Abstract
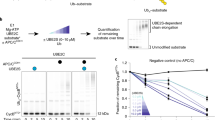
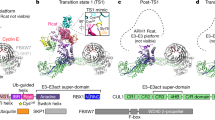
SUMO-1 (for small ubiquitin-related modifier) belongs to the ubiquitin (Ub) and ubiquitin-like (Ubl) protein family. SUMO conjugation occurs on specific lysine residues within protein targets, regulating pathways involved in differentiation, apoptosis, the cell cycle and responses to stress by altering protein function through changes in activity or cellular localization or by protecting substrates from ubiquitination1,2. Ub/Ubl conjugation occurs in sequential steps and requires the concerted action of E2 conjugating proteins and E3 ligases1,2. In addition to being a SUMO E3, the nucleoporin Nup358/RanBP2 localizes SUMO-conjugated RanGAP1 to the cytoplasmic face of the nuclear pore complex by means of interactions in a complex that also includes Ubc9, the SUMO E2 conjugating protein3,4,5,6. Here we describe the 3.0-Å crystal structure of a four-protein complex of Ubc9, a Nup358/RanBP2 E3 ligase domain (IR1-M) and SUMO-1 conjugated to the carboxy-terminal domain of RanGAP1. Structural insights, combined with biochemical and kinetic data obtained with additional substrates, support a model in which Nup358/RanBP2 acts as an E3 by binding both SUMO and Ubc9 to position the SUMO–E2-thioester in an optimal orientation to enhance conjugation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Johnson, E. S. Protein modification by SUMO. Annu. Rev. Biochem. 73, 355–382 (2004)
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998)
Matunis, M. J., Coutavas, E. & Blobel, G. A novel ubiquitin-like modification modulates the partitioning of the Ran-GTPase-activating protein RanGAP1 between the cytosol and the nuclear pore complex. J. Cell Biol. 135, 1457–1470 (1996)
Mahajan, R., Delphin, C., Guan, T., Gerace, L. & Melchior, F. A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell 88, 97–107 (1997)
Saitoh, H., Pu, R., Cavenagh, M. & Dasso, M. RanBP2 associates with Ubc9p and a modified form of RanGAP1. Proc. Natl Acad. Sci. USA 94, 3736–3741 (1997)
Zhang, H., Saitoh, H. & Matunis, M. J. Enzymes of the SUMO modification pathway localize to filaments of the nuclear pore complex. Mol. Cell. Biol. 22, 6498–6508 (2002)
Deshaies, R. J. SCF and Cullin/Ring H2-based ubiquitin ligases. Annu. Rev. Cell Dev. Biol. 15, 435–467 (1999)
Huibregtse, J. M., Scheffner, M., Beaudenon, S. & Howley, P. M. A family of proteins structurally and functionally related to the E6-AP ubiquitin-protein ligase. Proc. Natl Acad. Sci. USA 92, 2563–2567 (1995)
Bernier-Villamor, V., Sampson, D. A., Matunis, M. J. & Lima, C. D. Structural basis for E2-mediated SUMO conjugation revealed by a complex between ubiquitin-conjugating enzyme Ubc9 and RanGAP1. Cell 108, 345–356 (2002)
Johnson, E. S. & Gupta, A. A. An E3-like factor that promotes SUMO conjugation to the yeast septins. Cell 106, 735–744 (2001)
Kahyo, T., Nishida, T. & Yasuda, H. Involvement of PIAS1 in the sumoylation of tumor suppressor p53. Mol. Cell 8, 713–718 (2001)
Pichler, A., Gast, A., Seeler, J. S., Dejean, A. & Melchior, F. The nucleoporin RanBP2 has SUMO1 E3 ligase activity. Cell 108, 109–120 (2002)
Kagey, M. H., Melhuish, T. A. & Wotton, D. The polycomb protein Pc2 is a SUMO E3. Cell 113, 127–137 (2003)
Yokoyama, N. et al. A giant nucleopore protein that binds Ran/TC4. Nature 376, 184–188 (1995)
Wu, J., Matunis, M. J., Kraemer, D., Blobel, G. & Coutavas, E. Nup358, a cytoplasmically exposed nucleoporin with peptide repeats, Ran-GTP binding sites, zinc fingers, a cyclophilin A homologous domain, and a leucine-rich region. J. Biol. Chem. 270, 14209–14213 (1995)
Joseph, J., Liu, S. T., Jablonski, S. A., Yen, T. J. & Dasso, M. The RanGAP1-RanBP2 complex is essential for microtubule-kinetochore interactions in vivo . Curr. Biol. 14, 611–617 (2004)
Saitoh, H., Pizzi, M. D. & Wang, J. Perturbation of SUMOlation enzyme Ubc9 by distinct domain within nucleoporin RanBP2/Nup358. J. Biol. Chem. 277, 4755–4763 (2002)
Pichler, A., Knipscheer, P., Saitoh, H., Sixma, T. K. & Melchior, F. The RanBP2 SUMO E3 ligase is neither HECT nor RING type. Nature Struct. Mol. Biol. 11, 984–991 (2004)
Tatham, M. H. et al. Unique binding interactions among Ubc9, SUMO and RanBP2 reveal a mechanism for SUMO paralog selection. Nature Struct. Mol. Biol. 12, 67–74 (2004)
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991)
Lois, L. M. & Lima, C. D. Structures of the Small Ubiquitin-like MOdifier E1 activating enzyme provide insights into SUMO activation and the mechanism for E2 recruitment to E1. EMBO J. 24, 439–451 (2005)
Hamilton, K. S. et al. Structure of a conjugating enzyme-ubiquitin thiolester intermediate reveals a novel role for the ubiquitin tail. Structure 9, 897–904 (2001)
Wu, P. Y. et al. A conserved catalytic residue in the ubiquitin-conjugating enzyme family. EMBO J. 22, 5241–5250 (2003)
Song, J., Durrin, L. K., Wilkinson, T. A., Krontiris, T. G. & Chen, Y. Identification of a SUMO-binding motif that recognizes SUMO-modified proteins. Proc. Natl Acad. Sci. USA 101, 14373–14378 (2004)
Hannich, J. T. et al. Defining the SUMO-modified proteome by multiple approaches in Saccharomyces cerevisiae . J. Biol. Chem. 280, 4102–4110 (2005)
Huang, L. et al. Structure of an E6AP-UbcH7 complex: insights into ubiquitination by the E2–E3 enzyme cascade. Science 286, 1321–1326 (1999)
Leverson, J. D. et al. The APC11 RING-H2 finger mediates E2-dependent ubiquitination. Mol. Biol. Cell 11, 2315–2325 (2000)
Pickart, C. M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001)
Strickland, S., Palmer, G. & Massey, V. Determination of dissociation constants and specific rate constants of enzyme–substrate (or protein–ligand) interactions from rapid reaction kinetic data. J. Biol. Chem. 250, 4048–4052 (1975)
Acknowledgements
We thank M. J. Matunis for the original clone containing Nup358/RanBP2 (residues 2,596–2,836), and K. R. Rajashankar and A. Yunus for discussion and for reagents that contributed to this work. Use of the Advanced Photon Source (APS) is supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences. Use of the SGX Collaborative Access Team beamline facilities at Sector 31 of the APS was provided by Structural GenomiX, Inc., which constructed and operates the facility. D.R. and C.D.L. were supported in part by a National Institutes of Health grant. C.D.L. acknowledges support from the Rita Allen Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Coordinates have been deposited with the Protein Data Bank under accession number 1Z5S. Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Table S1
Crystallographic data, phasing, and refinement statistics. (DOC 53 kb)
Supplementary Figure S1
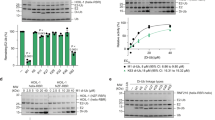
Activities of the Nup358/RanBP2 E3 with RanGAP1 and RanGAP1 (F562A). a) Activity under multiple turnover conditions with RanGAP1 and RanGAP1(F562A). b) Activity under single turnover conditions with RanGAP1 and RanGAP1(F562A). (PDF 61 kb)
Supplementary Figure S2
Biochemical activities of Nup358/RanBP2 with SUMO-2 and SUMO-3. a) For SUMO-2 conjugation. b) For SUMO-3 conjugation. c) Table of rate constants and relative rates for reactions in a and b. (PDF 29 kb)
Rights and permissions
About this article
Cite this article
Reverter, D., Lima, C. Insights into E3 ligase activity revealed by a SUMO–RanGAP1–Ubc9–Nup358 complex. Nature 435, 687–692 (2005). https://doi.org/10.1038/nature03588
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03588
This article is cited by
-
Mechanisms and functions of SUMOylation in health and disease: a review focusing on immune cells
Journal of Biomedical Science (2024)
-
Structural basis for the E3 ligase activity enhancement of yeast Nse2 by SUMO-interacting motifs
Nature Communications (2021)
-
Viruses, SUMO, and immunity: the interplay between viruses and the host SUMOylation system
Journal of NeuroVirology (2021)
-
Arkadia/RNF111 is a SUMO-targeted ubiquitin ligase with preference for substrates marked with SUMO1-capped SUMO2/3 chain
Nature Communications (2019)
-
Active Site Gate Dynamics Modulate the Catalytic Activity of the Ubiquitination Enzyme E2-25K
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.