Abstract
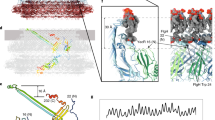
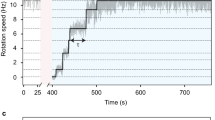
Many essential cellular processes are carried out by complex biological machines located in the cell membrane. The bacterial flagellar motor is a large membrane-spanning protein complex that functions as an ion-driven rotary motor to propel cells through liquid media1,2,3. Within the motor, MotB is a component of the stator that couples ion flow to torque generation and anchors the stator to the cell wall4,5. Here we have investigated the protein stoichiometry, dynamics and turnover of MotB with single-molecule precision in functioning bacterial flagellar motors in Escherichia coli. We monitored motor function by rotation of a tethered cell body6, and simultaneously measured the number and dynamics of MotB molecules labelled with green fluorescent protein (GFP–MotB) in the motor by total internal reflection fluorescence microscopy. Counting fluorophores by the stepwise photobleaching of single GFP molecules showed that each motor contains ∼22 copies of GFP–MotB, consistent with ∼11 stators each containing two MotB molecules. We also observed a membrane pool of ∼200 GFP–MotB molecules diffusing at ∼0.008 µm2 s-1. Fluorescence recovery after photobleaching and fluorescence loss in photobleaching showed turnover of GFP–MotB between the membrane pool and motor with a rate constant of the order of 0.04 s-1: the dwell time of a given stator in the motor is only ∼0.5 min. This is the first direct measurement of the number and rapid turnover of protein subunits within a functioning molecular machine.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Macnab, R. M. in Escherichia coli and Salmonella: Cellular and Molecular Biology (ed. Neidhardt, F. C.) 123–145 (American Society for Microbiology, Washington DC, 1996)
Berry, R. M. & Armitage, J. P. The bacterial flagella motor. Adv. Microb. Physiol. 41, 291–337 (1999)
Berg, H. C. The rotary motor of bacterial flagella. Annu. Rev. Biochem. 72, 19–54 (2003)
Block, S. M. & Berg, H. C. Successive incorporation of force-generating units in the bacterial rotary motor. Nature 309, 470–472 (1984)
Blair, D. F. & Berg, H. C. Restoration of torque in defective flagellar motors. Science 242, 1678–1681 (1988)
Silverman, M. & Simon, M. Flagellar rotation and the mechanism of bacterial motility. Nature 249, 73–74 (1974)
Tsien, R. Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998)
Mashanov, G. I., Tacon, D., Peckham, M. & Molloy, J. E. The spatial and temporal dynamics of pleckstrin homology domain binding at the plasma membrane measured by imaging single molecules in live mouse myoblasts. J. Biol. Chem. 279, 15274–15280 (2004)
Mullineaux, C. W. & Sarcina, M. Probing the dynamics of photosynthetic membranes with fluorescence recovery after photobleaching. Trends Plant Sci. 7, 237–240 (2002)
Ray, N., Nenninger, A., Mullineaux, C. W. & Robinson, C. Location and mobility of twin-arginine translocase subunits in the Escherichia coli plasma membrane. J. Biol. Chem. 280, 17961–17968 (2005)
Goodwin, J. S. & Kenworthy, A. K. Photobleaching approaches to investigate diffusional mobility and trafficking of Ras in living cells. Methods 37, 154–164 (2005)
Mullineaux, C. W., Nenninger, A., Ray, N. & Robinson, C. Diffusion of green fluorescent protein in three cell environments in Escherichia coli. J. Bacteriol. 188, 3442–3448 (2006)
Berry, R. M., Turner, L. & Berg, H. C. Mechanical limits of bacterial flagellar motors probed by electrorotation. Biophys. J. 69, 280–286 (1995)
Khan, S., Dapice, M. & Reese, T. S. Effects of mot gene expression on the structure of the flagellar motor. J. Mol. Biol. 202, 575–584 (1988)
Reid, S. W. et al. The maximum number of torque-generating units in the flagellar motor of Escherichia coli is at least 11. Proc. Natl Acad. Sci. USA 103, 8066–8071 (2006)
Kojima, S. & Blair, D. F. Solubilization and purification of the MotA/MotB complex of Escherichia coli. Biochemistry 43, 26–34 (2004)
Sowa, Y. et al. Direct observation of steps in rotation of the bacterial flagellar motor. Nature 437, 916–919 (2005)
Sourjik, V. & Berg, H. C. Localization of components of the chemotaxis machinery of Escherichia coli using fluorescent protein fusions. Mol. Microbiol. 37, 740–751 (2000)
Turner, L., Ryu, W. S. & Berg, H. C. Real-time imaging of fluorescent flagellar filaments. J. Bacteriol. 182, 2793–2801 (2000)
Dickson, R. M., Cubitt, A. B., Tsien, R. Y. & Moerner, W. E. On/off blinking and switching behaviour of single molecules of green fluorescent protein. Nature 388, 355–358 (1997)
Chu, S. Biology and polymer physics at the single-molecule level. Phil. Trans. R. Soc. Lond. A 361, 689–698 (2003)
Leake, M. C., Wilson, D., Gautel, M. & Simmons, R. M. The elasticity of single titin molecules using a two-bead optical tweezers assay. Biophys. J. 87, 1112–1135 (2004)
Svoboda, K., Schmidt, C. F., Schnapp, B. J. & Block, S. M. Direct observation of kinesin stepping by optical trapping interferometry. Nature 365, 721–727 (1993)
Mashanov, G. I., Tacon, D., Knight, A. E., Peckham, M. & Molloy, J. E. Visualizing single molecules inside living cells using total internal reflection fluorescence microscopy. Methods 29, 142–152 (2003)
Deich, J., Judd, E. M., McAdams, H. H. & Moerner, W. E. Visualization of the movement of single histidine kinase molecules in live Caulobacter cells. Proc. Natl Acad. Sci. USA 101, 15921–15926 (2004)
Van Way, S. M., Hosking, E. R., Braun, T. F. & Manson, M. D. Mot protein assembly into the bacterial flagellum: a model based on mutational analysis of the motB gene. J. Mol. Biol. 297, 7–24 (2000)
Demot, R. & Vanderleyden, J. The C-terminal sequence conservation between OMPA-related outer membrane proteins and MotB suggests a common function in both gram-positive and gram-negative bacteria, possibly in the interaction of these domains peptidoglycan. Mol. Microbiol. 12, 333–336 (1994)
Acknowledgements
We thank D. Blair for antibodies to flagellin and MotB. The research of M.C.L., J.H.C., R.M.B. and J.P.A. was supported by combined UK research councils via an Interdisciplinary Research Collaboration in Bionanotechnology (IRC), that of G.H.W. by the Biotechnology and Biological Sciences Research Council (BBSRC), and that of F.B. by a Clarendon Scholarship. Author Contributions Fluorescence experiments were carried out by M.C.L. and J.H.C. in the laboratory of R.M.B.; strain construction was done by J.H.C. and G.H.W. in the laboratory of J.P.A.; data analysis was done by M.C.L., R.M.B. and G.H.W.; and simulations were carried out by F.B. and M.C.L. The experiment was designed by M.C.L., R.M.B. and J.P.A.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains Supplementary Methods and Supplementary Figures 1–13. (PDF 1180 kb)
Supplementary Video Legends
This file contains text to accompany the below Supplementary Videos. (PDF 10 kb)
Supplementary Video 1
Brightfield video for a freely rotating tethered GFP-MotB cell and fixed cell in the same field of view. (MPG 141 kb)
Supplementary Video 2
Continuous TIRF bleach of cells in Supplementary Video 1. (MPG 638 kb)
Supplementary Video 3
Continuous TIRF illumination on a different pre-bleached fixed GFP-MotB cells showing diffusing GFP-MotB in the membrane. (AVI 212 kb)
Supplementary Video 4
Continuous TIRF illumination on a different pre-bleached fixed GFP-MotB cells showing diffusing GFP-MotB in the membrane. (AVI 329 kb)
Supplementary Video 5
Brightfield video for a freely rotating tethered GFP-MotB cell. (MPG 232 kb)
Supplementary Video 6
Continuous TIRF bleach of cell in Supplementary Video 5. (MPG 143 kb)
Supplementary Video 7
Continuous TIRF bleach of cell in Supplementary Video 4, 60 min after the end of the initial bleach. (MPG 143 kb)
Rights and permissions
About this article
Cite this article
Leake, M., Chandler, J., Wadhams, G. et al. Stoichiometry and turnover in single, functioning membrane protein complexes. Nature 443, 355–358 (2006). https://doi.org/10.1038/nature05135
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05135
This article is cited by
-
Direct observation of autoubiquitination for an integral membrane ubiquitin ligase in ERAD
Nature Communications (2024)
-
Precise programming of multigene expression stoichiometry in mammalian cells by a modular and programmable transcriptional system
Nature Communications (2023)
-
Single-molecule dynamics show a transient lipopolysaccharide transport bridge
Nature (2023)
-
Integration of protein context improves protein-based COVID-19 patient stratification
Clinical Proteomics (2022)
-
A multi-state dynamic process confers mechano-adaptation to a biological nanomachine
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.