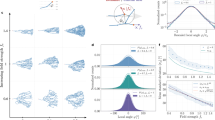
Abstract
Many organs are composed of tubular networks that arise by branching morphogenesis in which cells bud from an epithelium and organize into a tube1,2,3. Fibroblast growth factors (FGFs) and other signalling molecules have been shown to guide branch budding and outgrowth4,5,6,7, but it is not known how epithelial cells coordinate their movements and morphogenesis. Here we use genetic mosaic analysis in Drosophila melanogaster to show that there are two functionally distinct classes of cells in budding tracheal branches: cells at the tip that respond directly to Branchless FGF and lead branch outgrowth, and trailing cells that receive a secondary signal to follow the lead cells and form a tube. These roles are not pre-specified; rather, there is competition between cells such that those with the highest FGF receptor activity take the lead positions, whereas those with less FGF receptor activity assume subsidiary positions and form the branch stalk. Competition appears to involve Notch-mediated lateral inhibition that prevents extra cells from assuming the lead. There may also be cooperation between budding cells, because in a mosaic epithelium, cells that cannot respond to the chemoattractant, or respond only poorly, allow other cells in the epithelium to move ahead of them.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hogan, B. L. & Kolodziej, P. A. Organogenesis: molecular mechanisms of tubulogenesis. Nature Rev. Genet. 3, 513–523 (2002)
Metzger, R. J. & Krasnow, M. A. Genetic control of branching morphogenesis. Science 284, 1635–1639 (1999)
Affolter, M. et al. Tube or not tube: remodeling epithelial tissues by branching morphogenesis. Dev. Cell 4, 11–18 (2003)
Sutherland, D., Samakovlis, C. & Krasnow, M. A. branchless encodes a Drosophila FGF homolog that controls tracheal cell migration and the pattern of branching. Cell 87, 1091–1101 (1996)
Park, W. Y., Miranda, B., Lebeche, D., Hashimoto, G. & Cardoso, W. V. FGF-10 is a chemotactic factor for distal epithelial buds during lung development. Dev. Biol. 201, 125–134 (1998)
Weaver, M., Dunn, N. R. & Hogan, B. L. Bmp4 and Fgf10 play opposing roles during lung bud morphogenesis. Development 127, 2695–2704 (2000)
Pepicelli, C. V., Kispert, A., Rowitch, D. H. & McMahon, A. P. GDNF induces branching and increased cell proliferation in the ureter of the mouse. Dev. Biol. 192, 193–198 (1997)
Samakovlis, C. et al. Development of the Drosophila tracheal system occurs by a series of morphologically distinct but genetically coupled branching events. Development 122, 1395–1407 (1996)
Ghabrial, A., Luschnig, S., Metzstein, M. M. & Krasnow, M. A. Branching morphogenesis of the Drosophila tracheal system. Annu. Rev. Cell Dev. Biol. 19, 623–647 (2003)
Klambt, C., Glazer, L. & Shilo, B. Z. breathless, a Drosophila FGF receptor homolog, is essential for migration of tracheal and specific midline glial cells. Genes Dev. 6, 1668–1678 (1992)
Klambt, C. The Drosophila gene pointed encodes two ETS-like proteins which are involved in the development of the midline glial cells. Development 117, 163–176 (1993)
Jarecki, J., Johnson, E. & Krasnow, M. A. Oxygen regulation of airway branching in Drosophila is mediated by branchless FGF. Cell 99, 211–220 (1999)
Llimargas, M. Wingless and its signalling pathway have common and separable functions during tracheal development. Development 127, 4407–4417 (2000)
Chihara, T. & Hayashi, S. Control of tracheal tubulogenesis by Wingless signaling. Development 127, 4433–4442 (2000)
Llimargas, M. The Notch pathway helps to pattern the tips of the Drosophila tracheal branches by selecting cell fates. Development 126, 2355–2364 (1999)
Ikeya, T. & Hayashi, S. Interplay of Notch and FGF signaling restricts cell fate and MAPK activation in the Drosophila trachea. Development 126, 4455–4463 (1999)
Steneberg, P., Hemphala, J. & Samakovlis, C. Dpp and Notch specify the fusion cell fate in the dorsal branches of the Drosophila trachea. Mech. Dev. 87, 153–163 (1999)
Gabay, L., Seger, R. & Shilo, B. Z. MAP kinase in situ activation atlas during Drosophila embryogenesis. Development 124, 3535–3541 (1997)
Ribeiro, C., Ebner, A. & Affolter, M. In vivo imaging reveals different cellular functions for FGF and Dpp signaling in tracheal branching morphogenesis. Dev. Cell 2, 677–683 (2002)
Vincent, S., Wilson, R., Coelho, C., Affolter, M. & Leptin, M. The Drosophila protein Dof is specifically required for FGF signaling. Mol. Cell 2, 515–525 (1998)
Imam, F., Sutherland, D., Huang, W. & Krasnow, M. A. stumps, a Drosophila gene required for fibroblast growth factor (FGF)-directed migrations of tracheal and mesodermal cells. Genetics 152, 307–318 (1999)
Hacohen, N., Kramer, S., Sutherland, D., Hiromi, Y. & Krasnow, M. A. sprouty encodes a novel antagonist of FGF signaling that patterns apical branching of the Drosophila airways. Cell 92, 253–263 (1998)
Casci, T., Vinos, J. & Freeman, M. Sprouty, an intracellular inhibitor of Ras signaling. Cell 96, 655–665 (1999)
Ohshiro, T., Emori, Y. & Saigo, K. Ligand-dependent activation of breathless FGF receptor gene in Drosophila developing trachea. Mech. Dev. 114, 3–11 (2002)
Gerhardt, H. G., Alitalo, K., Shima, D., Betsholtz, C. et al. VEGF guides angiogenic sproutting utilizing endothelial tip cell filopodia. J. Cell Biol. 161, 1163–1177 (2003)
Shakya, R., Watanabe, T. & Costantini, F. The role of GDNF/Ret signaling in ureteric bud cell fate and branching morphogenesis. Dev. Cell 8, 65–74 (2005)
Cabernard, C. & Affolter, M. Distinct roles for two receptor tyrosine kinases in epithelial branching morphogenesis in Drosophila. Dev. Cell 9, 831–842 (2005)
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993)
Golic, K. G. Site-specific recombination between homologous chromosomes in Drosophila. Science 252, 958–961 (1991)
Luschnig, S., Batz, T., Armbruster, K. & Krasnow, M. A. serpentine and vermiform encode matrix proteins with chitin binding and deacetylation domains that limit tracheal tube length in Drosophila. Curr. Biol. 16, 186–194 (2006)
Acknowledgements
We thank B. Levi for help with isolation of ‘no mutant terminal cell’ mutants and for discussion, and S. Artavanis, M. Galko, S. Luschnig and S. Toering for fly stocks and antisera. This work was supported by an NIH NRSA fellowship (A.S.G.) and a grant from the N.I.H. M.A.K. is an investigator of the Howard Hughes Medical Institute. Author Contributions A.S.G. conceived, designed and performed experiments and analysed data. M.A.K. advised on the above. A.S.G. and M.A.K. wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Methods
This file contains additional details of the methods used in this study. (DOC 8 kb)
Supplementary Table 1
Distribution of positively-marked cell clones in mosaic dorsal branches. (DOC 10 kb)
Supplementary Table 2
Distribution of control tracheal clones (btl+/+) in wild-type (btl+/+) animals. Each row represents the structure of a mosaic DB of the genotype indicated in the title of the table, and each cell in a row describes the tracheal cell at that position in the DB. (DOC 591 kb)
Supplementary Table 3
Distribution of btl724/724 tracheal clones in heterozygous (btl+/724) animals. Each row represents the structure of a mosaic DB of the genotype indicated in the title of the table, and each cell in a row describes the tracheal cell at that position in the DB. (DOC 334 kb)
Supplementary Table 4
Distribution of btl788/788 tracheal clones in heterozygyous (btl+/788) animals. Each row represents the structure of a mosaic DB of the genotype indicated in the title of the table, and each cell in a row describes the tracheal cell at that position in the DB. (DOC 351 kb)
Supplementary Table 5
Distribution of pntΔ88/Δ88 tracheal clones in heterozygous (pnt+/Δ88) animals. Each row represents the structure of a mosaic DB of the genotype indicated in the title of the table, and each cell in a row describes the tracheal cell at that position in the DB. (DOC 311 kb)
Supplementary Table 6
Distribution of btlBN/BN tracheal clones in heterozygous (btl+/BN) animals. Each row represents the structure of a mosaic DB of the genotype indicated in the title of the table, and each cell in a row describes the tracheal cell at that position in the DB. (DOC 420 kb)
Supplementary Table 7
Distribution of wild-type (btl+/+) tracheal clones in heterozygous (btl+/788) animals. Each row represents the structure of a mosaic DB of the genotype indicated in the title of the table, and each cell in a row describes the tracheal cell at that position in the DB. (DOC 450 kb)
Supplementary Table 8
Distribution of wild-type (btl+/+) tracheal clones in heterozygous (btl+/1187) animals. Each row represents the structure of a mosaic DB of the genotype indicated in the title of the table, and each cell in a row describes the tracheal cell at that position in the DB. (DOC 431 kb)
Supplementary Table 9
Distribution of spryΔ5/Δ5 tracheal clones in heterozygous (spry+/Δ5) animals. Each row represents the structure of a mosaic DB of the genotype indicated in the title of the table, and each cell in a row describes the tracheal cell at that position in the DB. (DOC 346 kb)
Rights and permissions
About this article
Cite this article
Ghabrial, A., Krasnow, M. Social interactions among epithelial cells during tracheal branching morphogenesis. Nature 441, 746–749 (2006). https://doi.org/10.1038/nature04829
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04829
This article is cited by
-
Role of senescent cells in the motile behavior of active, non-senescent cells in confluent populations
Scientific Reports (2022)
-
Numerical Simulations of Stochastic Differential Equation Model for Spatiotemporal Biological Interactions
International Journal of Applied and Computational Mathematics (2022)
-
Branching pattern and morphogenesis of medusa tentacles in the jellyfish Cladonema pacificum (Hydrozoa, Cnidaria)
Zoological Letters (2019)
-
Anxa4 mediated airway progenitor cell migration promotes distal epithelial cell fate specification
Scientific Reports (2018)
-
Heterogeneity in VEGFR3 levels drives lymphatic vessel hyperplasia through cell-autonomous and non-cell-autonomous mechanisms
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.