Abstract
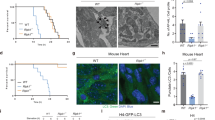
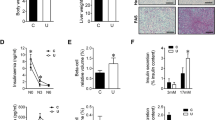
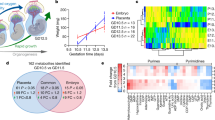
At birth the trans-placental nutrient supply is suddenly interrupted, and neonates face severe starvation until supply can be restored through milk nutrients1. Here, we show that neonates adapt to this adverse circumstance by inducing autophagy. Autophagy is the primary means for the degradation of cytoplasmic constituents within lysosomes2,3,4. The level of autophagy in mice remains low during embryogenesis; however, autophagy is immediately upregulated in various tissues after birth and is maintained at high levels for 3–12 h before returning to basal levels within 1–2 days. Mice deficient for Atg5, which is essential for autophagosome formation, appear almost normal at birth but die within 1 day of delivery. The survival time of starved Atg5-deficient neonates (∼ 12 h) is much shorter than that of wild-type mice (∼ 21 h) but can be prolonged by forced milk feeding. Atg5-deficient neonates exhibit reduced amino acid concentrations in plasma and tissues, and display signs of energy depletion. These results suggest that the production of amino acids by autophagic degradation of ‘self’ proteins, which allows for the maintenance of energy homeostasis, is important for survival during neonatal starvation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Medina, J. M., Vicario, C., Juanes, M. & Fernandez, E. in Perinatal Biochemistry (eds Herrera, E. & Knopp, R.) 233–258 (CRC Press, Boca Raton, 1992)
Cuervo, A. M. Autophagy: in sickness and in health. Trends Cell Biol. 14, 70–77 (2004)
Levine, B. & Klionsky, D. J. Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev. Cell 6, 463–477 (2004)
Mizushima, N., Ohsumi, Y. & Yoshimori, T. Autophagosome formation in mammalian cells. Cell Struct. Funct. 27, 421–429 (2002)
Klionsky, D. J. et al. A unified nomenclature for yeast autophagy-related genes. Dev. Cell 5, 539–545 (2003)
Tsukada, M. & Ohsumi, Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 333, 169–174 (1993)
Otto, G. P., Wu, M. Y., Kazgan, N., Anderson, O. R. & Kessin, R. H. Macroautophagy is required for multicellular development of the social amoeba Dictyostelium discoideum. J. Biol. Chem. 278, 17636–17645 (2003)
Juhasz, G., Csikos, G., Sinka, R., Erdelyi, M. & Sass, M. The Drosophila homolog of Aut1 is essential for autophagy and development. FEBS Lett. 543, 154–158 (2003)
Scott, R. C., Schuldiner, O. & Neufeld, T. P. Role and regulation of starvation-induced autophagy in the Drosophila fat body. Dev. Cell 7, 167–178 (2004)
Melendez, A. et al. Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 301, 1387–1391 (2003)
Doelling, J. H., Walker, J. M., Friedman, E. M., Thompson, A. R. & Veirstra, R. D. The APG8/12-activating enzyme APG7 is required for proper nutrient recycling and senescence in Arabidopsis thaliana. J. Biol. Chem. 277, 33105–33114 (2002)
Hanaoka, H. et al. Leaf senescence and starvation-induced chlorosis are accelerated by the disruption of an Arabidopsis autophagy gene. Plant Physiol. 129, 1181–1193 (2002)
Yue, Z., Jin, S., Yang, C., Levine, A. J. & Heintz, N. Beclin1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc. Natl Acad. Sci. USA 100, 15077–15082 (2003)
Qu, X. et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J. Clin. Invest. 112, 1809–1820 (2003)
Mizushima, N., Yamamoto, A., Matsui, M., Yoshimori, T. & Ohsumi, Y. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol. Biol. Cell 15, 1101–1111 (2004)
Mizushima, N. Methods for monitoring autophagy. Int. J. Biochem. Cell Biol. 36, 2491–2502 (2004)
Kabeya, Y. et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 19, 5720–5728 (2000)
Ichimura, Y. et al. A ubiquitin-like system mediates protein lipidation. Nature 408, 488–492 (2000)
Kabeya, Y. et al. LC3, GABARAP and GATE16 localize to autophagosomal membrane depending on form-II formation. J. Cell Sci. 117, 2805–2812 (2004)
Mizushima, N. et al. A protein conjugation system essential for autophagy. Nature 395, 395–398 (1998)
Mizushima, N., Sugita, H., Yoshimori, T. & Ohsumi, Y. A new protein conjugation system in human. The counterpart of the yeast Apg12p conjugation system essential for autophagy. J. Biol. Chem. 273, 33889–33892 (1998)
Mizushima, N. et al. Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells. J. Cell Biol. 152, 657–667 (2001)
Brun, S. et al. Activators of peroxisome proliferator-activated receptor-alpha induce the expression of the uncoupling protein-3 gene in skeletal muscle: a potential mechanism for the lipid intake-dependent activation of uncoupling protein-3 gene expression at birth. Diabetes 48, 1217–1222 (1999)
Hardie, D. G. Minireview: the AMP-activated protein kinase cascade: the key sensor of cellular energy status. Endocrinology 144, 5179–5183 (2003)
Carling, D. The AMP-activated protein kinase cascade—a unifying system for energy control. Trends Biochem. Sci. 29, 18–24 (2004)
Yamamoto, A. et al. Stacks of flattened smooth endoplasmic reticulum highly enriched in inositol 1,4,5-trisphosphate (InsP3) receptor in mouse cerebellar Purkinje cells. Cell Struct. Funct. 16, 419–432 (1991)
Wood, S. A., Allen, N. D., Rossant, J., Auerbach, A. & Nagy, A. Non-injection methods for the production of embryonic stem cell-embryo chimaeras. Nature 365, 87–89 (1993)
Acknowledgements
We thank M. Miwa and H. Satake for technical assistance. We also thank S. Sugano for donation of the pEF321-T plasmid; K. Ono and K. Tanaka for histological examination of the brain; M. Tamagawa for instruction in electrocardiogram recording; and S. Nishio, N. Tsunekawa and M. Terai for discussions. Amino acid measurements were carried out with the aid of the Center for Analytical Instruments at the National Institute for Basic Biology. This work was supported in part by Grants-in-aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure S1
Representative histological sections of haematoxylin and eosin-stained brain from wild type and Atg5-/- newborns. (JPG 64 kb)
Supplementary Figure S2
The restriction map of the wild-type Atg5 allele, the targeting construct, and the mutated allele. (JPG 23 kb)
Supplementary Table
Plasma and tissue amino acid concentrations in newborn mice under fasting conditions at 0 h and at 10 h after the caesarean delivery under fasting condition. (DOC 30 kb)
Rights and permissions
About this article
Cite this article
Kuma, A., Hatano, M., Matsui, M. et al. The role of autophagy during the early neonatal starvation period. Nature 432, 1032–1036 (2004). https://doi.org/10.1038/nature03029
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature03029
This article is cited by
-
Autophagy-modulating biomaterials: multifunctional weapons to promote tissue regeneration
Cell Communication and Signaling (2024)
-
In vitro modulation of mTOR and mGlur5 influence α-synuclein accumulation
Molecular Brain (2024)
-
Reduction of DHHC5-mediated beclin 1 S-palmitoylation underlies autophagy decline in aging
Nature Structural & Molecular Biology (2024)
-
Dissecting the multifaceted roles of autophagy in cancer initiation, growth, and metastasis: from molecular mechanisms to therapeutic applications
Medical Oncology (2024)
-
The impact of VPS35 D620N mutation on alternative autophagy and its reversal by estrogen in Parkinson's disease
Cellular and Molecular Life Sciences (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.