Abstract
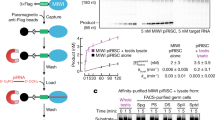
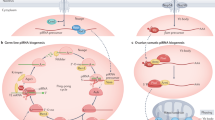
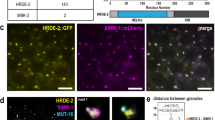
Small RNAs associate with Argonaute proteins and serve as sequence-specific guides to regulate messenger RNA stability, protein synthesis, chromatin organization and genome structure1,2,3. In animals, Argonaute proteins segregate into two subfamilies4. The Argonaute subfamily acts in RNA interference and in microRNA-mediated gene regulation using 21–22-nucleotide RNAs as guides. The Piwi subfamily is involved in germline-specific events such as germline stem cell maintenance and meiosis. However, neither the biochemical function of Piwi proteins nor the nature of their small RNA guides is known. Here we show that MIWI, a murine Piwi protein, binds a previously uncharacterized class of ∼29–30-nucleotide RNAs that are highly abundant in testes. We have therefore named these Piwi-interacting RNAs (piRNAs). piRNAs show distinctive localization patterns in the genome, being predominantly grouped into 20–90-kilobase clusters, wherein long stretches of small RNAs are derived from only one strand. Similar piRNAs are also found in human and rat, with major clusters occurring in syntenic locations. Although their function must still be resolved, the abundance of piRNAs in germline cells and the male sterility of Miwi mutants suggest a role in gametogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hannon, G. J. RNA interference. Nature 418, 244–251 (2002)
Meister, G. & Tuschl, T. Mechanisms of gene silencing by double-stranded RNA. Nature 431, 343–349 (2004)
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004)
Carmell, M. A., Xuan, Z., Zhang, M. Q. & Hannon, G. J. The Argonaute family: tentacles that reach into RNAi, developmental control, stem cell maintenance, and tumorigenesis. Genes Dev. 16, 2733–2742 (2002)
Cox, D. N. et al. A novel class of evolutionarily conserved genes defined by piwi are essential for stem cell self-renewal. Genes Dev. 12, 3715–3727 (1998)
Lin, H. & Spradling, A. C. A novel group of pumilio mutations affects the asymmetric division of germline stem cells in the Drosophila ovary. Development 124, 2463–2476 (1997)
Cox, D. N., Chao, A. & Lin, H. piwi encodes a nucleoplasmic factor whose activity modulates the number and division rate of germline stem cells. Development 127, 503–514 (2000)
Pal-Bhadra, M., Bhadra, U. & Birchler, J. A. RNAi related mechanisms affect both transcriptional and posttranscriptional transgene silencing in Drosophila. Mol. Cell 9, 315–327 (2002)
Pal-Bhadra, M. et al. Heterochromatic silencing and HP1 localization in Drosophila are dependent on the RNAi machinery. Science 303, 669–672 (2004)
Aravin, A. A. et al. Dissection of a natural RNA silencing process in the Drosophila melanogaster germ line. Mol. Cell. Biol. 24, 6742–6750 (2004)
Aravin, A. A. et al. Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr. Biol. 11, 1017–1027 (2001)
Kalmykova, A. I., Klenov, M. S. & Gvozdev, V. A. Argonaute protein PIWI controls mobilization of retrotransposons in the Drosophila male germline. Nucleic Acids Res. 33, 2052–2059 (2005)
Savitsky, M., Kwon, D., Georgiev, P., Kalmykova, A. & Gvozdev, V. Telomere elongation is under the control of the RNAi-based mechanism in the Drosophila germline. Genes Dev. 20, 345–354 (2006)
Vagin, V. V. et al. The RNA interference proteins and vasa locus are involved in the silencing of retrotransposons in the female germline of Drosophila melanogaster. RNA Biol. 1, 51–55 (2004)
Sarot, E., Payen-Groschene, G., Bucheton, A. & Pelisson, A. Evidence for a piwi-dependent RNA silencing of the gypsy endogenous retrovirus by the Drosophila melanogaster flamenco gene. Genetics 166, 1313–1321 (2004)
Deng, W. & Lin, H. miwi, a murine homolog of piwi, encodes a cytoplasmic protein essential for spermatogenesis. Dev. Cell 2, 819–830 (2002)
Kuramochi-Miyagawa, S. et al. Mili, a mammalian member of piwi family gene, is essential for spermatogenesis. Development 131, 839–849 (2004)
Ambros, V. & Lee, R. C. Identification of microRNAs and other tiny noncoding RNAs by cDNA cloning. Methods Mol. Biol. 265, 131–158 (2004)
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005)
Aravin, A. A. et al. The small RNA profile during Drosophila melanogaster development. Dev. Cell 5, 337–350 (2003)
Sijen, T. et al. On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 107, 465–476 (2001)
Bourc'his, D. & Bestor, T. H. Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature 431, 96–99 (2004)
Webster, K. E. et al. Meiotic and epigenetic defects in Dnmt3L-knockout mouse spermatogenesis. Proc. Natl Acad. Sci. USA 102, 4068–4073 (2005)
Peters, A. H. et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 (2001)
Caudy, A. A., Myers, M., Hannon, G. J. & Hammond, S. M. Fragile X-related protein and VIG associate with the RNA interference machinery. Genes Dev. 16, 2491–2496 (2002)
Acknowledgements
We thank N. Sheth for help with the informatics analysis of piRNA populations, and W. R. McCombie for pilot sequencing. C. Cordon-Cardo and M. Golding provided tissue samples. We thank C. Mello, L. Murchison, S. Cheloufi and O. Tam for helpful discussions. This work was supported in part by grants from the NIH (G.J.H.). A.G. is a Florence Gould Fellow of the Watson School of Biological Sciences. G.J.H. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The GenBank accession numbers for piR-1, piR-2 and piR-3 are DQ539889, DQ539890 and DQ539891, respectively. Mouse piRNA accession numbers range from DQ539889 to DQ569912; human piRNA accession numbers range from DQ569913 to DQ601958; and rat piRNA accession numbers range from DQ601959 to DQ628526. Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Methods
Full description of the materials and methods used in this study. (DOC 29 kb)
Supplementary Tables 1–4
Supplementary Table 1 indicates the number of piRNA clones mapping to each category of sequences in the mouse genome. Supplementary Tables 2, 3 and 4 describe the piRNA clusters in mouse, human and rat, respectively. (DOC 901 kb)
Supplementary Figures 1–2
Supplementary Figure 1 shows that piRNAs are degraded by RNaseA, but not DNase I. Supplementary Figure 2 describes the general features of the mouse piRNA clones. (PDF 228 kb)
Supplementary Figures 3–6
Supplementary Figures 3, 4, 5 and 6 represent the density of mouse piRNAs along chromosomes 1 and 2, 3 and 4, 5 and 6, 7 and 8, respectively. (PDF 3383 kb)
Supplementary Figures 7–11
Supplementary Figures 7, 8, 9, 10 and 11 represent the density of mouse piRNAs along chromosomes 9 and 10, 11 and 12, 13 and 14, 15 and 16, 18 and 19, respectively. (PDF 3145 kb)
Rights and permissions
About this article
Cite this article
Girard, A., Sachidanandam, R., Hannon, G. et al. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442, 199–202 (2006). https://doi.org/10.1038/nature04917
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04917
This article is cited by
-
PiRNA hsa_piR_019949 promotes chondrocyte anabolic metabolism by inhibiting the expression of lncRNA NEAT1
Journal of Orthopaedic Surgery and Research (2024)
-
Activation of human endogenous retroviruses and its physiological consequences
Nature Reviews Molecular Cell Biology (2024)
-
The emerging role of the piRNA/PIWI complex in respiratory tract diseases
Respiratory Research (2023)
-
Regulation of Miwi-mediated mRNA stabilization by Ck137956/Tssa is essential for male fertility
BMC Biology (2023)
-
The epigenetic regulatory mechanism of PIWI/piRNAs in human cancers
Molecular Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.