Abstract
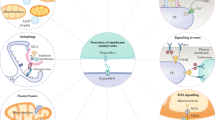
Eukaryotic cells have systems of internal organelles to synthesize lipids and membrane proteins, to release secreted proteins, to take up nutrients and to degrade membrane-bound and internalized molecules. Proteins and lipids move from organelle to organelle using transport vesicles. The accuracy of this traffic depends upon organelles being correctly recognized. In general, organelles are identified by the activated GTPases and specific lipid species that they display. These short-lived determinants provide organelles with an identity that is both unique and flexible. Recent studies have helped to establish how cells maintain and restrict these determinants and explain how this system is exploited by invading pathogens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bonifacino, J.S. & Glick, B. S. The mechanisms of vesicle budding and fusion. Cell 116, 153–166 (2004).
Zerial, M. & McBride, H. Rab proteins as membrane organizers. Nature Rev. Mol. Cell Biol. 2, 107–117 (2001).
Vetter, I.R. & Wittinghofer, A. The guanine nucleotide-binding switch in three dimensions. Science 294, 1299–1304 (2001).
Pereira-Leal, J.B. & Seabra, M.C. Evolution of the Rab family of small GTP-binding proteins. J. Mol. Biol. 313, 889–901 (2001).
Pfeffer, S.R. Rab GTPases: specifying and deciphering organelle identity and function. Trends Cell Biol. 11, 487–491 (2001).
Matanis, T. et al. Bicaudal-D regulates COPI-independent Golgi-ER transport by recruiting the dynein–dynactin motor complex. Nature Cell Biol. 4, 986–992 (2002).
Short, B., Preisinger, C., Schaletzky, J., Kopajtich, R. & Barr, F.A. The Rab6 GTPase regulates recruitment of the dynactin complex to Golgi membranes. Curr. Biol. 12, 1792–1795 (2002).
Fridmann-Sirkis, Y., Siniossoglou, S. & Pelham, H.R. TMF is a golgin that binds Rab6 and influences Golgi morphology. BMC Cell Biol. 5, 18 (2004).
Simonsen, A. et al. EEA1 links PI(3)K function to Rab5 regulation of endosome fusion. Nature 394, 494–498 (1998).
Seabra, M.C. & Coudrier, E. Rab GTPases and myosin motors in organelle motility. Traffic 5, 393–399 (2004).
Pfeffer, S. & Aivazian, D. Targeting Rab GTPases to distinct membrane compartments. Nature Rev. Mol. Cell Biol. 5, 886–896 (2004).
Seabra, M.C. & Wasmeier, C. Controlling the location and activation of Rab GTPases. Curr. Opin. Cell Biol. 16, 451–457 (2004).
Chavrier, P. et al. Hypervariable C-terminal domain of rab proteins acts as a targeting signal. Nature 353, 769–772 (1991).
Ali, B.R., Wasmeier, C., Lamoreux, L., Strom, M. & Seabra, M.C. Multiple regions contribute to membrane targeting of Rab GTPases. J. Cell Sci. 117, 6401–6412 (2004).
Calero, M. & Collins, R.N. Saccharomyces cerevisiae Pra1p/Yip3p interacts with Yip1p and Rab proteins. Biochem. Biophys. Res. Commun. 290, 676–681 (2002).
Figueroa, C., Taylor, J. & Vojtek, A.B. Prenylated Rab acceptor protein is a receptor for prenylated small GTPases. J. Biol. Chem. 276, 28219–28225 (2001).
Sivars, U., Aivazian, D. & Pfeffer, S.R. Yip3 catalyses the dissociation of endosomal Rab-GDI complexes. Nature 425, 856–859 (2003).
Hutt, D.M., Da-Silva, L.F., Chang, L.H., Prosser, D.C. & Ngsee, J.K. PRA1 inhibits the extraction of membrane-bound rab GTPase by GDI1. J.Biol. Chem. 275, 18511–18519 (2000).
Shakoori, A. et al. Identification of a five-pass transmembrane protein family localizing in the Golgi apparatus and the ER. Biochem. Biophys. Res. Commun. 312, 850–857 (2003).
Geng, J., Shin, M.E., Gilbert, P.M., Collins, R.N. & Burd, C.G. Saccharomyces cerevisiae Rab-GDI displacement factor ortholog Yip3p forms distinct complexes with the Ypt1 Rab GTPase and the reticulon Rtn1p. Eukaryot. Cell 4, 1166–1174 (2005).
Heidtman, M., Chen, C.Z., Collins, R.N. & Barlowe, C. A role for Yip1p in COPII vesicle biogenesis. J. Cell Biol. 163, 57–69 (2003).
Barrowman, J., Wang, W., Zhang, Y. & Ferro-Novick, S. The Yip1p.Yif1p complex is required for the fusion competence of endoplasmic reticulum-derived vesicles. J. Biol. Chem. 278, 19878–19884 (2003).
Munro, S. Organelle identity and the targeting of peripheral membrane proteins. Curr. Opin. Cell Biol. 14, 506–514 (2002).
Munro, S. Organelle identity and the organization of membrane traffic. Nature Cell Biol. 6, 469–472 (2004).
Ortiz, D., Medkova, M., Walch-Solimena, C. & Novick, P. Ypt32 recruits the Sec4p guanine nucleotide exchange factor, Sec2p, to secretory vesicles; evidence for a Rab cascade in yeast. J. Cell Biol. 157, 1005–1015 (2002).
Wang, W. & Ferro-Novick, S. A Ypt32p exchange factor is a putative effector of Ypt1p. Mol. Biol. Cell 13, 3336–3343 (2002).
Sato, M. et al. Caenorhabditis elegans RME-6 is a novel regulator of RAB-5 at the clathrin-coated pit. Nature Cell Biol. 7, 559–569 (2005).
Lippe, R., Miaczynska, M., Rybin, V., Runge, A. & Zerial, M. Functional synergy between Rab5 effector Rabaptin-5 and exchange factor Rabex-5 when physically associated in a complex. Mol. Biol. Cell 12, 2219–2228 (2001).
Haas, A.K., Fuchs, E., Kopajtich, R. & Barr, F.A. A GTPase-activating protein controls Rab5 function in endocytic trafficking. Nature Cell Biol. 7, 887–993 (2005).
De Antoni, A., Schmitzova, J., Trepte, H.H., Gallwitz, D. & Albert, S. Significance of GTP hydrolysis in Ypt1p-regulated endoplasmic reticulum to Golgi transport revealed by the analysis of two novel Ypt1-GAPs. J. Biol. Chem. 277, 41023–41031 (2002).
Lafourcade, C., Galan, J.M., Gloor, Y., Haguenauer-Tsapis, R. & Peter, M. The GTPase-activating enzyme Gyp1p is required for recycling of internalized membrane material by inactivation of the Rab/Ypt GTPase Ypt1p. Mol. Cell. Biol. 24, 3815–3826 (2004).
Lee, M.C., Miller, E.A., Goldberg, J., Orci, L. & Schekman, R. Bi-directional protein transport between the ER and Golgi. Annu. Rev. Cell Dev. Biol. 20, 87–123 (2004).
Gillingham, A.K., Tong, A.H., Boone, C. & Munro, S. The GTPase Arf1p and the ER to Golgi cargo receptor Erv14p cooperate to recruit the golgin Rud3p to the cis-Golgi. J. Cell Biol. 167, 281–292 (2004).
Donaldson, J.G., Honda, A. & Weigert, R. Multiple activities for Arf1 at the Golgi complex. Biochim. Biophys. Acta 1744, 364–373 (2005).
Krauss, M. et al. ARF6 stimulates clathrin/AP-2 recruitment to synaptic membranes by activating phosphatidylinositol phosphate kinase type Iγ. J. Cell Biol. 162, 113–124 (2003).
Setty, S.R., Shin, M.E., Yoshino, A., Marks, M.S. & Burd, C.G. Golgi recruitment of GRIP domain proteins by Arf-like GTPase 1 is regulated by Arf-like GTPase 3. Curr. Biol. 13, 401–404 (2003).
Panic, B., Whyte, J.R. & Munro, S. The ARF-like GTPases Arl1p and Arl3p act in a pathway that interacts with vesicle-tethering factors at the Golgi apparatus. Curr. Biol. 13, 405–410 (2003).
Pasqualato, S., Renault, L. & Cherfils, J. Arf, Arl, Arp and Sar proteins: a family of GTP-binding proteins with a structural device for ‘front-back’ communication. EMBO Rep. 3, 1035–1041 (2002).
Goldberg, J. Structural basis for activation of ARF GTPase: mechanisms of guanine nucleotide exchange and GTP-myristoyl switching. Cell 95, 237–248 (1998).
Robert, C.H., Cherfils, J., Mouawad, L. & Perahia, D. Integrating three views of Arf1 activation dynamics. J. Mol. Biol. 337, 969–983 (2004).
Jackson, C.L. & Casanova, J.E. Turning on ARF: the Sec7 family of guanine-nucleotide-exchange factors. Trends Cell Biol. 10, 60–67 (2000).
Garcia-Mata, R. & Sztul, E. The membrane-tethering protein p115 interacts with GBF1, an ARF guanine-nucleotide-exchange factor. EMBO Rep. 4, 320–325 (2003).
Chantalat, S. et al. A novel Golgi membrane protein is a partner of the ARF exchange factors Gea1p and Gea2p. Mol. Biol. Cell 14, 2357–2371 (2003).
Chantalat, S. et al. The Arf activator Gea2p and the P-type ATPase Drs2p interact at the Golgi in Saccharomyces cerevisiae. J. Cell Sci. 117, 711–722 (2004).
Donaldson, J.G. & Jackson, C.L. Regulators and effectors of the ARF GTPases. Curr. Opin. Cell Biol. 12, 475–482 (2000).
Gommel, D.U. et al. Recruitment to Golgi membranes of ADP-ribosylation factor 1 is mediated by the cytoplasmic domain of p23. EMBO J. 20, 6751–6760 (2001).
Honda, A., Al-Awar, O.S., Hay, J.C. & Donaldson, J.G. Targeting of Arf-1 to the early Golgi by membrin, an ER-Golgi SNARE. J. Cell Biol. 168, 1039–1051 (2005).
Behnia, R., Panic, B., Whyte, J.R. & Munro, S. Targeting of the Arf-like GTPase Arl3p to the Golgi requires N-terminal acetylation and the membrane protein Sys1p. Nature Cell Biol. 6, 405–413 (2004).
Setty, S.R., Strochlic, T.I., Tong, A.H., Boone, C. & Burd, C.G. Golgi targeting of ARF-like GTPase Arl3p requires its Nα-acetylation and the integral membrane protein Sys1p. Nature Cell Biol. 6, 414–419 (2004).
Liu, W., Duden, R., Phair, R.D. & Lippincott-Schwartz, J. ArfGAP1 dynamics and its role in COPI coat assembly on Golgi membranes of living cells. J. Cell Biol. 168, 1053–1063 (2005).
Bigay, J., Casella, J.F., Drin, G., Mesmin, B. & Antonny, B. ArfGAP1 responds to membrane curvature through the folding of a lipid packing sensor motif. EMBO J. 24, 2244–2253 (2005).
De Matteis, M.A. & Godi, A. PI-loting membrane traffic. Nature Cell Biol. 6, 487–492 (2004).
Wenk, M.R. & De Camilli, P. Protein–lipid interactions and phosphoinositide metabolism in membrane traffic: insights from vesicle recycling in nerve terminals. Proc. Natl Acad. Sci. USA 101, 8262–8269 (2004).
Stenmark, H., Aasland, R. & Driscoll, P.C. The phosphatidylinositol 3-phosphate-binding FYVE finger. FEBS Lett. 513, 77–84 (2002).
Ellson, C.D., Andrews, S., Stephens, L.R. & Hawkins, P.T. The PX domain: a new phosphoinositide-binding module. J. Cell Sci. 115, 1099–1105 (2002).
Efe, J.A., Botelho, R.J. & Emr, S.D. The Fab1 phosphatidylinositol kinase pathway in the regulation of vacuole morphology. Curr. Opin. Cell Biol. 17, 402–408 (2005).
Friant, S. et al. Ent3p Is a PtdIns(3,5)P2 effector required for protein sorting to the multivesicular body. Dev. Cell 5, 499–511 (2003).
Dove, S.K. et al. Svp1p defines a family of phosphatidylinositol 3,5-bisphosphate effectors. EMBO J. 23, 1922–1933 (2004).
Levine, T.P. & Munro, S. Targeting of Golgi-specific pleckstrin homology domains involves both PtdIns 4-kinase-dependent and -independent components. Curr. Biol. 12, 695–704 (2002).
Wang, Y.J. et al. Phosphatidylinositol 4 phosphate regulates targeting of clathrin adaptor AP-1 complexes to the Golgi. Cell 114, 299–310 (2003).
Balla, A., Tuymetova, G., Tsiomenko, A., Varnai, P. & Balla, T. A plasma membrane pool of phosphatidylinositol 4-phosphate is generated by phosphatidylinositol 4-kinase type-III alpha: studies with the PH domains of the oxysterol binding protein and FAPP1. Mol.Biol. Cell 16, 1282–1295 (2005).
Shin, H.W. & Nakayama, K. Dual control of membrane targeting by PtdIns(4)P and ARF. Trends Biochem. Sci. 29, 513–515 (2004).
Honing, S. et al. Phosphatidylinositol-(4,5)-bisphosphate regulates sorting signal recognition by the clathrin-associated adaptor complex AP2. Mol. Cell 18, 519–531 (2005).
Yin, H.L. & Janmey, P.A. Phosphoinositide regulation of the actin cytoskeleton. Annu. Rev. Physiol. 65, 761–789 (2003).
Caroni, P. Actin cytoskeleton regulation through modulation of PI(4,5)P2 rafts. EMBO J. 20, 4332–4336 (2001).
van Rheenen, J., Achame, E. M., Janssen, H., Calafat, J. & Jalink, K. PIP2 signaling in lipid domains: a critical re-evaluation. EMBO J. 24, 1664–1673 (2005).
Murray, J.T., Panaretou, C., Stenmark, H., Miaczynska, M. & Backer, J.M. Role of Rab5 in the recruitment of hVps34/p150 to the early endosome. Traffic 3, 416–427 (2002).
Kihara, A., Noda, T., Ishihara, N. & Ohsumi, Y. Two distinct Vps34 phosphatidylinositol 3-kinase complexes function in autophagy and carboxypeptidase Y sorting in Saccharomyces cerevisiae. J. Cell Biol. 152, 519–530 (2001).
Stefan, C.J., Audhya, A. & Emr, S.D. The yeast synaptojanin-like proteins control the cellular distribution of phosphatidylinositol (4,5)-bisphosphate. Mol. Biol. Cell 13, 542–557 (2002).
Roy, A. & Levine, T.P. Multiple pools of phosphatidylinositol 4-phosphate detected using the pleckstrin homology domain of Osh2p. J. Biol. Chem. 279, 44683–44689 (2004).
Faulhammer, F. et al. Cell growth-dependent coordination of lipid signaling and glycosylation is mediated by interactions between Sac1p and Dpm1p. J. Cell Biol. 168, 185–191 (2005).
Ivetac, I. et al. The type Iα inositol polyphosphate 4-phosphatase generates and terminates phosphoinositide 3-kinase signals on endosomes and the plasma membrane. Mol. Biol. Cell 16, 2218–2233 (2005).
Munro, S., The Golgi apparatus: defining the identity of Golgi membranes. Curr. Opin. Cell Biol. 17, 395–401 (2005).
Murray, D. & Honig, B. Electrostatic control of the membrane targeting of C2 domains. Mol. Cell 9, 145–154 (2002).
Natarajan, P., Wang, J., Hua, Z. & Graham, T.R. Drs2p-coupled aminophospholipid translocase activity in yeast Golgi membranes and relationship to in vivo function. Proc. Natl Acad. Sci. USA 101, 10614–10619 (2004).
Pomorski, T. et al. Drs2p-related P-type ATPases Dnf1p and Dnf2p are required for phospholipid translocation across the yeast plasma membrane and serve a role in endocytosis. Mol. Biol. Cell 14, 1240–1254 (2003).
Ichimura, Y. et al. A ubiquitin-like system mediates protein lipidation. Nature 408, 488–492 (2000).
Bijlmakers, M.J. & Marsh, M. The on-off story of protein palmitoylation. Trends Cell Biol. 13, 32–42 (2003).
Smotrys, J.E. & Linder, M.E. Palmitoylation of intracellular signaling proteins: regulation and function. Annu. Rev. Biochem. 73, 559–587 (2004).
Drenan, R.M. et al. Palmitoylation regulates plasma membrane-nuclear shuttling of R7BP, a novel membrane anchor for the RGS7 family. J. Cell Biol. 169, 623–633 (2005).
Rocks, O. et al. An acylation cycle regulates localization and activity of palmitoylated Ras isoforms. Science 307, 1746–1752 (2005).
Salcedo, S.P. & Holden, D.W. Bacterial interactions with the eukaryotic secretory pathway. Curr. Opin. Microbiol. 8, 92–98 (2005).
Pizarro-Cerda, J. & Cossart, P. Subversion of phosphoinositide metabolism by intracellular bacterial pathogens. Nature Cell Biol. 6, 1026–1033 (2004).
Meresse, S. et al. Controlling the maturation of pathogen-containing vacuoles: a matter of life and death. Nature Cell Biol. 1, E183–E188 (1999).
Buttner, D. & Bonas, U. Port of entry — the type III secretion translocon. Trends Microbiol. 10, 186–192 (2002).
Zhou, D. & Galan, J. Salmonella entry into host cells: the work in concert of type III secreted effector proteins. Microbes Infect. 3, 1293–1298 (2001).
Gouin, E., Welch, M.D. & Cossart, P. Actin-based motility of intracellular pathogens. Curr. Opin. Microbiol. 8, 35–45 (2005).
Via, L.E. et al. Arrest of mycobacterial phagosome maturation is caused by a block in vesicle fusion between stages controlled by rab5 and rab7. J. Biol. Chem. 272, 13326–13331 (1997).
Vergne, I. et al. Mechanism of phagolysosome biogenesis block by viable Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 102, 4033–4038 (2005).
Hernandez, L.D., Hueffer, K., Wenk, M.R. & Galan, J.E. Salmonella modulates vesicular traffic by altering phosphoinositide metabolism. Science 304, 1805–1807 (2004).
Roy, C.R. Bacterial subversion of the host secretory pathway. Proc. Natl Acad. Sci. USA 102, 1271–1272 (2005).
Nagai, H., Kagan, J.C., Zhu, X., Kahn, R.A. & Roy, C.R. A bacterial guanine nucleotide exchange factor activates ARF on Legionella phagosomes. Science 295, 679–682 (2002).
Grieshaber, S.S., Grieshaber, N.A. & Hackstadt, T. Chlamydia trachomatis uses host cell dynein to traffic to the microtubule-organizing center in a p50 dynamitin-independent process. J. Cell Sci. 116, 3793–3802 (2003).
Guignot, J. et al. Microtubule motors control membrane dynamics of Salmonella-containing vacuoles. J. Cell Sci. 117, 1033–1045 (2004).
Harrison, R.E. et al. Salmonella impairs RILP recruitment to Rab7 during maturation of invasion vacuoles. Mol. Biol. Cell 15, 3146–3154 (2004).
Boucrot, E., Henry, T., Borg, J.P., Gorvel, J.P. & Meresse, S. The intracellular fate of Salmonella depends on the recruitment of kinesin. Science 308, 1174–1178 (2005).
Smith, G.A. & Enquist, L.W. Break ins and break outs: viral interactions with the cytoskeleton of mammalian cells. Annu. Rev. Cell Dev. Biol. 18, 135–161 (2002).
Belov, G.A., Fogg, M.H. & Ehrenfeld, E. Poliovirus proteins induce membrane association of GTPase ADP-ribosylation factor. J. Virol. 79, 7207–7216 (2005).
Loewen, C.J., Roy, A. & Levine, T.P. A conserved ER targeting motif in three families of lipid binding proteins and in Opi1p binds VAP. EMBO J. 22, 2025–2035 (2003).
Sollner, T. et al. SNAP receptors implicated in vesicle targeting and fusion. Nature 362, 318–324 (1993).
Banfield, D.K. SNARE complexes — is there sufficient complexity for vesicle targeting specificity? Trends Biochem. Sci. 26, 67–68 (2001).
Short, B., Haas, A. & Barr, F.A. Golgins and GTPases, giving identity and structure to the Golgi apparatus. Biochim. Biophys. Acta 1744, 383–395 (2005).
Acknowledgements
We are indebted to M. Freeman, A. Gillingham, P. Langridge and K. Röper for comments on the manuscript. R.B. is supported by the Cambridge European Trust.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Author Information Reprints and permissions information is available at npg.nature.com/reprintsandpermissions.
Rights and permissions
About this article
Cite this article
Behnia, R., Munro, S. Organelle identity and the signposts for membrane traffic. Nature 438, 597–604 (2005). https://doi.org/10.1038/nature04397
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04397
This article is cited by
-
Mechanisms of SNARE proteins in membrane fusion
Nature Reviews Molecular Cell Biology (2024)
-
Rab GTPases and phosphoinositides fine-tune SNAREs dependent targeting specificity of intracellular vesicle traffic
Nature Communications (2024)
-
Phosphoinositides as membrane organizers
Nature Reviews Molecular Cell Biology (2022)
-
Functional analyses of phosphatidylserine/PI(4)P exchangers with diverse lipid species and membrane contexts reveal unanticipated rules on lipid transfer
BMC Biology (2021)
-
The functional universe of membrane contact sites
Nature Reviews Molecular Cell Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.