Abstract
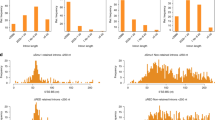
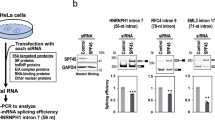
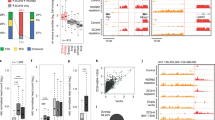
The human and mouse genomes share a number of long, perfectly conserved nucleotide sequences, termed ultraconserved elements1. Whereas these regions can act as transcriptional enhancers when upstream of genes, those within genes are less well understood. In particular, the function of ultraconserved elements that overlap alternatively spliced exons of genes encoding RNA-binding proteins is unknown1,2. Here we report that in every member of the human SR family of splicing regulators, highly or ultraconserved elements are alternatively spliced, either as alternative ‘poison cassette exons’ containing early in-frame stop codons, or as alternative introns in the 3′ untranslated region. These alternative splicing events target the resulting messenger RNAs for degradation by means of an RNA surveillance pathway called nonsense-mediated mRNA decay. Mouse orthologues of the human SR proteins exhibit the same unproductive splicing patterns. Three SR proteins have been previously shown to direct splicing of their own transcripts, and one of these is known to autoregulate its expression by coupling alternative splicing with decay3,4,5; our results suggest that unproductive splicing is important for regulation of the entire SR family. We find that unproductive splicing associated with conserved regions has arisen independently in different SR genes, suggesting that splicing factors may readily acquire this form of regulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Bejerano, G. et al. Ultraconserved elements in the human genome. Science 304, 1321–1325 (2004)
Bejerano, G. et al. A distal enhancer and an ultraconserved exon are derived from a novel retroposon. Nature 441, 87–90 (2006)
Jumaa, H. & Nielsen, P. J. The splicing factor SRp20 modifies splicing of its own mRNA and ASF/SF2 antagonizes this regulation. EMBO J. 16, 5077–5085 (1997)
Lejeune, F., Cavaloc, Y. & Stevenin, J. Alternative splicing of intron 3 of the serine/arginine-rich protein 9G8 gene. Identification of flanking exonic splicing enhancers and involvement of 9G8 as a trans-acting factor. J. Biol. Chem. 276, 7850–7858 (2001)
Sureau, A., Gattoni, R., Dooghe, Y., Stevenin, J. & Soret, J. SC35 autoregulates its expression by promoting splicing events that destabilize its mRNAs. EMBO J. 20, 1785–1796 (2001)
Sanford, J. R., Ellis, J. & Caceres, J. F. Multiple roles of arginine/serine-rich splicing factors in RNA processing. Biochem. Soc. Trans. 33, 443–446 (2005)
Nagy, E. & Maquat, L. E. A rule for termination-codon position within intron-containing genes: when nonsense affects RNA abundance. Trends Biochem. Sci. 23, 198–199 (1998)
Paillusson, A., Hirschi, N., Vallan, C., Azzalin, C. M. & Muhlemann, O. A GFP-based reporter system to monitor nonsense-mediated mRNA decay. Nucleic Acids Res. 33, e54 (2005)
Lareau, L. F., Green, R. E., Bhatnagar, R. S. & Brenner, S. E. The evolving roles of alternative splicing. Curr. Opin. Struct. Biol. 14, 273–282 (2004)
Ge, H., Zuo, P. & Manley, J. L. Primary structure of the human splicing factor ASF reveals similarities with Drosophila regulators. Cell 66, 373–382 (1991)
Cartegni, L., Wang, J., Zhu, Z., Zhang, M. Q. & Krainer, A. R. ESEfinder: A web resource to identify exonic splicing enhancers. Nucleic Acids Res. 31, 3568–3571 (2003)
Fairbrother, W. G., Yeh, R. F., Sharp, P. A. & Burge, C. B. Predictive identification of exonic splicing enhancers in human genes. Science 297, 1007–1013 (2002)
Hughes, J. R. et al. Annotation of cis-regulatory elements by identification, subclassification, and functional assessment of multispecies conserved sequences. Proc. Natl Acad. Sci. USA 102, 9830–9835 (2005)
Lewis, B. P., Green, R. E. & Brenner, S. E. Evidence for the widespread coupling of alternative splicing and nonsense-mediated mRNA decay in humans. Proc. Natl Acad. Sci. USA 100, 189–192 (2003)
Hillman, R. T., Green, R. E. & Brenner, S. E. An unappreciated role for RNA surveillance. Genome Biol. 5, R8 (2004)
Duncan, P. I., Stojdl, D. F., Marius, R. M. & Bell, J. C. In vivo regulation of alternative pre-mRNA splicing by the Clk1 protein kinase. Mol. Cell. Biol. 17, 5996–6001 (1997)
Garcia-Sacristan, A., Fernandez-Nestosa, M. J., Hernandez, P., Schvartzman, J. B. & Krimer, D. B. Protein kinase clk/STY is differentially regulated during erythroleukemia cell differentiation: a bias toward the skipped splice variant characterizes postcommitment stages. Cell Res. 15, 495–503 (2005)
Wollerton, M. C., Gooding, C., Wagner, E. J., Garcia-Blanco, M. A. & Smith, C. W. Autoregulation of polypyrimidine tract binding protein by alternative splicing leading to nonsense-mediated decay. Mol. Cell 13, 91–100 (2004)
Yeo, G. W., Van Nostrand, E., Holste, D., Poggio, T. & Burge, C. B. Identification and analysis of alternative splicing events conserved in human and mouse. Proc. Natl Acad. Sci. USA 102, 2850–2855 (2005)
Kumar, S. & Lopez, A. J. Negative feedback regulation among SR splicing factors encoded by Rbp1 and Rbp1-like in Drosophila. EMBO J. 24, 2646–2655 (2005)
Zorio, D. A., Lea, K. & Blumenthal, T. Cloning of Caenorhabditis U2AF65: an alternatively spliced RNA containing a novel exon. Mol. Cell. Biol. 17, 946–953 (1997)
Morrison, M., Harris, K. S. & Roth, M. B. smg mutants affect the expression of alternatively spliced SR protein mRNAs in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 94, 9782–9785 (1997)
Kalyna, M., Lopato, S., Voronin, V. & Barta, A. Evolutionary conservation and regulation of particular alternative splicing events in plant SR proteins. Nucleic Acids Res. 34, 4395–4405 (2006)
Iida, K. & Go, M. Survey of conserved alternative splicing events of mRNAs encoding SR proteins in land plants. Mol. Biol. Evol. 23, 1085–1094 (2006)
Sigal, A. et al. Variability and memory of protein levels in human cells. Nature 444, 643–646 (2006)
Pan, Q. et al. Quantitative microarray profiling provides evidence against widespread coupling of alternative splicing with nonsense-mediated mRNA decay to control gene expression. Genes Dev. 20, 153–158 (2006)
Wheeler, D. L. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 34, D173–D180 (2006)
Pearson, W. R. & Lipman, D. J. Improved tools for biological sequence comparison. Proc. Natl Acad. Sci. USA 85, 2444–2448 (1988)
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002)
John, B. et al. Human microRNA targets. PLoS Biol. 2, e363 (2004)
Acknowledgements
We thank present and past members of the Brenner laboratory and J. Pleiss for discussions and comments on the manuscript, Q. Meng and D. Rio for expertise, A. Fisher for assistance, and the Feldman, Luan, Rio and Tjian laboratories for use of their equipment. This work was supported by NIH grants and a Sloan fellowship to S.E.B. L.F.L. was supported by an NIH training grant. R.E.G. is supported by an NSF postdoctoral fellowship in Biological Informatics.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Competing Interests Statement: Our previous work in this area has resulted in three patent filings. US 7149631 ‘Alternative splicing and nonsense-mediated decay: computational methods and gene regulation’; application 10/637482 (pending) ‘RNA surveillance among curated proteins’; application 11/054155 (pending) ‘Nonsense suppressor agents in treatment of cutaneous and gastrointestinal disorders’. The first of these describes a computational method for identifying NMD-target isoforms and may cover the method used in this manuscript. The other patent filings discuss genes affected by alternative splicing and NMD, and therapeutic treatment. None of these patents mentions the specific results in this paper.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Results, Supplementary Figures S1-S5 with Legends, Supplementary Tables 1-5 and. additional references. (PDF 860 kb)
Rights and permissions
About this article
Cite this article
Lareau, L., Inada, M., Green, R. et al. Unproductive splicing of SR genes associated with highly conserved and ultraconserved DNA elements. Nature 446, 926–929 (2007). https://doi.org/10.1038/nature05676
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05676
This article is cited by
-
Shifts in isoform usage underlie transcriptional differences in regulatory T cells in type 1 diabetes
Communications Biology (2023)
-
TEQUILA-seq: a versatile and low-cost method for targeted long-read RNA sequencing
Nature Communications (2023)
-
RNA splicing dysregulation and the hallmarks of cancer
Nature Reviews Cancer (2023)
-
RNA editing: new roles in feedback and feedforward control
Cell Research (2023)
-
Plant serine/arginine-rich proteins: versatile players in RNA processing
Planta (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.