Abstract
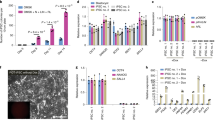
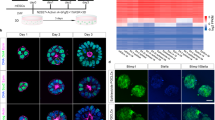
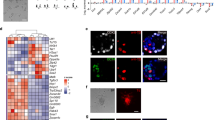
Although the first mouse embryonic stem (ES) cell lines were derived 25 years ago1,2 using feeder-layer-based blastocyst cultures, subsequent efforts to extend the approach to other mammals, including both laboratory and domestic species, have been relatively unsuccessful. The most notable exceptions were the derivation of non-human primate ES cell lines3 followed shortly thereafter by their derivation of human ES cells4. Despite the apparent common origin and the similar pluripotency of mouse and human embryonic stem cells, recent studies have revealed that they use different signalling pathways to maintain their pluripotent status. Mouse ES cells depend on leukaemia inhibitory factor and bone morphogenetic protein, whereas their human counterparts rely on activin (INHBA)/nodal (NODAL) and fibroblast growth factor (FGF). Here we show that pluripotent stem cells can be derived from the late epiblast layer of post-implantation mouse and rat embryos using chemically defined, activin-containing culture medium that is sufficient for long-term maintenance of human embryonic stem cells. Our results demonstrate that activin/Nodal signalling has an evolutionarily conserved role in the derivation and the maintenance of pluripotency in these novel stem cells. Epiblast stem cells provide a valuable experimental system for determining whether distinctions between mouse and human embryonic stem cells reflect species differences or diverse temporal origins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981)
Martin, G. R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc. Natl Acad. Sci. USA 78, 7634–7638 (1981)
Thomson, J. A. et al. Isolation of a primate embryonic stem cell line. Proc. Natl Acad. Sci. USA 92, 7844–7848 (1995)
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998)
Johansson, B. M. & Wiles, M. V. Evidence for involvement of activin A and bone morphogenetic protein 4 in mammalian mesoderm and hematopoietic development. Mol. Cell. Biol. 15, 141–151 (1995)
Brook, F. A. et al. The derivation of highly germline-competent embryonic stem cells containing NOD-derived genome. Diabetes 52, 205–208 (2003)
Vallier, L., Alexander, M. & Pedersen, R. A. Activin/Nodal and FGF pathways cooperate to maintain pluripotency of human embryonic stem cells. J. Cell Sci. 118, 4495–4509 (2005)
Pelton, T. A., Sharma, S., Schulz, T. C., Rathjen, J. & Rathjen, P. D. Transient pluripotent cell populations during primitive ectoderm formation: correlation of in vivo and in vitro pluripotent cell development. J. Cell Sci. 115, 329–339 (2002)
Rathjen, J. et al. Formation of a primitive ectoderm like cell population, EPL cells, from ES cells in response to biologically derived factors. J. Cell Sci. 112, 601–612 (1999)
Resnick, J. L., Bixler, L. S., Cheng, L. & Donovan, P. J. Long-term proliferation of mouse primordial germ cells in culture. Nature 359, 550–551 (1992)
Matsui, Y., Zsebo, K. & Hogan, B. L. Derivation of pluripotential embryonic stem cells from murine primordial germ cells in culture. Cell 70, 841–847 (1992)
De Felici, M. & McLaren, A. Isolation of mouse primordial germ cells. Exp. Cell Res. 142, 476–482 (1982)
Xu, R. H. et al. BMP4 initiates human embryonic stem cell differentiation to trophoblast. Nature Biotechnol. 20, 1261–1264 (2002)
Camus, A., Perea-Gomez, A., Moreau, A. & Collignon, J. Absence of Nodal signaling promotes precocious neural differentiation in the mouse embryo. Dev. Biol. 295, 743–755 (2006)
Mesnard, D., Guzman-Ayala, M. & Constam, D. B. Nodal specifies embryonic visceral endoderm and sustains pluripotent cells in the epiblast before overt axial patterning. Development 133, 2497–2505 (2006)
Nichols, J., Chambers, I., Taga, T. & Smith, A. Physiological rationale for responsiveness of mouse embryonic stem cells to gp130 cytokines. Development 128, 2333–2339 (2001)
Conlon, F. L. et al. A primary requirement for nodal in the formation and maintenance of the primitive streak in the mouse. Development 120, 1919–1928 (1994)
Song, J. et al. The type II activin receptors are essential for egg cylinder growth, gastrulation, and rostral head development in mice. Dev. Biol. 213, 157–169 (1999)
Gu, Z. et al. The type I serine/threonine kinase receptor ActRIA (ALK2) is required for gastrulation of the mouse embryo. Development 126, 2551–2561 (1999)
James, D., Noggle, S. A., Swigut, T. & Brivanlou, A. H. Contribution of human embryonic stem cells to mouse blastocysts. Dev. Biol. 295, 90–102 (2006)
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005)
Loh, Y. H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nature Genet. 38, 431–440 (2006)
Rugg-Gunn, P. J., Ferguson-Smith, A. C. & Pedersen, R. A. Epigenetic status of human embryonic stem cells. Nature Genet. 37, 585–587 (2005)
Sun, B. W. et al. Temporal and parental-specific expression of imprinted genes in a newly derived Chinese human embryonic stem cell line and embryoid bodies. Hum. Mol. Genet. 15, 65–75 (2006)
Beddington, R. S. & Robertson, E. J. An assessment of the developmental potential of embryonic stem cells in the midgestation mouse embryo. Development 105, 733–737 (1989)
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nature Genet. 24, 372–376 (2000)
Acknowledgements
We thank A. McLaren for her support. We thank A. Smith for CGR8 cells, A. Nagy for R1 cells, P. Andrew for the SSEA-1 antibody, L. Wicker for access to NOD mice, and S. Thiru for advice in the teratoma analysis. This work was supported by an MRC International Appointments Initiative grant (R.A.P), MRC/Juvenile Diabetes Research Foundation Centre funding (R.A.P., I.G.M.B.), Remedi (I.G.M.B.), the Wellcome Trust Functional Genomics Initiative on Stem Cells (L.E.S, M.W.B.T.), the Addenbrookes National Institute for Health Research Biomedical Research Centre, and a Diabetes UK Career Development fellowship (L.V.). We dedicate this paper to the memory of our colleague Isabelle Bouhon.
Author Contributions L.V. conceived the experiment and did the molecular analysis; I.G.M.B. derived and cultured the EpiSCs; R.A.P. performed the epiblast dissections and together with I.G.M.B. carried out the aggregation, chimaera and clonal assays; L.E.S. and M.T. obtained the microarray data; P.R-G. performed the epigenetic analysis; B.S. performed the blastocyst immunosurgery and ICM cultures; S.M.C.d.S.L. did the Stella–GFP dissection and Blimp1 staining; S.K.H. provided PCR analysis of chimaeras; A.C. did karyotyping of human ES cell lines; L.A-R. carried out the teratoma work; L.V., R.A.P. and I.G.M.B. analysed the data and co-wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information 1
This file contains Supplementary Data, Supplementary Methods, Supplementary Figures 1-7 with Legends and additional references. (PDF 5757 kb)
Rights and permissions
About this article
Cite this article
Brons, I., Smithers, L., Trotter, M. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007). https://doi.org/10.1038/nature05950
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05950
This article is cited by
-
Establishment of African pygmy mouse induced pluripotent stem cells using defined doxycycline inducible transcription factors
Scientific Reports (2024)
-
Conservation and divergence of canonical and non-canonical imprinting in murids
Genome Biology (2023)
-
c-Jun as a one-way valve at the naive to primed interface
Cell & Bioscience (2023)
-
Modelling in vitro gametogenesis using induced pluripotent stem cells: a review
Cell Regeneration (2023)
-
A new era of stem cell and developmental biology: from blastoids to synthetic embryos and beyond
Experimental & Molecular Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.