Abstract
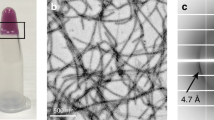
Amyloid or amyloid-like fibrils are elongated, insoluble protein aggregates, formed in vivo1 in association with neurodegenerative diseases or in vitro2 from soluble native proteins, respectively. The underlying structure of the fibrillar or ‘cross-β’ state has presented long-standing, fundamental puzzles of protein structure. These include whether fibril-forming proteins have two structurally distinct stable states, native and fibrillar, and whether all or only part of the native protein refolds as it converts to the fibrillar state. Here we show that a designed amyloid-like fibril of the well-characterized enzyme RNase A contains native-like molecules capable of enzymatic activity. In addition, these functional molecular units are formed from a core RNase A domain and a swapped complementary domain. These findings are consistent with the zipper-spine model3 in which a cross-β spine is decorated with three-dimensional domain-swapped functional units, retaining native-like structure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Koo, E. H., Lansbury, P. T. Jr & Kelly, J. W. Amyloid diseases: abnormal protein aggregation in neurodegeneration. Proc. Natl Acad. Sci. USA 96, 9989–9990 (1999)
Dobson, C. M. Protein misfolding, evolution and disease. Trends Biochem. Sci. 24, 329–332 (1999)
Liu, Y., Gotte, G., Libonati, M. & Eisenberg, D. A domain-swapped RNase A dimer with implications for amyloid formation. Nature Struct. Biol. 8, 211–214 (2001)
Sunde, M. et al. Common core structure of amyloid fibrils by synchrotron X-ray diffraction. J. Mol. Biol. 273, 729–739 (1997)
Anfinsen, C. B. Principles that govern the folding of protein chains. Science 181, 223–230 (1973)
Cohen, A. A. & Calkins, E. Electron microscopic observations on a fibrous component in amyloid of diverse origins. Nature 183, 1202–1203 (1959)
Jimenez, J. L. et al. Cryo-electron microscopy structure of an SH3 amyloid fibril and model of the molecular packing. EMBO J. 18, 815–821 (1999)
Jimenez, J. L. et al. The protofilament structure of insulin amyloid fibrils. Proc. Natl Acad. Sci. USA 99, 9196–9201 (2002)
Kishimoto, A. et al. β-Helix is a likely core structure of yeast prion Sup amyloid fibrils. Biochem. Biophys. Res. Commun. 315, 739–745 (2004)
Govaerts, C., Wille, H., Prusiner, S. B. & Cohen, F. E. Evidence for assembly of prions with left-handed β-helices into trimers. Proc. Natl Acad. Sci. USA 101, 8342–8347 (2004)
Perutz, M. F., Finch, J. T., Berriman, J. & Lesk, A. Amyloid fibrils are water-filled nanotubes. Proc. Natl Acad. Sci. USA 99, 5591–5595 (2002)
Pickersgill, R. W. A primordial structure underlying amyloid. Structure 11, 137–138 (2003)
Wetzel, R. Ideas of order for amyloid fibril structure. Structure 10, 1031–1036 (2002)
Eakin, C. M., Attenello, F. J., Morgan, C. J. & Miranker, A. D. Oligomeric assembly of native-like precursors precedes amyloid formation by β-2 microglobulin. Biochemistry 43, 7808–7815 (2004)
Schlunegger, M. P., Bennett, M. J. & Eisenberg, D. Oligomer formation by 3D domain swapping: a model for protein assembly and misassembly. Adv. Protein Chem. 50, 61–122 (1997)
Janowski, R. et al. Human cystatin C, an amyloidogenic protein, dimerizes through three-dimensional domain swapping. Nature Struct. Biol. 8, 316–320 (2001)
Nilsson, M. et al. Prevention of domain swapping inhibits dimerization and amyloid fibril formation of cystatin C: use of engineered disulfide bridges, antibodies, and carboxymethylpapain to stabilize the monomeric form of cystatin C. J. Biol. Chem. 279, 24236–24245 (2004)
Sanders, A. et al. Cystatin forms a tetramer through structural rearrangement of domain-swapped dimers prior to amyloidogenesis. J. Mol. Biol. 336, 165–178 (2004)
Staniforth, R. A. et al. Three-dimensional domain swapping in the folded and molten-globule states of cystatins, an amyloid-forming structural superfamily. EMBO J. 20, 4774–4781 (2001)
Sela, M., White, F. H. & Anfinsen, C. B. Reductive cleavage of disulfide bridges in ribonuclease. Science 125, 691–692 (1957)
Crestfield, A. M., Stein, W. H. & Moore, S. Properties and conformation of the histidine residues at the active site of ribonuclease. J. Biol. Chem. 228, 2421–2428 (1963)
Gotte, G., Bertoldi, M. & Libonati, M. Structural versatility of bovine ribonuclease A. Distinct conformers of trimeric and tetrameric aggregates of the enzyme. Eur. J. Biochem. 265, 680–687 (1999)
Nelson, R. et al. Structure of the amyloid spine. Nature 435, 773–777 (2005)
Liu, Y. & Eisenberg, D. 3D domain swapping: as domains continue to swap. Protein Sci. 11, 1285–1299 (2002)
Booth, D. R. et al. Instability, unfolding and aggregation of human lysozyme variants underlying amyloid fibrillogenesis. Nature 385, 787–793 (1997)
Bennett, M. J., Schlunegger, M. P. & Eisenberg, D. 3D domain swapping: a mechanism for oligomer assembly. Protein Sci. 4, 2455–2468 (1995)
Acknowledgements
We thank M. Phillips for electron microscopy, M. Bennett, T. Yeates and D. H. Anderson for discussion, and M. Nebohacova and L. Simpson for assistance with blue-native gels. This study received financial support from the NIH, NSF and the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The structure of Q10-H119A RNase A has been deposited in the Protein Data Bank with accession code 2APQ. The model of Fig. 4 has been deposited in the Protein Data Bank with accession code 2APU. Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file includes Supplementary Methods, additional control experiments and legends for Supplementary Table S1, Supplementary Figure S1 and Supplementary Figure S2. (DOC 1824 kb)
Supplementary Table S2
Data Collection for Q10-H119A (DOC 28 kb)
Supplementary Figure S1
Cross- diffraction from partially oriented Q10-H119A RNase A fibrils. (JPG 62 kb)
Supplementary Figure S2
Silver-stained, non-denaturing gel comparing lyophilized Q10 -expanded RNase A with lyophilized wild-type RNase A. (JPG 79 kb)
Rights and permissions
About this article
Cite this article
Sambashivan, S., Liu, Y., Sawaya, M. et al. Amyloid-like fibrils of ribonuclease A with three-dimensional domain-swapped and native-like structure. Nature 437, 266–269 (2005). https://doi.org/10.1038/nature03916
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03916
This article is cited by
-
Breakdown of supersaturation barrier links protein folding to amyloid formation
Communications Biology (2021)
-
Nanoscale Dynamics of Protein Assembly Networks in Supersaturated Solutions
Scientific Reports (2017)
-
Domain-swap polymerization drives the self-assembly of the bacterial flagellar motor
Nature Structural & Molecular Biology (2016)
-
Amyloid formation of growth hormone in presence of zinc: Relevance to its storage in secretory granules
Scientific Reports (2016)
-
The C-terminus hot spot region helps in the fibril formation of bacteriophage-associated hyaluronate lyase (HylP2)
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.