Abstract
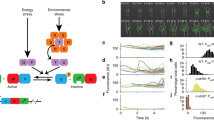
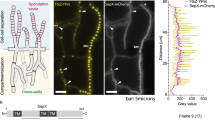
Sporulation in Bacillus subtilis serves as a model for the development of two different cell types from a single cell1. Although much information has been accumulated about the mechanisms that initiate the developmental programmes2, important questions remain that can be answered only by quantitative analysis. Here we develop, with the help of existing and new experimental results, a mathematical model that reproduces published in vitro experiments and explains how the activation of the key transcription factor is regulated. The model identifies the difference in volume between the two cell types as the primary trigger for determining cell fate. It shows that this effect depends on the allosteric behaviour of a key protein kinase and on a low rate of dephosphorylation by the corresponding phosphatase; both predicted effects are confirmed experimentally.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hilbert, D. W. & Piggot, P. J. Compartmentalization of gene expression during Bacillus subtilis spore formation. Microbiol. Mol. Biol. Rev. 68, 234–262 (2004)
Yudkin, M. D. & Clarkson, J. Differential gene expression in genetically identical sister cells: the initiation of sporulation in Bacillus subtilis. Mol. Microbiol. 56, 578–589 (2005)
Duncan, L., Alper, S., Arigoni, F., Losick, R. & Stragier, P. Activation of cell-specific transcription by a serine phosphatase at the site of asymmetric division. Science 270, 641–644 (1995)
Lord, M., Barillà, D. & Yudkin, M. D. Replacement of vegetative σA by sporulation-specific σF as a component of the RNA polymerase holoenzyme in sporulating Bacillus subtilis. J. Bacteriol. 181, 2346–2350 (1999)
Diederich, B. et al. Role of interactions between SpoIIAA and SpoIIAB in regulating cell-specific transcription factor σF of Bacillus subtilis. Genes Dev. 8, 2653–2663 (1994)
Clarkson, J., Shu, J.-C., Harris, D. A., Campbell, I. D. & Yudkin, M. D. Fluorescence and kinetic analysis of the SpoIIAB phosphorylation reaction, a key regulator of sporulation in Bacillus subtilis. Biochemistry 43, 3120–3128 (2004)
Clarkson, J., Campbell, I. D. & Yudkin, M. D. Physical evidence for the induced release of the Bacillus subtilis transcription factor, σF, from its inhibitory complex. J. Mol. Biol. 340, 203–209 (2004)
Clarkson, J., Campbell, I. D. & Yudkin, M. D. Efficient regulation of σF, the first sporulation-specific sigma factor in B. subtilis. J. Mol. Biol. 342, 1187–1195 (2004)
Shu, J.-C., Clarkson, J. & Yudkin, M. D. Studies of SpoIIAB mutant proteins elucidate the mechanisms that regulate the developmental transcription factor σF in Bacillus subtilis. Biochem. J. 384, 169–178 (2004)
Garsin, D. A., Duncan, L., Paskowitz, D. M. & Losick, R. The kinase activity of the antisigma factor SpoIIAB is required for activation as well as inhibition of transcription factor σF during sporulation in Bacillus subtilis. J. Mol. Biol. 284, 569–578 (1998)
Lee, C.-S., Lucet, I. & Yudkin, M. D. Fate of the SpoIIAB*-ADP liberated after SpoIIAB phosphorylates SpoIIAA of Bacillus subtilis. J. Bacteriol. 182, 6250–6253 (2000)
Lucet, I., Borriss, R. & Yudkin, M. D. Purification, kinetic properties, and intracellular concentration of SpoIIE, an integral membrane protein that regulates sporulation in Bacillus subtilis. J. Bacteriol. 181, 3242–3245 (1999)
Schroeter, R., Schlisio, S., Lucet, I., Yudkin, M. D. & Borriss, R. The Bacillus subtilis regulator protein SpoIIE shares functional and structural similarities with eukaryotic protein phosphatases 2C. FEMS Microbiol. Lett. 174, 117–123 (1999)
Eisenstadt, E., Fisher, S., Der, C. L. & Silver, S. Manganese transport in Bacillus subtilis W23 during growth and sporulation. J. Bacteriol. 113, 1363–1372 (1973)
Magnin, T., Lord, M. & Yudkin, M. D. Contribution of partner switching and SpoIIAA cycling to regulation of σF activity in sporulating Bacillus subtilis. J. Bacteriol. 179, 3922–3927 (1997)
Fujita, M. & Sadaie, Y. Rapid isolation of RNA polymerase from sporulating cells of Bacillus subtilis. Gene 221, 185–190 (1998)
Kaern, M., Elston, T. C., Blake, W. J. & Collins, J. J. Stochasticity in gene expression: from theories to phenotypes. Nature Rev. Genet. 6, 451–464 (2005)
Frandsen, N., Barák, I., Karmazyn-Campelli, C. & Stragier, P. Transient gene asymmetry during sporulation and establishment of cell specificity in Bacillus subtilis. Genes Dev. 13, 394–399 (1999)
Dworkin, J. & Losick, R. Differential gene expression governed by chromosomal spatial asymmetry. Cell 107, 339–346 (2001)
Pan, Q., Garsin, D. A. & Losick, R. Self-reinforcing activation of a cell-specific transcription factor by proteolysis of an anti-sigma factor in Bacillus subtilis. Mol. Cell 8, 873–883 (2001)
Carniol, K., Eichenberger, P. & Losick, R. A threshold mechanism governing activation of the developmental regulatory protein σF in Bacillus subtilis. J. Biol. Chem. 279, 14860–14870 (2004)
King, N., Dreesen, O., Stragier, P., Pogliano, K. & Losick, R. Septation, dephosphorylation, and the activation of σF during sporulation in Bacillus subtilis. Genes Dev. 13, 1156–1167 (1999)
Acknowledgements
We thank J.-C. Shu for providing unpublished SPR and AA phosphorylation data. This work was supported by the BBSRC UK. D.I. is a Junior Research Fellow at St John's College, Oxford, and is supported by a EPSRC DTA studentship. I.D.C. acknowledges financial support from the Wellcome Trust and the NIH-funded Cell Migration Consortium. Author Contributions D.I. developed the model and conducted the simulations. J.C. performed the experiments and all the authors contributed to the concepts and writing of the paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains details on the derivation of the mathematical model and of parameter values and a discussion of the wider relevance of the model. The also contains Supplementary Figures 1–12 and Supplementary Tables 1–3. (PDF 762 kb)
Rights and permissions
About this article
Cite this article
Iber, D., Clarkson, J., Yudkin, M. et al. The mechanism of cell differentiation in Bacillus subtilis. Nature 441, 371–374 (2006). https://doi.org/10.1038/nature04666
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04666
This article is cited by
-
Extreme slow growth as alternative strategy to survive deep starvation in bacteria
Nature Communications (2019)
-
Microbial enhanced oil recovery—a modeling study of the potential of spore-forming bacteria
Computational Geosciences (2016)
-
Prediction stability in a data-based, mechanistic model of σF regulation during sporulation in Bacillus subtilis
Scientific Reports (2013)
-
Assembly reflects evolution of protein complexes
Nature (2008)
-
A quantitative study of the benefits of co‐regulation using the spoIIA operon as an example
Molecular Systems Biology (2006)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.