Abstract
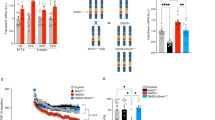
Trisomy 21 results in Down's syndrome, but little is known about how a 1.5-fold increase in gene dosage produces the pleiotropic phenotypes of Down's syndrome. Here we report that two genes, DSCR1 and DYRK1A , lie within the critical region of human chromosome 21 and act synergistically to prevent nuclear occupancy of NFATc transcription factors, which are regulators of vertebrate development. We use mathematical modelling to predict that autoregulation within the pathway accentuates the effects of trisomy of DSCR1 and DYRK1A, leading to failure to activate NFATc target genes under specific conditions. Our observations of calcineurin-and Nfatc-deficient mice, Dscr1- and Dyrk1a–overexpressing mice, mouse models of Down's syndrome and human trisomy 21 are consistent with these predictions. We suggest that the 1.5-fold increase in dosage of DSCR1 and DYRK1A cooperatively destabilizes a regulatory circuit, leading to reduced NFATc activity and many of the features of Down's syndrome. More generally, these observations suggest that the destabilization of regulatory circuits can underlie human disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
McAdams, H. H. & Shapiro, L. A bacterial cell-cycle regulatory network operating in time and space. Science 301, 1874–1877 (2003)
Davidson, E. H. et al. A genomic regulatory network for development. Science 295, 1669–1678 (2002)
Korenberg, J. R. et al. Down syndrome: toward a molecular definition of the phenotype. Am. J. Med. Genet. 7 (Suppl.), 91–97 (1990)
Holtzman, D. M. & Epstein, C. J. The molecular genetics of Down syndrome. Mol. Genet. Med. 2, 105–120 (1992)
Korenberg, J. R. et al. Down syndrome phenotypes: the consequences of chromosomal imbalance. Proc. Natl Acad. Sci. USA 91, 4997–5001 (1994)
Antonarakis, S. E., Lyle, R., Dermitzakis, E. T., Reymond, A. & Deutsch, S. Chromosome 21 and Down syndrome: from genomics to pathophysiology. Nature Rev. Genet. 5, 725–738 (2004)
Graef, I. A., Chen, F. & Crabtree, G. R. NFAT signaling in vertebrate development. Curr. Opin. Genet. Dev. 11, 505–512 (2001)
Crabtree, G. R. & Olson, E. N. NFAT signaling: choreographing the social lives of cells. Cell 109 (Suppl.), S67–S79 (2002)
Flanagan, W. M., Corthesy, B., Bram, R. J. & Crabtree, G. R. Nuclear association of a T-cell transcription factor blocked by FK-506 and cyclosporin A. Nature 352, 803–807 (1991)
Beals, C. R., Sheridan, C. M., Turck, C. W., Gardner, P. & Crabtree, G. R. Nuclear export of NF-ATc enhanced by glycogen synthase kinase-3. Science 275, 1930–1934 (1997)
Graef, I. A. et al. L-type calcium channels and GSK-3 regulate the activity of NF-ATc4 in hippocampal neurons. Nature 401, 703–708 (1999)
Epstein, C. J. in The Consequences of Chromosome Imbalance (eds Barlow, P. W., Green, P. B. & Wylie, C. C.) (Cambridge Univ. Press, Cambridge, 1986)
Richtsmeier, J. T., Baxter, L. L. & Reeves, R. H. Parallels of craniofacial maldevelopment in Down syndrome and Ts65Dn mice. Dev. Dyn. 217, 137–145 (2000)
Torfs, C. P. & Christianson, R. E. Anomalies in Down syndrome individuals in a large population-based registry. Am. J. Med. Genet. 77, 431–438 (1998)
Zeng, H. et al. Forebrain-specific calcineurin knockout selectively impairs bidirectional synaptic plasticity and working/episodic-like memory. Cell 107, 617–629 (2001)
Graef, I. A. et al. Neurotrophins and netrins require calcineurin/NFAT signaling to stimulate outgrowth of embryonic axons. Cell 113, 657–670 (2003)
Benedito, A. B. et al. The transcription factor NFAT3 mediates neuronal survival. J. Biol. Chem. 280, 2818–2825 (2005)
Baxter, L. L., Moran, T. H., Richtsmeier, J. T., Troncoso, J. & Reeves, R. H. Discovery and genetic localization of Down syndrome cerebellar phenotypes using the Ts65Dn mouse. Hum. Mol. Genet. 9, 195–202 (2000)
Kegley, K. M., Gephart, J., Warren, G. L. & Pavlath, G. K. Altered primary myogenesis in NFATC3-/- mice leads to decreased muscle size in the adult. Dev. Biol. 232, 115–126 (2001)
Graef, I. A., Chen, F., Chen, L., Kuo, A. & Crabtree, G. R. Signals transduced by Ca2+/calcineurin and NFATc3/c4 pattern the developing vasculature. Cell 105, 863–875 (2001)
Chang, C. P. et al. A field of myocardial-endocardial NFAT signaling underlies heart valve morphogenesis. Cell 118, 649–663 (2004)
Chang, C. P. et al. Calcineurin is required in urinary tract mesenchyme for the development of the pyeloureteral peristaltic machinery. J. Clin. Invest. 113, 1051–1058 (2004)
Hogan, P. G., Chen, L., Nardone, J. & Rao, A. Transcriptional regulation by calcium, calcineurin, and NFAT. Genes Dev. 17, 2205–2232 (2003)
Rothermel, B. et al. A protein encoded within the Down syndrome critical region is enriched in striated muscles and inhibits calcineurin signaling. J. Biol. Chem. 275, 8719–8725 (2000)
Fuentes, J. J. et al. DSCR1, overexpressed in Down syndrome, is an inhibitor of calcineurin-mediated signaling pathways. Hum. Mol. Genet. 9, 1681–1690 (2000)
Reeves, R. H. et al. A mouse model for Down syndrome exhibits learning and behaviour deficits. Nature Genet. 11, 177–184 (1995)
Sago, H. et al. Ts1Cje, a partial trisomy 16 mouse model for Down syndrome, exhibits learning and behavioural abnormalities. Proc. Natl Acad. Sci. USA 95, 6256–6261 (1998)
Richtsmeier, J. T., Zumwalt, A., Carlson, E. J., Epstein, C. J. & Reeves, R. H. Craniofacial phenotypes in segmentally trisomic mouse models for Down syndrome. Am. J. Med. Genet. 107, 317–324 (2002)
O'Doherty, A. et al. An aneuploid mouse strain carrying human chromosome 21 with down syndrome phenotypes. Science 309, 2033–2037 (2005)
Olson, L. E., Richtsmeier, J. T., Leszl, J. & Reeves, R. H. A chromosome 21 critical region does not cause specific Down syndrome phenotypes. Science 306, 687–690 (2004)
Kentrup, H. et al. Dyrk, a dual specificity protein kinase with unique structural features whose activity is dependent on tyrosine residues between subdomains VII and VIII. J. Biol. Chem. 271, 3488–3495 (1996)
Fotaki, V. et al. Dyrk1A haploinsufficiency affects viability and causes developmental delay and abnormal brain morphology in mice. Mol. Cell. Biol. 22, 6636–6647 (2002)
Altafaj, X. et al. Neurodevelopmental delay, motor abnormalities and cognitive deficits in transgenic mice overexpressing Dyrk1A (minibrain), a murine model of Down's syndrome. Hum. Mol. Genet. 10, 1915–1923 (2001)
Garaschuk, O., Linn, J., Eilers, J. & Konnerth, A. Large-scale oscillatory calcium waves in the immature cortex. Nature Neurosci. 3, 452–459 (2000)
Borodinsky, L. N. et al. Activity-dependent homeostatic specification of transmitter expression in embryonic neurons. Nature 429, 523–530 (2004)
Woods, Y. L. et al. The kinase DYRK phosphorylates protein-synthesis initiation factor eIF2Bɛ at Ser539 and the microtubule-associated protein tau at Thr212: potential role for DYRK as a glycogen synthase kinase 3-priming kinase. Biochem. J. 355, 609–615 (2001)
Timmerman, L. A., Clipstone, N. A., Ho, S. N., Northrop, J. P. & Crabtree, G. R. Rapid shuttling of NF-AT in discrimination of Ca2+ signals and immunosuppression. Nature 383, 837–840 (1996)
Ranger, A. M. et al. The transcription factor NF-ATc is essential for cardiac valve formation. Nature 392, 186–190 (1998)
de la Pompa, J. L. et al. Role of the NF-ATc transcription factor in morphogenesis of cardiac valves and septum. Nature 392, 182–186 (1998)
Northrop, J. P. et al. NF-AT components define a family of transcription factors targeted in T-cell activation. Nature 369, 497–502 (1994)
Fiering, S. et al. Single cell assay of a transcription factor reveals a threshold in transcription activated by signals emanating from the T-cell antigen receptor. Genes Dev. 4, 1823–1834 (1990)
Miyakawa, T. et al. Conditional calcineurin knockout mice exhibit multiple abnormal behaviors related to schizophrenia. Proc. Natl Acad. Sci. USA 100, 8987–8992 (2003)
Acknowledgements
We thank S. L. Schreiber, W. Mobley and K. Tanda for discussion and comments on the manuscript; W. Becker for Dyrk1a expression constructs; E. Olson for the anti-DCSR1 (MCIP1) antibody; R. S. Williams for the Dscr1 (MCIP1) expression construct; K. Stankunas, G. Krampitz and C. Shang for help with histology on Dyrk1a and Dscr1 transgenic mice; E. Wang for mass spectrometric analysis; W. Mobley and K. Zhan for providing Ts65Dn mice; and F. Wang, members of the Crabtree laboratory, J. Lee, M. Dionne and S. Arron for discussions. We thank the Stanford Center for Innovation in In Vivo Imaging (NCI Small Animal Imaging Resource Program Grant), the Stanford Imaging Facility and the Stanford Proteomics and Integrated Research Facility. These studies were supported by the Howard Hughes Medical Institute and NIH grants to G.R.C., and by the Christopher Reeve Paralysis Foundation (I.A.G.). M.M.W. is supported by a Stanford Graduate Fellowship and an HHMI predoctoral fellowship, C.-P.C. by grants from AHA and the NIH, J.R.A. by a postdoctoral fellowship from the Berry Foundation, J.J.H. and S.K.K. by the ADA, H.W. by a Damon Runyon Cancer Research Foundation postdoctoral fellowship and a Muscular Dystrophy Association research development grant, and T.M. by KAKENHI from JSPS and MEXT and by a grant from JST BIRD. Author Contributions The order of listing of the authors J.R.A., M.M.W., A.P. and I.A.G. does in no way reflect their relative contribution to this work. I.A.G. and G.R.C. are responsible for the original concept. I.A.G. generated the Nfatc mutant mice (Fig. 1, Supplementary Figs 2–5 and Table 1), J.R.N. the Cnb1 mutant mice (Supplementary Fig. 8c), and H.W. and L.C. the Dyrk1a/Dscr1 transgenic mice (Fig. 3). I.A.G., M.M.W., C.-P.C., X.G., J.R.N., J.J.H., S.K.K., N.Y. and T.M. analysed mutant mice (Figs 1, 3 and Supplementary Figs 2–5 and Table 1). M.M.W. performed the skull morphometry studies (Fig. 1a–c and Supplementary Figs 2–4) and helped with the analysis of Ts1Cje and Ts65Dn mice (Supplementary Fig. 9). I.A.G. performed the neuron signalling experiments (Fig. 2a, b, e, f and Supplementary Figs 7, 8b), biochemical analysis of human Down's syndrome samples (Fig. 4c), calcineurin mutant mice (Supplementary Fig. 8c), Ts1Cje and Ts65Dn mice (Supplementary Fig. 9) and Dyrk1a/Dscr1-overexpressing mice (Fig. 3a) as well as the Nfatc4 promoter studies (Supplementary Fig. 8). A.P. generated and solved the mathematical model (Fig. 4a, b, d and Supplementary Discussion B). U.F. provided the clinical samples (Fig. 4c). J.R.A. conducted the in vitro kinase (Fig. 2c, d and Supplementary Fig. 7) and 293T (Supplementary Fig. 6) assays and DSCR1/DYRK1A quantifications (used in Supplementary Discussion B), made the anti-DYRK1A antiserum and helped H.W. and I.A.G. to genotype some of the Dyrk1a/Dscr1-overexpressing mice. G.R.C., I.A.G., J.A.A., M.M.W. and A.P. wrote the manuscript and I.A.G., M.M.W., A.P. and G.R.C. generated the figures.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figure 1
Increased expression of DSCR1 and DYRK1a on chromosome 21 destabilizes the NFAT genetic circuit (PDF 354 kb)
Supplementary Figure 2
Craniofacial phenotype of NFATc2/c4 DKO mice (PDF 1726 kb)
Supplementary Figure 3
Measurement of cranial dimensions (PDF 1661 kb)
Supplementary Figure 4
Measurement of mandible dimensions. (PDF 1017 kb)
Supplementary Figure 5
NFATc mutant mice have defects in placental vascularization, annular pancreas and agangionic megacolon. (PDF 3846 kb)
Supplementary Figure 6
a, DYRK1a is localized to the nucleus. b, DYRK1a and DSCR1 inhibit NFAT transcriptional activity in a dose-dependent manner. c, DYRK1a kinase activity prevents nuclear accumulation of NFATc1. (PDF 927 kb)
Supplementary Figure 7
a, Diagram of NFATc4 constructs. b, Serine-to-alanine mutations of in the SRR- and SP-region of NFATc4 result in Ca2+-independent nuclear localization of EGFP-NFATc4. c, DYRK1a targets the third serine in the SP1 region, permitting processive phosphorylation of the second, then the first serine by GSK-3. (PDF 900 kb)
Supplementary Figure 8
a, NFAT binding sites in the NFATc4 promoter. b, The NFATc4 promoter in cortical neurons is regulated by CaN/NFAT activity. c, positive feedback regulation of NFATc1 and c4. (PDF 522 kb)
Supplementary Figure 9
Immunoblot of DYRK1a, HSP-90, NFATc4 and DSCR1 in whole cell extracts from E11.5 heads and E13.5 cerebral cortex from trisomic and control Ts1Cje embyros. (PDF 2604 kb)
Supplementary Discussion A
Discussion of the Down syndrome critical region and additional references. (PDF 98 kb)
Supplementary Discussion B
A Mathematical Model of the NFAT Genetic Circuit (PDF 296 kb)
Supplementary Methods
This file contains additional details of the methods used in this study. (PDF 148 kb)
Rights and permissions
About this article
Cite this article
Arron, J., Winslow, M., Polleri, A. et al. NFAT dysregulation by increased dosage of DSCR1 and DYRK1A on chromosome 21. Nature 441, 595–600 (2006). https://doi.org/10.1038/nature04678
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04678
This article is cited by
-
Harmine is an effective therapeutic small molecule for the treatment of cardiac hypertrophy
Acta Pharmacologica Sinica (2022)
-
The chromosome 21 kinase DYRK1A: emerging roles in cancer biology and potential as a therapeutic target
Oncogene (2022)
-
Prolonged contextual fear memory in AMPA receptor palmitoylation-deficient mice
Neuropsychopharmacology (2022)
-
Neurogenetic disorders across the lifespan: from aberrant development to degeneration
Nature Reviews Neurology (2022)
-
Embryonic organizer formation disorder leads to multiorgan dysplasia in Down syndrome
Cell Death & Disease (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.