiarachchi, G. H. C. M., Deng, X.-W. & Holm, M. Arabidopsis CONSTANS-LIKE3 is a positive regulator of red light signaling and root growth. Plant Cell 18, 70–84 (2006)
Robson, F. et al. Functional importance of conserved domains in the flowering-time gene CONSTANS demonstrated by analysis of mutant alleles and transgenic plants. Plant J. 28, 619–631 (2001)
Staswick, P. E. & Tiryaki, I. The oxylipin signal jasmonic acid is activated by an enzyme that conjugates it to isoleucine in Arabidopsis. Plant Cell 16, 2117–2127 (2004)
Thines, B. et al. JAZ repressor proteins are targets of the SCFCOI1 complex during jasmonate signalling. Nature advance online publication. doi: 10.1038/nature05960 (this issue)
Färber, K., Schumann, B., Miersch, O. & Roos, W. Selective desensitization of jasmonate- and pH-dependent signaling in the induction of benzophenanthridine biosynthesis in cells of Eschscholzia californica. Phytochemistry 62, 491–500 (2003)
Dharmasiri, N., Dharmasiri, S. & Estelle, M. The F-box protein TIR1 is an auxin receptor. Nature 435, 441–445 (2005)
Kepinski, S. & Leyser, O. The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 435, 446–451 (2005)
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007)
Weiler, E. W. et al. The Pseudomonas phytotoxin coronatine mimics octadecanoid signalling molecules of higher plants. FEBS Lett. 345, 9–13 (1994)
Axel, M., Mathias, M. & Wilhelm, B. Structural and biological diversity of cyclic octadecanoids, jasmonates, and mimetics. J. Plant Growth Regul. 23, 170–178 (2004)
Stintzi, A., Weber, H., Reymond, P., Browse, J. & Farmer, E. E. Plant defense in the absence of jasmonic acid: the role of cyclopentenones. Proc. Natl Acad. Sci. USA 98, 12837–12842 (2001)
Taki, N. et al. 12-oxo-phytodienoic acid triggers expression of a distinct set of genes and plays a role in wound-induced gene expression in Arabidopsis. Plant Physiol. 139, 1268–1283 (2005)
Uppalapati, S. R. et al. The phytotoxin coronatine and methyl jasmonate impact multiple phytohormone pathways in tomato. Plant J. 42, 201–217 (2005)
Mita, S., Suzuki-Fujii, K. & Nakamura, K. Sugar-inducible expression of a gene for beta-amylase in Arabidopsis thaliana. Plant Physiol. 107, 895–904 (1995)
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998)
Hammarstrom, M., Hellgren, N., van Den Berg, S., Berglund, H. & Hard, T. Rapid screening for improved solubility of small human proteins produced as fusion proteins in Escherichia coli.. Protein Sci. 11, 313–321 (2002)
Schiestl, R. H. & Gietz, R. D. High efficiency transformation of intact yeast cells using single stranded nucleic acids as a carrier. Curr. Genet. 16, 339–346 (1989)
Adie, B. A. T. et al. ABA is an essential signal for plant resistance to pathogens affecting JA biosynthesis and the activation of defenses in Arabidopsis. Plant Cell 19, 1665–1681 (2007)
Solano, R., Stepanova, A., Chao, Q. & Ecker, J. R. Nuclear events in ethylene signaling: a transcriptional cascade mediated by ETHYLENE-INSENSITIVE3 and ETHYLENE-RESPONSE-FACTOR1. Genes Dev. 12, 3703–3714 (1998)
Thijs, G. et al. A higher-order background model improves the detection of promoter regulatory elements by Gibbs sampling. Bioinformatics 17, 1113–1122 (2001)
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003)
Deblaere, R. et al. Efficient octopine Ti plasmid-derived vectors for Agrobacterium-mediated gene transfer to plants. Nucleic Acids Res. 13, 4777–4788 (1985)
Valli, A., Martin-Hernandez, A. M., Lopez-Moya, J. J. & Garcia, J. A. RNA silencing suppression by a second copy of the P1 serine protease of Cucumber vein yellowing ipomovirus, a member of the family Potyviridae that lacks the cysteine protease HCPro. J. Virol. 80, 10055–10063 (2006)
Ponce, M. R., Robles, P. & Micol, J. L. High-throughput genetic mapping in Arabidopsis thaliana. Mol. Gen. Genet. 261, 408–415 (1999)
Ponce, M. R., Robles, P., Lozano, F. M., Brotons, M. A. & Micol, J. L. Low-resolution mapping of untagged mutations. Methods Mol. Biol. 323, 105–113 (2006)
Bell, C. J. & Ecker, J. R. Assignment of 30 microsatellite loci to the linkage map of Arabidopsis. Genomics 19, 137–144 (1994)
Benjamani, Y. & Hochberg, Y. Controlling the false discovery rate. J. R. Stat. Soc. Ser. C Appl. Stat. 57, 289–300 (1995)
Saeed, A. I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003)
Chini, A. & Loake, G. J. Motifs specific for the ADR1 NBS-LRR protein family in Arabidopsis are conserved among NBS-LRR sequences from both dicotyledonous and monocotyledonous plants. Planta 4, 597–601 (2005)
Acknowledgements
We thank C. Castresana, J. A. García, J. Paz-Ares and S. Prat for critical reading of the manuscript and V. Rubio , J. C. del Pozo and J. M. Iglesias-Pedraz for advice on pull-down and proteasome degradation experiments. We also thank M. Laos and S. Gutiérrez for technical assistance. We are grateful to B. Thines and J. Browse for sharing results before publication. We also thank J. Turner for providing coi1-1 seeds, P. Staswick for jar1 and NASC for T-DNA insertions lines and full-length cDNAs. This work was supported by funding from the Ministerio de Educación y Ciencia of Spain (to R.S., J.L.M. and M.R.P.), and the Comunidad de Madrid and European Comission (to R.S.). A.C. was supported by an EMBO long-term Fellowship, and S.F. by a Portuguese FCT fellowship.
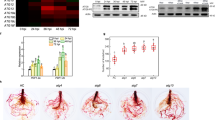
Author Contributions A.C. was responsible for experiments of Fig. 1b–h and Fig. 2c–i, S.F. performed all pull-down experiments, G.F., in vitro degradation experiments and EMSA, B.A., two-hybrid assays, J.M.C., G.G.-C. and I.L.-V., microarray analysis and A.C., O.L., F.M.L., J.L.M. and M.R.P. mapped jai3-1. R.S. designed experiments, supervised the work and wrote the manuscript. All authors discussed the results and commented on the manuscript.
MIAMExpress Accession numbers for all microarray experiments are: E-ATMX-15, E-ATMX-16, E-ATMX-17 and E-ATMX-18. Accession numbers for JAZ family members are: JAZ1, At1g19180; JAZ2, At1g74950; JAI3(JAZ3), At3g17860; JAZ4, At1g48500; JAZ5, At1g17380; JAZ6, At1g72450; JAZ7, At2g34600; JAZ8, At1g30135; JAZ9, At1g70700; JAZ10, At5g13220; JAZ11, At3g43440; and JAZ12, At5g20900.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S7 with Legends and Supplementary Tables S1-S3. (PDF 1990 kb)
Rights and permissions
About this article
Cite this article
Chini, A., Fonseca, S., Fernández, G. et al. The JAZ family of repressors is the missing link in jasmonate signalling. Nature 448, 666–671 (2007). https://doi.org/10.1038/nature06006
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06006
This article is cited by
-
Petal abscission is promoted by jasmonic acid-induced autophagy at Arabidopsis petal bases
Nature Communications (2024)
-
LiCOI1 mediates the biosynthesis of monoterpenes induced by methyl jasmonate in Lilium ‘Siberia’
Horticulture, Environment, and Biotechnology (2024)
-
Temporal profiling of physiological, histological, and transcriptomic dissection during auxin-induced adventitious root formation in tetraploid Robinia pseudoacacia micro-cuttings
Planta (2024)
-
The N-terminal α2 helix element is critical for the activity of the rice transcription factor MYC2
Plant Molecular Biology (2024)
-
Genome-wide identification of the TIFY family reveals JAZ subfamily function in response to hormone treatment in Betula platyphylla
BMC Plant Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.