Abstract
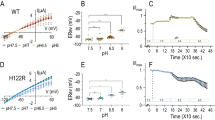
Although membrane proteins often rely on ionizable residues for structure and function, their ionization states under physiological conditions largely elude experimental estimation. To gain insight into the effect of the local microenvironment on the proton affinity of ionizable residues, we have engineered individual lysines, histidines and arginines along the α-helical lining of the transmembrane pore of the nicotinic acetylcholine receptor. We can detect individual proton binding–unbinding reactions electrophysiologically at the level of a single proton on a single side chain as brief blocking–unblocking events of the passing cation current. Kinetic analysis of these fluctuations yields the position-dependent rates of proton transfer, from which the corresponding pKa values and shifts in pKa can be calculated. Here we present a self-consistent, residue-by-residue description of the microenvironment around the pore-lining transmembrane α-helices (M2) in the open-channel conformation, in terms of the excess free energy that is required to keep the engineered basic side chains protonated relative to bulk water. A comparison with closed-channel data leads us to propose that the rotation of M2, which is frequently invoked as a hallmark of the gating mechanism of Cys-loop receptors, is minimal, if any.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Perutz, M. F. Electrostatic effects in proteins. Science 201, 1187–1191 (1978)
Warshel, A. Electrostatic basis of structure–function correlation in proteins. Acc. Chem. Res. 14, 284–290 (1981)
Davis, M. E. & McCammon, J. A. Electrostatics in biomolecular structure and dynamics. Chem. Rev. 90, 509–521 (1990)
Honig, B. & Nicholls, A. Classical electrostatics in biology and chemistry. Science 268, 1144–1149 (1995)
Imoto, K. et al. Rings of negatively charged amino acids determine the acetylcholine receptor channel conductance. Nature 335, 645–648 (1988)
Kelley, S. P., Dunlop, J. I., Kirkness, E. F. & Lambert, J. J. A cytoplasmic region determines single-channel conductance in 5-HT3 receptors. Nature 424, 321–324 (2003)
Heinemann, S. H., Terlau, H., Stühmer, W., Imoto, K. & Numa, S. Calcium channel characteristics conferred on the sodium channel by single mutations. Nature 356, 441–443 (1992)
Yang, J., Ellinor, P. T., Sather, W. A., Zhang, J. F. & Tsien, R. W. Molecular determinants of Ca2+ selectivity and ion permeation in L-type Ca2+ channels. Nature 366, 158–161 (1993)
Keramidas, A., Moorhouse, A. J., Pierce, K. D., Schofield, P. R. & Barry, P. H. Cation-selective mutations in the M2 domain of the inhibitory glycine receptor channel reveal determinants of ion-charge selectivity. J. Gen. Physiol. 119, 393–410 (2002)
Lu, Z. & MacKinnon, R. Electrostatic tuning of Mg2+ affinity in an inward-rectifier K+ channel. Nature 371, 243–246 (1994)
Khan, A., Romantseva, L., Lam, A., Lipkind, G. & Fozzard, H. A. Role of outer ring carboxylates of the rat skeletal muscle sodium channel pore in proton block. J. Physiol. Lond. 543, 71–84 (2002)
Schulte, U. & Fakler, B. Gating of inward-rectifier K+ channels by intracellular pH. Eur. J. Biochem. 267, 5837–5841 (2000)
Aggarwal, S. K. & MacKinnon, R. Contribution of the S4 segment to gating charge in the Shaker K+ channel. Neuron 16, 1169–1177 (1996)
Seoh, S.-A., Sigg, D., Papazian, D. M. & Bezanilla, F. Voltage-sensing residues in the S2 and S4 segments of the Shaker K+ channel. Neuron 16, 1159–1167 (1996)
Schutz, C. N. & Warshel, A. What are the dielectric ‘constants’ of proteins and how to validate electrostatic models? Proteins 44, 400–417 (2001)
Mehler, E. L., Fuxreiter, M., Simon, I. & García-Moreno, E. B. The role of hydrophobic microenvironments in modulating pKa shifts in proteins. Proteins 48, 283–292 (2002)
Parzefall, F., Wilhelm, R., Heckmann, M. & Dudel, J. Single channel currents at six microsecond resolution elicited by acetylcholine in mouse myoballs. J. Physiol. Lond. 512, 181–188 (1998)
Dao-pin, S., Anderson, D. E., Baase, W. A., Dahlquist, F. W. & Matthews, B. W. Structural and thermodynamic consequences of burying a charged residue within the hydrophobic core of T4 lysozyme. Biochemistry 30, 11521–11529 (1991)
Prod'hom, B., Pietrobon, D. & Hess, P. Direct measurement of proton transfer rates to a group controlling the dihydropyridine-sensitive Ca2+ channel. Nature 329, 243–246 (1987)
Root, M. J. & MacKinnon, R. Two identical noninteracting sites in an ion channel revealed by proton transfer. Science 265, 1852–1856 (1994)
Lester, H. The permeation pathway of neurotransmitter-gated ion channels. Annu. Rev. Biophys. Biomol. Struct. 21, 267–292 (1992)
Unwin, N. Acetylcholine receptor channel imaged in the open state. Nature 373, 37–43 (1995)
Miyazawa, A., Fujiyoshi, Y. & Unwin, N. Structure and gating mechanism of the acetylcholine receptor pore. Nature 423, 949–955 (2003)
Horenstein, J., Wagner, D. A., Czajkowski, C. & Akabas, M. H. Protein mobility and GABA-induced conformational changes in GABAA receptor pore-lining M2 segment. Nature Neurosci. 4, 477–485 (2001)
Corringer, P. J. et al. Mutational analysis of the charge selectivity filter of the α7 nicotinic acetylcholine receptor. Neuron 22, 831–843 (1999)
Adcock, C., Smith, G. R. & Sansom, M. S. P. Electrostatics and the ion selectivity of ligand-gated channels. Biophys. J. 75, 1211–1222 (1998)
Stopar, D., Sprujit, R. B., Wolfs, C. J. & Hemminga, M. A. Local dynamics of the M13 major coat protein in different membrane-mimicking systems. Biochemistry 35, 15467–15473 (1996)
Grosman, C., Zhou, M. & Auerbach, A. Mapping the conformational wave of acetylcholine receptor channel gating. Nature 403, 773–776 (2000)
Cymes, G. D., Grosman, C. & Auerbach, A. Structure of the transition state of gating in the acetylcholine receptor channel pore: a φ-value analysis. Biochemistry 41, 5548–5555 (2002)
Grosman, C. Free-energy landscapes of ion-channel gating are malleable: changes in the number of bound ligands are accompanied by changes in the location of the transition state in acetylcholine-receptor channels. Biochemistry 42, 14977–14987 (2003)
Goren, E. N., Reeves, D. C. & Akabas, M. H. Loose protein packing around the extracellular half of the GABAA receptor β1 subunit M2 channel-lining segment. J. Biol. Chem. 279, 11198–11205 (2004)
Kim, S., Chamberlain, A. K. & Bowie, J. U. A model of the closed form of the nicotinic acetylcholine receptor M2 channel pore. Biophys. J. 87, 792–799 (2004)
Dahan, D. et al. A fluorophore attached to nicotinic acetylcholine receptor βM2 detects productive binding of agonist to the αδ site. Proc. Natl Acad. Sci. USA 101, 10195–10200 (2004)
Wilson, G. G. & Karlin, A. The location of the gate in the acetylcholine receptor channel. Neuron 20, 1269–1281 (1998)
White, B. H. & Cohen, J. B. Agonist-induced changes in the structure of the acetylcholine receptor M2 regions revealed by photoincorporation of an uncharged nicotinic noncompetitive antagonist. J. Biol. Chem. 267, 15770–15783 (1992)
Arévalo, E., Chiara, D. C., Forman, S. A., Cohen, J. B. & Miller, K. W. Gating-enhanced accessibility of hydrophobic sites within the transmembrane region of the nicotinic acetylcholine receptor's δ subunit. A time-resolved photolabeling study. J. Biol. Chem. 280, 13631–13640 (2005)
O'Mara, M., Barry, P. H. & Chung, S. H. A model of the glycine receptor deduced from Brownian dynamics. Proc. Natl Acad. Sci. USA 100, 4310–4315 (2003)
Corry, B., Vora, T. & Chung, S. H. Electrostatic basis of valence selectivity in cationic channels. Biochim. Biophys. Acta 1711, 72–86 (2005)
Akabas, M. H., Kaufmann, C., Archdeacon, P. & Karlin, A. Identification of acetylcholine receptor channel-lining residues in the entire M2 segment of the α subunit. Neuron 13, 919–927 (1994)
Qin, F. Restoration of single-channel currents using the segmental k-means method based on hidden Markov modeling. Biophys. J. 86, 1488–1501 (2004)
Qin, F., Auerbach, A. & Sachs, F. Estimating single-channel kinetic parameters from idealized patch-clamp data containing missed events. Biophys. J. 70, 264–280 (1996)
Ohno, K. et al. Congenital myasthenic syndrome caused by prolonged acetylcholine receptor channel openings due to a mutation in the M2 domain of the epsilon subunit. Proc. Natl Acad. Sci. USA 92, 758–762 (1995)
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graphics 14, 33–38 (1996)
Acknowledgements
We thank S. Sine for wild-type muscle AChR subunit cDNA; J. Jasielec and J. Gasser for technical assistance; S. Varma and B. García-Moreno E. for discussions; and E. Tajkhorshid for introducing us to the VMD imaging program. This work was supported by a grant from the NIH (to C.G.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains Supplementary Methods, Supplementary Discussion, Supplementary Figures 1–3 and accompanying Supplementary Figure Legends. (PDF 562 kb)
Rights and permissions
About this article
Cite this article
Cymes, G., Ni, Y. & Grosman, C. Probing ion-channel pores one proton at a time. Nature 438, 975–980 (2005). https://doi.org/10.1038/nature04293
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04293
This article is cited by
-
Subunit gating resulting from individual protonation events in Kir2 channels
Nature Communications (2023)
-
Structural mechanisms of activation and desensitization in neurotransmitter-gated ion channels
Nature Structural & Molecular Biology (2016)
-
Side chain flexibility and the pore dimensions in the GABAA receptor
Journal of Computer-Aided Molecular Design (2016)
-
Crystal structure of a human GABAA receptor
Nature (2014)
-
The unanticipated complexity of the selectivity-filter glutamates of nicotinic receptors
Nature Chemical Biology (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.