Abstract
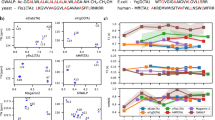
Kinetic data on a number of protein–protein associations have provided evidence for the initial formation of a pre-equilibrium encounter complex that subsequently relaxes to the final stereospecific complex1. Site-directed mutagenesis2,3,4 and brownian dynamics simulations5,6,7 have suggested that the rate of association can be modulated by perturbations in charge distribution outside the direct interaction surfaces. Furthermore, rate enhancement through non-specific binding may occur by either a reduction in dimensionality8 or the presence of a short-range, non-specific attractive potential9. Here, using paramagnetic relaxation enhancement, we directly demonstrate the existence and visualize the distribution of an ensemble of transient, non-specific encounter complexes under equilibrium conditions for a relatively weak protein–protein complex between the amino-terminal domain of enzyme I and the phosphocarrier protein HPr. Neither the stereospecific complex10 alone nor any single alternative conformation can account fully for the intermolecular paramagnetic relaxation enhancement data. Restrained rigid-body simulated annealing refinement against the paramagnetic relaxation enhancement data enables us to obtain an atomic probability distribution map of the non-specific encounter complex ensemble that qualitatively correlates with the electrostatic surface potentials on the interacting proteins. Qualitatively similar results are presented for two other protein–protein complexes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Fersht, A. R. Structure and Mechanism in Protein Science: a Guide to Enzyme Catalysis and Protein Folding (Freeman & Co, New York, 1999)
Schreiber, G. & Fersht, A. R. Rapid electrostatically assisted association of proteins. Nature Struct. Biol. 3, 427–431 (1996)
Vijaykumar, M. et al. Electrostatic enhancement of diffusion-controlled protein–protein association: comparison of theory and experiment on barnase and barstar. J. Mol. Biol. 278, 1015–1024 (1998)
Selzer, T., Albeck, S. & Schreiber, G. Rational design of faster associating and tighter binding protein complexes. Nature Struct. Biol. 7, 537–541 (2000)
Northrup, S. H., Boles, J. O. & Reynolds, J. C. L. Brownian dynamics of cytochrome c and cytochrome c peroxidase association. Science 241, 67–70 (1988)
Gabdoulline, R. R. & Wade, R. C. Biomolecular diffusional association. Curr. Opin. Struct. Biol. 12, 204–213 (2002)
Spaar, A., Dammer, C., Gabdoulline, R. R., Wade, R. C. & Helms, V. Diffusional encounter of barnase and barstar. Biophys. J. 90, 1913–1924 (2006)
Adams, G. & Debruck, M. in Structural Chemistry and Molecular Biology (eds Rich, A. & Davidson, N.) 198–215 (Freeman & Co, San Francisco, CA, 1968)
Zhou, H-X. & Szabo, A. Enhancement of association rates by nonspecific binding to DNA and cell membranes. Phys. Rev. Lett. 93, 178101 (2004)
Garrett, D. S., Seok, Y-J., Peterkofsky, A., Gronenborn, A. M. & Clore, G. M. Solution structure of the 40,000 M r phosphoryl transfer complex between the N-terminal domain of enzyme I and HPr. Nature Struct. Biol. 6, 166–173 (1999)
Postma, P. W., Lengeler, J. W. & Jacobson, G. R. in Escherichia coli and Salmonella: Cellular and Molecular Biology (ed. Neidhardt, F. C.) 1149–1174 (ASM, Washington DC, 1996)
Iwahara, J. & Clore, G. M. Detecting transient intermediates in macromolecular binding by paramagnetic NMR. Nature 440, 1227–1230 (2006)
Iwahara, J., Schwieters, C. D. & Clore, G. M. Characterization of nonspecific protein–DNA interactions by 1H paramagnetic relaxation enhancement. J. Am. Chem. Soc. 126, 12800–12808 (2004)
Ebright, Y. W., Chen, Y., Pendergrast, P. S. & Ebright, R. H. Incorporation of an EDTA–metal complex at a rationally selected site within a protein: application to EDTA–iron DNA affinity cleaving with catabolite gene activator protein (CAP) and Cro. Biochemistry 31, 10664–10670 (1992)
Iwahara, J., Schwieters, C. D. & Clore, G. M. Ensemble approach for NMR structure refinement against 1H paramagnetic relaxation enhancement data arising from a flexible paramagnetic group attached to a macromolecules. J. Am. Chem. Soc. 126, 5879–5896 (2004)
Donaldson, L. W. et al. Structural characterization of proteins with an attached ATCUN motif by paramagnetic relaxation enhancement NMR spectroscopy. J. Am. Chem. Soc. 123, 9843–9847 (2001)
Pintacuda, G. & Otting, G. Identification of protein surfaces by NMR measurements with a paramagnetic Gd(iii) chelate. J. Am. Chem. Soc. 124, 372–373 (2002)
Schwieters, C. D., Kuszewski, J. J. & Clore, G. M. Using Xplor-NIH for NMR molecular structure determination. Prog. NMR Spectrosc. 48, 47–62 (2006)
Brünger, A. T., Clore, G. M., Gronenborn, A. M. & Nilges, M. Assessing the quality of solution nuclear magnetic resonance structures by complete cross-validation. Science 261, 328–331 (1993)
Schwieters, C. D. & Clore, G. M. Reweighted atomic densities to represent ensembles of NMR structures. J. Biomol. NMR 23, 221–225 (2002)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Jones, S. & Thornton, J. M. principles of protein–protein interactions. Proc. Natl Acad. Sci. USA 93, 13–20 (1996)
Cornilescu, G. et al. Solution structure of the phosphoryl transfer complex between the cytoplasmic A domain of the mannitol transporter IIMannitol and HPr of the Eschrichia coli phosphotransferase system. J. Biol. Chem. 277, 42289–42298 (2002)
Williams, D. C., Cai, M., Suh, Y-J., Peterkofsky, A. & Clore, G. M. Solution NMR structure of the 48 kDa IIAMannose–HPr complex of the Escherichia coli mannose phosphotransferase system. J. Biol. Chem. 280, 20775–20784 (2005)
Nilges, M., Gronenborn, A. M., Brünger, A. T. & Clore, G. M. Determination of three-dimensional structures of proteins by simulated annealing with interproton distance restraints: application to crambin, potato carboxypeptidase inhibitor and barley serine proteinase inhibitor 2. Protein Eng. 2, 27–38 (1988)
Kuszewski, J., Gronenborn, A. M. & Clore, G. M. Improving the packing and accuracy of NMR structures with a pseudopotential for the radius of gyration. J. Am. Chem. Soc. 121, 2337–2338 (1999)
DeLano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific, San Carlos, CA, USA, 2002)
Laskowski, R. A. SURFNET: a program for visualizing molecular surfaces, cavities and intermolecular interactions. J. Mol. Graph. 13, 323–330 (1995)
Acknowledgements
We thank C. Schwieters, A. Szabo and C. Bewley for discussions; and C. Byeon for assistance with initial sample preparation. This work was supported by funds from the Intramural Program of the NIH, NIDDK and the AIDS Targeted Antiviral program of the Office of the Director of the NIH (to G.M.C.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains Supplementary Figures 1–6, Supplementary Methods, Supplementary Table and additional references. (PDF 453 kb)
Supplementary Movie 1
Movie representation of left-hand panels of Supplementary Figure 4a. (MOV 105070 kb)
Supplementary Movie 2
Movie representation of right-hand panels of Supplementary Figure 4a. (MOV 91291 kb)
Rights and permissions
About this article
Cite this article
Tang, C., Iwahara, J. & Clore, G. Visualization of transient encounter complexes in protein–protein association. Nature 444, 383–386 (2006). https://doi.org/10.1038/nature05201
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05201
This article is cited by
-
Measuring transverse relaxation in highly paramagnetic systems
Journal of Biomolecular NMR (2020)
-
Understanding molecular mechanisms in cell signaling through natural and artificial sequence variation
Nature Structural & Molecular Biology (2019)
-
Development of R7BP inhibitors through cross-linking coupled mass spectrometry and integrated modeling
Communications Biology (2019)
-
Transition path times of coupled folding and binding reveal the formation of an encounter complex
Nature Communications (2018)
-
Structures of chaperone-substrate complexes docked onto the export gate in a type III secretion system
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.