Abstract
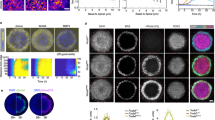
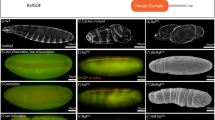
A central question in the development of multicellular organisms pertains to the timing and mechanisms of specification of the embryonic axes. In many organisms, specification of the dorsoventral axis requires signalling by proteins of the Transforming growth factor-β and Wnt families1,2,3. Here we show that maternal transcripts of the zebrafish Nodal-related morphogen, Squint (Sqt), can localize to two blastomeres at the four-cell stage and predict the dorsal axis. Removal of cells containing sqt transcripts from four-to-eight-cell embryos or injection of antisense morpholino oligonucleotides targeting sqt into oocytes can cause a loss of dorsal structures. Localization of sqt transcripts is independent of maternal Wnt pathway function and requires a highly conserved sequence in the 3′ untranslated region. Thus, the dorsoventral axis is apparent by early cleavage stages and may require the maternally encoded morphogen Sqt and its associated factors. Because the 3′ untranslated region of the human nodal gene can also localize exogenous sequences to dorsal cells, this mechanism may be evolutionarily conserved.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
12 December 2012
Nature 438, 1030–1035 (2005); doi:10.1038/nature04184 In Fig. 3p and q and Supplementary Table 3 of this Letter, the data for sqt MO2 and sqt MO3 were inadvertently mislabelled and should be swapped. The text on page 1033 that describes Fig. 3p and q should therefore read ‘sqt MO2’ not ‘sqt MO3’ on lines 4 and 13 of the second full paragraph of the left column.
References
Moon, R. T. & Kimelman, D. From cortical rotation to organizer gene expression: toward a molecular explanation of axis specification in Xenopus. BioEssays 20, 536–545 (1998)
Sokol, S. Y. Wnt signalling and dorso-ventral axis specification in vertebrates. Curr. Opin. Genet. Dev. 9, 405–410 (1999)
Holley, S. A. & Ferguson, E. L. Fish are like flies are like frogs: conservation of dorsal–ventral patterning mechanisms. BioEssays 19, 281–284 (1997)
Riechmann, V. & Ephrussi, A. Axis formation during Drosophila oogenesis. Curr. Opin. Genet. Dev. 11, 374–383 (2001)
Heasman, J. Patterning the Xenopus blastula. Development 124, 4179–4191 (1997)
Schier, A. F. & Talbot, W. S. Molecular genetics of axis formation in zebrafish. Annu. Rev. Genet. published online 9 August 2005 (doi:10.1146/annurev.genet.37.110801.143752) .
Weaver, C. & Kimelman, D. Move it or lose it: axis specification in Xenopus. Development 131, 3491–3499 (2004)
Jesuthasan, S. & Strahle, U. Dynamic microtubules and specification of the zebrafish embryonic axis. Curr. Biol. 7, 31–42 (1997)
Mizuno, T., Yamaha, E. & Yamazaki, F. Localized axis determinant in the early cleavage embryo of goldfish, Carassius auratus. Dev. Growth Differ. 206, 389–396 (1997)
Oppenheimer, J. The development of isolated blastoderms of Fundulus heteroclitus. J. Exp. Zool. 72, 247–269 (1936)
Rebagliati, M. R., Toyama, R., Fricke, C., Haffter, P. & Dawid, I. B. Zebrafish nodal-related genes are implicated in axial patterning and establishing left–right asymmetry. Dev. Biol. 199, 261–272 (1998)
Erter, C. E., Solnica-Krezel, L. & Wright, C. V. Zebrafish nodal-related 2 encodes an early mesendodermal inducer signalling from the extraembryonic yolk syncytial layer. Dev. Biol. 204, 361–372 (1998)
Feldman, B. et al. Zebrafish organizer development and germ-layer formation require nodal-related signals. Nature 395, 181–185 (1998)
Gore, A. V. & Sampath, K. Localization of transcripts of the zebrafish morphogen Squint is dependent on egg activation and the microtubule cytoskeleton. Mech. Dev. 112, 153–156 (2002)
Klein, S. L. The first cleavage furrow demarcates the dorsal–ventral axis in Xenopus embryos. Dev. Biol. 120, 299–304 (1987)
Feng, B., Schwarz, H. & Jesuthasan, S. Furrow-specific endocytosis during cytokinesis of zebrafish blastomeres. Exp. Cell Res. 279, 14–20 (2002)
Lambert, J. D. & Nagy, L. M. Asymmetric inheritance of centrosomally localized mRNAs during embryonic cleavages. Nature 420, 682–686 (2002)
Yamanaka, Y. et al. A novel homeobox gene, dharma, can induce the organizer in a non-cell-autonomous manner. Genes Dev. 12, 2345–2353 (1998)
Dougan, S. T., Warga, R. M., Kane, D. A., Schier, A. F. & Talbot, W. S. The role of the zebrafish nodal-related genes squint and cyclops in patterning of mesendoderm. Development 130, 1837–1851 (2003)
Kishimoto, Y., Lee, K. H., Zon, L., Hammerschmidt, M. & Schulte-Merker, S. The molecular nature of zebrafish swirl: BMP2 function is essential during early dorsoventral patterning. Development 124, 4457–4466 (1997)
Feldman, B. & Stemple, D. L. Morpholino phenocopies of sqt, oep, and ntl mutations. Genesis 30, 175–177 (2001)
Tao, Q. et al. Maternal wnt11 activates the canonical wnt signalling pathway required for axis formation in Xenopus embryos. Cell 120, 857–871 (2005)
Kelly, C., Chin, A. J., Leatherman, J. L., Kozlowski, D. J. & Weinberg, E. S. Maternally controlled β-catenin-mediated signalling is required for organizer formation in the zebrafish. Development 127, 3899–3911 (2000)
Abdelilah, S., Solnica-Krezel, L., Stainier, D. Y. & Driever, W. Implications for dorsoventral axis determination from the zebrafish mutation janus. Nature 370, 468–471 (1994)
Helde, K. A., Wilson, E. T., Cretekos, C. J. & Grunwald, D. J. Contribution of early cells to the fate map of the zebrafish gastrula. Science 265, 517–520 (1994)
Zernicka-Goetz, M. Patterning of the embryo: the first spatial decisions in the life of a mouse. Development 129, 815–829 (2002)
Bullock, S. L. & Ish-Horowicz, D. Conserved signals and machinery for RNA transport in Drosophila oogenesis and embryogenesis. Nature 414, 611–616 (2001)
Sampath, K. et al. Induction of the zebrafish ventral brain and floorplate requires cyclops/nodal signalling. Nature 395, 185–189 (1998)
Muller, F., Lele, Z., Varadi, L., Menczel, L. & Orban, L. Efficient transient expression system based on square pulse electroporation and in vivo luciferase assay of fertilized fish eggs. FEBS Lett. 324, 27–32 (1993)
Higgins, D. G., Bleasby, A. J. & Fuchs, R. CLUSTAL V: improved software for multiple sequence alignment. Comput. Appl. Biosci. 8, 189–191 (1992)
Acknowledgements
We thank M. Balasubramanian, R. Dosch, R. Dunn, S. Jesuthasan, E. Raz, S. Roy, A. Schier, and members of the Sampath laboratory for discussions and suggestions; S. Koshida, E. Yamaha and H. Takeda for embryo operation techniques; A. Chang and the Qian Hu fish farm for cyprinids; H. Ngoc Bao Quach for technical assistance; M. Hidayat for sequencing; and J. Bradner, C. Hu and C. Goh and the TLL fish facility for fish care. This work was supported by NIH grants (E.W.) and by TLL, Singapore (K.S.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The cyprinid sqt sequences have been deposited in GenBank (accession numbers DQ080241, DQ080242 and DQ080243). Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figure 1
Real-Time PCR analysis of sqt transcripts (a) and ef1-α (b) at various stages of development. (PDF 354 kb)
Supplementary Figure 2
Colocalization of β-catenin with β-Gal human nodal 3’UTR. (PDF 76 kb)
Supplementary Table 1
Percentage embryos expressing localized sqt through cleavage, blastula and early gastrula stages. (DOC 20 kb)
Supplementary Table 2
Marker gene expression in operated embryos. (DOC 24 kb)
Supplementary Table 3
Marker gene expression in morpholino injected oocytes or fertilized embryos (DOC 28 kb)
Supplementary Video 1
Dynamic localization of fluorescent sqt RNA in zebrafish embryos at early cleavage stages. (MOV 2570 kb)
Supplementary Video 2
Dynamic localization of fluorescent sqt RNA in zebrafish embryos at early cleavage stages. (AVI 517 kb)
Supplementary Video 3
Uniform distribution of fluorescent lacZ RNA with the β-globin 3’ UTR. (MOV 976 kb)
Supplementary Video 4
Localization of fluorescent sqt RNA is inhibited by the microtubule poison, nocodazole. (MOV 3388 kb)
Supplementary Video 5
The 3’UTR of sqt RNA is required for localization. The embryo was injected with sqt RNA lacking its 3’UTR. (AVI 3076 kb)
Supplementary Video 6
Exogenous 3’UTRs do not localize sqt RNA. The embryo was injected with labelled sqt coding region RNA fused with the β-globin 3’UTR. (MOV 1176 kb)
Supplementary Video 7
Fluorescent lacZ RNA fused to sqt 3’UTR is localized by early cleavage stages. (MOV 3337 kb)
Supplementary Video 8
Fluorescent lacZ RNA fused to sqt 3’UTR is localized by early cleavage stages. (MOV 2176 kb)
Supplementary Video 9
Fluorescent sqt RNA with the first 50 nucleotides of its 3’UTR is localized by the 4-cell stage (MOV 2363 kb)
Supplementary Video 10
Fluorescent sqt RNA with the first 50 nucleotides of its 3’UTR is localized by the 4-cell stage. (MOV 3411 kb)
Supplementary Video 11
Fluorescent lacZ RNA with the human nodal 3’UTR is localized by the 4-cell stage. (MOV 2774 kb)
Supplementary Video 12
Fluorescent lacZ RNA with the human nodal 3’UTR is localized by the 4-cell stage. Lateral view shown. (AVI 656 kb)
Supplementary Legends
Text to accompany the above Supplementary Figures and Supplementary videos. (DOC 24 kb)
Rights and permissions
About this article
Cite this article
Gore, A., Maegawa, S., Cheong, A. et al. The zebrafish dorsal axis is apparent at the four-cell stage. Nature 438, 1030–1035 (2005). https://doi.org/10.1038/nature04184
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04184
This article is cited by
-
Dynamics of maternal gene expression in Rhodnius prolixus
Scientific Reports (2022)
-
MiR-430 Can Affect the Mesoderm Formation and Metamorphosis of Paralichthys olivaceus by Targeting lefty Gene
Journal of Ocean University of China (2020)
-
Correction: Corrigendum: The zebrafish dorsal axis is apparent at the four-cell stage
Nature (2013)
-
Osteogenic transcription factor Runx2 is a maternal determinant of dorsoventral patterning in zebrafish
Nature Cell Biology (2008)
-
Maternal nodal and zebrafish embryogenesis
Nature (2007)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.