Abstract
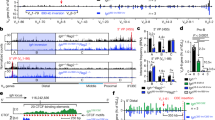
The T-helper-cell 1 and 2 (TH1 and TH2) pathways, defined by cytokines interferon-γ (IFN-γ) and interleukin-4 (IL-4), respectively, comprise two alternative CD4+ T-cell fates, with functional consequences for the host immune system. These cytokine genes are encoded on different chromosomes. The recently described TH2 locus control region (LCR) coordinately regulates the TH2 cytokine genes by participating in a complex between the LCR and promoters of the cytokine genes Il4, Il5 and Il13. Although they are spread over 120 kilobases, these elements are closely juxtaposed in the nucleus in a poised chromatin conformation. In addition to these intrachromosomal interactions, we now describe interchromosomal interactions between the promoter region of the IFN-γ gene on chromosome 10 and the regulatory regions of the TH2 cytokine locus on chromosome 11. DNase I hypersensitive sites that comprise the TH2 LCR developmentally regulate these interchromosomal interactions. Furthermore, there seems to be a cell-type-specific dynamic interaction between interacting chromatin partners whereby interchromosomal interactions are apparently lost in favour of intrachromosomal ones upon gene activation. Thus, we provide an example of eukaryotic genes located on separate chromosomes associating physically in the nucleus via interactions that may have a function in coordinating gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Cremer, T. & Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nature Rev. Genet. 2, 292–301 (2001)
Spector, D. L. The dynamics of chromosome organization and gene regulation. Annu. Rev. Biochem. 72, 573–608 (2003)
Brown, K. E. et al. Association of transcriptionally silent genes with Ikaros complexes at centromeric heterochromatin. Cell 91, 845–854 (1997)
Brown, K. E., Baxter, J., Graf, D., Merkenschlager, M. & Fisher, A. G. Dynamic repositioning of genes in the nucleus of lymphocytes preparing for cell division. Mol. Cell 3, 207–217 (1999)
Grogan, J. L. et al. Early transcription and silencing of cytokine genes underlie polarization of T helper cell subsets. Immunity 14, 205–215 (2001)
Ragoczy, T., Telling, A., Sawado, T., Groudine, M. & Koxak, S. T. A genetic analysis of chromosome territory looping: diverse roles for distal regulatory elements. Chromosome Res. 11, 513–525 (2003)
Francastel, C., Walters, M. C., Groudine, M. & Martin, D. I. A functional enhancer suppresses silencing of a transgene and prevents its localization close to centromeric heterochromatin. Cell 99, 259–269 (1999)
Schubeler, D. et al. Nuclear localization and histone acetylation: a pathway for chromatin opening and transcriptional activation of the human β-globin locus. Genes Dev. 14, 940–950 (2000)
Gerasimova, T. I., Byrd, K. & Corces, V. A chromatin insulator determines the nuclear localization of DNA. Mol. Cell 6, 1025–1035 (2000)
Cobb, B. S. et al. Targeting Ikaros to pericentromeric heterochromatin by direct DNA binding. Genes Dev. 14, 2146–2160 (2000)
Carmo-Fonseca, M., Platani, M. & Swedlow, J. R. Macromolecular mobility inside the cell nucleus. Trends Cell Biol. 12, 491–495 (2002)
Carmo-Fonseca, M. The contribution of nuclear compartmentalization to gene regulation. Cell 108, 513–521 (2002)
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001)
Mahy, N. L., Perry, P. E., Goilchrist, S., Baldock, R. A. & Bickmore, W. A. Spatial organization of active and inactive genes and noncoding DNA within chromosome territories. J. Cell Biol. 157, 579–589 (2004)
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002)
Tolhuis, B., Palstra, R.-J., Splinter, E., Grosveld, F. & de Laat, W. Looping and interaction between Hypersensitive sites in the active β-globin locus. Mol. Cell 10, 1453–1465 (2002)
Palstra, R.-J. et al. The β-globin nuclear compartment in development and erythroid differentiation. Nature Genet. 35, 190–194 (2003)
Spilianakis, C. & Flavell, R. A. Long-range intrachromosomal interactions in the T helper type 2 cytokine locus. Nature Immunol. 5, 1017–1027 (2004)
Takemoto, N. et al. TH2-specific DNase I-hypersensitive sites in the murine IL-13 and IL-4 intergenic region. Int. Immunol. 10, 1981–1985 (1998)
Agarwal, S. & Rao, A. Modulation of chromatin structure regulates cytokine gene expression during T cell differentiation. Immunity 9, 765–775 (1998)
Loots, G. G. et al. Identification of a coordinate regulator of interleukins-4, 13 and 5 by cross-species sequence comparisons. Science 288, 136–140 (2000)
Agarwal, S., Avni, O. & Rao, A. Cell-type-restricted binding of the transcription factor NFAT to a distal IL-4 enhancer in vivo . Immunity 12, 643–652 (2000)
Lee, G. R., Fields, P. E., Griffin, T. J. IV & Flavell, R. A. Regulation of the Th2 cytokine locus by a Locus Control Region. Immunity 19, 145–153 (2003)
Fields, P. E., Lee, G. R., Kim, S. T., Bartsevich, V. & Flavell, R. A. Th2-specific chromatin remodeling and enhancer activity in the Th2 cytokine locus control region. Immunity 21, 865–876 (2004)
Lee, D. U., Avni, O., Chen, L. & Rao, A. A distal enhancer in the IFN-γ locus revealed by genome sequence comparison. J. Biol. Chem. 279, 4802–4810 (2004)
Shnyreva, M. et al. Evolutionarily conserved sequence elements that positively regulate IFN-γ expression in T cells. Proc. Natl Acad. Sci. USA 101, 12622–12627 (2004)
Lee, G. R., Spilianakis, C. & Flavell, R. A. Hypersensitive site 7 of the TH2 locus control region is essential for expressing TH2 cytokine genes and for long-range intrachromosomal interactions. Nature Immunol. 6, 42–48 (2005)
Eivazova, E. R. & Aune, T. M. Dynamic alterations in the conformation of the Ifng gene region during T helper cell differentiation. Proc. Natl Acad. Sci. USA 101, 251–256 (2004)
Taddei, A., Maison, C., Roche, D. & Almouzni, G. Reversible disruption of pericentric heterochromatin and centromere function by inhibiting deacetylases. Nature Cell Biol. 3, 114–120 (2001)
Schmiedeberg, L., Weisshart, K., Diekmann, S., Meyer zu Hoerste, G. & Hemmerich, P. High- and low-mobility populations of HP1 in heterochromatin of mammalian cells. Mol. Biol. Cell 15, 2819–2833 (2004)
Eggenschwiler, J. et al. Mouse mutant embryos overexpressing IGF-II exhibit phenotypic features of the Beckwith–Wiedemann and Simpson–Golabi–Behmel syndromes. Genes Dev. 11, 3128–3142 (1997)
Dong, C. & Flavell, R. A. Cell fate decision: T-helper 1 and 2 subsets in immune responses. Arthritis Res. 2, 179–188 (2000)
Guo, L. et al. In TH2 cells the Il4 gene has a series of accessibility states associated with distinctive probabilities of IL-4 production. Proc. Natl Acad. Sci. USA 99, 10623–10628 (2002)
Kosak, S. T. & Groudine, M. Form follows function: the genomic organization of cellular differentiation. Genes Dev. 18, 1371–1384 (2004)
Guo, L. et al. In TH2 cells the IL-4 gene has a series of accessibility states associated with distinctive probabilities of IL-4 production. Proc. Natl Acad. Sci. USA 99, 10623–10628 (2002)
Parada, L. A., Sotiriou, S. & Misteli, T. Spatial genome organization. Exp. Cell Res. 296, 64–70 (2004)
Roix, J. J., McQueen, P. G., Munson, P. J., Parada, L. A. & Misteli, T. Spatial proximity of translocation-prone gene loci in human lymphomas. Nature Genet. 34, 287–291 (2003)
Avni, O. et al. TH cell differentiation is accompanied by dynamic changes in histone acetylation of cytokine genes. Nature Immunol. 3, 643–651 (2002)
Acknowledgements
We thank Wyeth Laboratories for donation of IL-12. We also thank F. Manzo for assistance with manuscript preparation, T. Gerasimova for suggestions with the FISH protocols, and D. Sakkas for use of his fluorescence microscope. We are also grateful to F. G. Grosveld and W. de Laat for originally providing us with detailed protocols and help with establishing the 3C technique. We would like to thank C. Szekely for assistance with graphs. C.S. is supported by a Cancer Research Institute fellowship; M.D.L is partly supported by a Human Frontiers Science Program long-term fellowship; T.T. is supported by a Ruth L. Kirchstein NIH/NRSA/NIA post-doctoral fellowship; R.A.F. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure S1
The IFN-γ and TH2 loci are colocalized in the nucleus as detected by two dimensional-fluorescence in situ hybridization (PPT 69 kb)
Supplementary Figure S2
Immuno-DNA FISH experiments reveal that the colocalized signals reside in euchromatin (PPT 902 kb)
Supplementary Figure S3 xm-repla ce_text {title}
Digestion of chromatinized genomic DNA within the nuclei of different cells types. (PPT 79 kb)
Supplementary Figure S4
IFNγ and TH2 loci colocalize in a more loose manner in the T cell nucleus. (PPT 45 kb)
Supplementary Figure Legends
Legends to accompany the above Supplementary Information (DOC 26 kb)
Supplementary Methods
Additional descripton of the methods used in this study, including Chromosome conformation capture analysis. (DOC 36 kb)
Rights and permissions
About this article
Cite this article
Spilianakis, C., Lalioti, M., Town, T. et al. Interchromosomal associations between alternatively expressed loci. Nature 435, 637–645 (2005). https://doi.org/10.1038/nature03574
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature03574
This article is cited by
-
Chromosome territory reorganization through artificial chromosome fusion is compatible with cell fate determination and mouse development
Cell Discovery (2023)
-
rDNA Transcription in Developmental Diseases and Stem Cells
Stem Cell Reviews and Reports (2023)
-
Association between fetal sex and maternal plasma microRNA responses to prenatal alcohol exposure: evidence from a birth outcome-stratified cohort
Biology of Sex Differences (2020)
-
Functions and properties of nuclear lncRNAs—from systematically mapping the interactomes of lncRNAs
Journal of Biomedical Science (2020)
-
Methods for mapping 3D chromosome architecture
Nature Reviews Genetics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.