Abstract
Tetraodon nigroviridis is a freshwater puffer fish with the smallest known vertebrate genome. Here, we report a draft genome sequence with long-range linkage and substantial anchoring to the 21 Tetraodon chromosomes. Genome analysis provides a greatly improved fish gene catalogue, including identifying key genes previously thought to be absent in fish. Comparison with other vertebrates and a urochordate indicates that fish proteins have diverged markedly faster than their mammalian homologues. Comparison with the human genome suggests ∼900 previously unannotated human genes. Analysis of the Tetraodon and human genomes shows that whole-genome duplication occurred in the teleost fish lineage, subsequent to its divergence from mammals. The analysis also makes it possible to infer the basic structure of the ancestral bony vertebrate genome, which was composed of 12 chromosomes, and to reconstruct much of the evolutionary history of ancient and recent chromosome rearrangements leading to the modern human karyotype.
Similar content being viewed by others
Main
Access to entire genome sequences is revolutionizing our understanding of how genetic information is stored and organized in DNA, and how it has evolved over time. The sequence of a genome provides exquisite detail of the gene catalogue within a species, and the recent analysis of near-complete genome sequences of three mammals (human1, mouse2 and rat3) shows the acceleration in the search for causal links between genotype and phenotype, which can then be related to physiological, ecological and evolutionary observations. The partial sequence of the compact puffer fish Takifugu rubripes genome was obtained recently and this survey provided a preliminary catalogue of fish genes4. However, the Takifugu assembly is highly fragmented and as a result important questions could not be addressed.
Here, we describe and analyse the genome sequence of the freshwater puffer fish Tetraodon nigroviridis with long-range linkage and extensive anchoring to chromosomes. Tetraodon resembles Takifugu in that it possesses one of the smallest known vertebrate genomes, but as a popular aquarium fish it is readily available and is easily maintained in tap water (see Supplementary Notes for naming conventions, natural habitat and phylogeny). The two puffer fish diverged from a common ancestor between 18–30 million years (Myr) ago and from the common ancestor with mammals about 450 Myr ago5. This long evolutionary distance provides a good contrast to distinguish conserved features from neutrally evolving DNA by sequence comparison. Tetraodon sequences in fact had an important role in providing a reliable estimate of the number of genes in the human genome6.
There has been a vigorous and unresolved debate as to whether a whole-genome duplication (WGD) occurred in the ray-finned fish (actinopterygians) lineage after its separation from tetrapods7,8,9. By exploiting the extensive anchoring of the Tetraodon sequence to chromosomes, we provide a definitive answer to this question. The distribution of duplicated genes in the genome reveals a striking pattern of chromosome pairing, and the correspondence of orthologues with the human genome show precisely the signatures expected from an ancient WGD followed by a massive loss of duplicated genes.
Moreover, we find that relatively few interchromosomal rearrangements occurred in the Tetraodon lineage over several hundred million years after the WGD. This allows us to propose a karyotype of the ancestral bony vertebrate (Osteichthyes) composed of 12 chromosomes, and to uncover many unknown evolutionary breakpoints that occurred in the human genome in the past 450 Myr.
The Tetraodon genome sequence
Sequencing and assembly
The Tetraodon genome was sequenced using the whole-genome shotgun (WGS) approach. Random paired-end sequences providing 8.3-fold redundant coverage were produced at Genoscope (GSC) and the Broad Institute of MIT and Harvard (see Supplementary Table SI1). From this, the assembly program Arachne10,11 constructed 49,609 contigs for a total of 312 megabases (Mb; Table 1), which it then connected into 25,773 scaffolds (or supercontigs) covering 342 Mb (including gaps; see Supplementary Information). Half of the assembly is in 102 scaffolds larger than 731 kilobases (kb; the N50 length) and the largest scaffold measures 7.6 Mb, the typical length of a Tetraodon chromosome arm.
We produced additional data to physically link scaffolds and anchor them to chromosomes. These data include probe hybridizations to arrayed bacterial artificial chromosome (BAC) libraries, restriction digest fingerprints of BAC clones, additional linking clone sequence, alignment to available Takifugu sequence and two-colour fluorescence in situ hybridization (FISH) (see Supplementary Information). The impact of these additional mapping data was twofold: first, we could join 2,563 scaffolds in 128 ‘ultracontigs’ that cover 81.3% of the assembly, and second, we were able to anchor the 39 ultracontigs among the largest (covering 64.6% of the assembly, with an N50 size of 8.7 Mb) to Tetraodon chromosomes (Fig. 1; see also Supplementary Table SI2 and Supplementary Notes).
Red areas indicate the location of 5S and 28S ribosomal RNA gene arrays on chromosome 10 and chromosome 15, respectively. Many chromosomes are subtelocentric; that is, they only possess a very short heterochromatic arm. The extent of 39 sequence-based ultracontigs that cover about 64% of their length is shown in blue. In addition, approximately 16% of the genome is contained in another 89 ultracontigs that are not yet anchored on chromosomes, and the remaining 20% of the genome is in 23,210 smaller scaffolds.
The accuracy of the assembly was experimentally tested and the inter-contig links found to be correct in >99% of cases. On the basis of a re-sequencing experiment, we estimate that the assembly covers >90% of the euchromatin of the Tetraodon genome (Supplementary Information). Finally, the overall genome size was directly measured by flow cytometry experiments on several fish; an average value of 340 Mb was obtained, consistent with the sequence assembly and smaller than the previously reported estimate of 350–400 Mb.
The Tetraodon draft sequence has roughly 60-fold greater continuity at the level of N50 ultracontig size than the Takifugu draft sequence (7.62 Mb versus 125 kb). Critically, the anchoring of the assembly provides a comprehensive view of a fish genome sequence organized in individual chromosomes.
Genome landscape
A consequence of the remarkably compact nature of the Tetraodon genome is that its G + C content is much higher than in the larger genomes of mammals. Although the G + C content is shifted markedly, it still shows the same asymmetric bell-shaped distribution with an excess of higher values as seen in human and mouse (Fig. 2a). (G + C)-rich regions tend to be gene-rich in mammals, and analysis of our data shows that this is also true for Tetraodon (Fig. 2b, c). The Tetraodon genome thus cannot be considered as a single homogeneous component but, as in mammals, it is a mosaic of relatively gene-rich and gene-poor regions.
a, Distribution in 5-kb non-overlapping windows across Tetraodon (red squares) and Takifugu (blue circles) scaffolds, and in 50-kb windows in human (black triangles) and mouse (green inverted triangles) chromosomes. Windows containing more than 25% ambiguous or unknown nucleotides (gaps) were excluded from the analysis. b, Cumulative sum of annotated coding bases in Tetraodon and Takifugu (5-kb non-overlapping windows) and human and mouse (50-kb windows) as a function of G + C content. c, In sharp contrast to Takifugu4 the density of genes increases with the G + C content (%) in Tetraodon (red circles) much more than in human (black triangles). d, The three major families of repeats in Tetraodon are not distributed uniformly in the genome: long terminal repeat (LTR) and LINE elements (red diamonds and green squares, respectively) concentrate in (G + C)-rich regions and SINE elements (blue circles) concentrate in (A + T)-rich regions. In contrast, the distribution of these elements is much more uniform in Takifugu (Supplementary Fig. S4).
Transposable elements are very rare in the Tetraodon genome12,13: we estimate here that they do not exceed 4,000 copies; however, with 73 different types, they are richly represented (Supplementary Notes and Supplementary Table SI3). In sharp contrast, the human and mouse genomes contain only ∼20 different types but are riddled with millions of transposable element copies. One of the intriguing features of the human genome is that the distribution of short interspersed nucleotide elements (SINEs) is biased towards (G + C)-rich regions, whereas long interspersed nucleotide elements (LINEs) favour (A + T)-rich regions. In Tetraodon, these preferences are precisely reverse: LINEs occur preferentially in (G + C)-rich regions and SINEs in (A + T)-rich regions (Fig. 2d). The reason for these differences is not clear.
The Tetraodon genome shows certain striking differences from the previously reported Takifugu genome sequence. Takifugu contains eightfold more copies of transposable elements4 than Tetraodon, which may contribute to its slightly larger genome size (approximately 370 Mb; see Supplementary Information). More surprisingly, the G + C content of Takifugu does not show the characteristic asymmetry seen in mammals and in Tetraodon (Fig. 2a) nor the biases in SINE and LINE distribution (Supplementary Fig. S4). Why would the (G + C)-rich component be lacking in the Takifugu sequence, when this fraction is gene dense in mammals and in Tetraodon? This cannot be ascribed to transposable elements, which represent less than 5% of the assembly in both of these puffer fish species. One possible explanation is that the (G + C)-rich fraction exists in Takifugu, but was markedly under-represented as a result of aspects of the cloning, sequencing or assembly process. The fact that Tetraodon (G + C)-rich regions contain an excess of genes with no apparent orthologues in the Takifugu genome supports this hypothesis. Indeed, the Tetraodon genome appears to contain ∼16.5% more coding exons than Takifugu (see below).
Tetraodon genes
Gene catalogue
The most prevalent features of the Tetraodon genome are protein-coding genes, which span 40% of the assembly. We constructed a catalogue of genes by adapting the GAZE14 computational framework (Supplementary Fig. S5) in order to combine three types of data: Tetraodon complementary DNA mapping, similarities to human, mouse and Takifugu proteins and genomes, and ab initio gene models (Supplementary Notes and Supplementary Tables SI4 and SI5).
The current Tetraodon catalogue is composed of 27,918 gene models, with 6.9 coding exons per gene on average (7.3 including untranslated regions (UTRs); Table 2). Assuming that fish and mammal genes possess similar gene structures, this suggests that some Tetraodon annotated genes are partial or fragmented because human and mouse genes respectively show 8.7 and 8.4 coding exons per gene2. Adjusting the gene count for such fragmentation (by multiplying by 6.9/8.6) would yield an estimated gene count of 22,400 genes, whereas accounting for unsequenced regions of the genome might increase the estimate slightly further. Although such estimates are somewhat imprecise, it seems likely that Tetraodon has between 20,000–25,000 protein coding genes.
The Tetraodon gene catalogue appears to be the most complete so far for a fish, with coding exons and UTRs totalling ∼36 Mb (∼ 11% of the genome; Table 2). The Takifugu paper4 reported an estimate of 35,180 genes, but it did not account for a high degree of fragmentation (∼ 4.3 exons per gene model). More recent, unpublished analyses have revised this number sharply downward (Table 2). The human and Tetraodon genomes have a similar distribution of exon sizes but markedly different distributions of intron size (Supplementary Fig. S6a). Although neither genome seems to tolerate introns below approximately 50–60 base pairs, Tetraodon has accumulated a much higher frequency of introns at this lower limit. Interestingly, this phenomenon is not uniform across the genome: there is an excess of genes with many small introns (Supplementary Fig. S6b), suggesting that intron sizes fluctuate in a regional fashion.
Proteome comparison between vertebrates
We examined in detail two gene families with unusual properties that represent challenges for automatic annotation procedures and have particular biological interest. The first is the family of selenoproteins, where the UGA codon encodes a rare cysteine analogue named selenocysteine (Sec) instead of signalling the end of translation as in all other genes15. We annotated 18 distinct families in Tetraodon based on similarities with the 19 protein families known in eukaryotes, and discovered a new selenoprotein that seems to be restricted to the actinopterygians among vertebrates and does not have a Cys counterpart in mammals. We also catalogued type I helical cytokines and their receptors (HCRI), a group of genes that were not found in the Takifugu genome4 because of their poor sequence conservation, leading to the hypothesis that fish may not possess this large family that includes hormones and interleukins. Tetraodon, in fact, contains 30 genes encoding HCRIs with a typical D200 domain (Supplementary Fig. S7) and represents all families previously described in mammals16.
InterPro17 domains were annotated in protein sequences predicted in the Tetraodon, Takifugu, human, mouse and the urochordate Ciona intestinalis18 genome using InterProScan19. We did not identify major differences between fish and mammal InterPro families, except for a few striking cases (Table 3): (1) collagen molecules are much more diverse in fish than in mammals, with one Tetraodon gene containing 20 von Willebrand type A domains, the largest number found so far in a single protein. (2) Some domains associated with sodium transport are noticeably enriched in fishes and Ciona, perhaps a reflection of their adaptation to saline aquatic environments that was lost in land vertebrates. (3) Purine nucleosidases usually involved in the recovery of purine nucleosides are more abundant in fish, including an allantoin pathway for purine degradation that is present in Tetraodon and absent in human. (4) Several hundred KRAB box transcriptional repressors involved in chromatin-mediated gene regulation exist in mammals and are totally absent in fish. (5) Proteins involved in general gene regulation are more abundant in vertebrates than in Ciona.
Protein annotation with gene ontology (GO) classifications20 shows only subtle differences between fish and mammals, as was already observed between human and mouse2. The largest differences between species are seen with the GO classification in molecular functions (Supplementary Fig. S9). Interestingly, the two puffer fish and Ciona often vary together, showing for instance a higher frequency of enzymatic and transporter functions, and a lower frequency of signal transducer and structural molecules than both mammals (human and mouse). These global observations are difficult to relate to evolutionary or physiological mechanisms but provide a framework to understand the emergence or decline of molecular functions in vertebrates.
Number of genes in mammals and teleosts
The total amount of coding sequence conserved between the two fish and the two mammalian genomes provides a measure of their respective coding capacity. The Exofish method6 is well suited to measure this, because it translates entire genomes in all six frames and identifies conserved coding regions (ecores) with a high specificity and independently of prior genome annotation (Table 4; see also Supplementary Information). The four vertebrate genomes contain remarkably similar numbers of ecores, apart from minor differences attributable to varying degrees of sequence completion. This suggests that they possess fairly similar numbers of genes. In fact, the gene count may be slightly less in mammals than in fish because the proportion of ecores corresponding to pseudogenes is higher in mammals21.
The human ecores can be used to search for previously unrecognized human genes. The discovery of new human genes is becoming an increasingly rare event, given the scale and intensity of international efforts to annotate the genome by systematic annotation pipelines and by human experts. Roughly 14,500 human ecores conserved with Tetraodon sequences do not overlap any ‘known’ features (genes or pseudogenes) in the human genome. Using these as anchors for local gene identification using the GAZE program, we identified 904 novel human gene predictions. Of these, 63% are also supported by expressed sequence tag (EST) data (from human or other species) and 50% contain predicted InterPro protein domains (Supplementary Table SI9). The most convincing evidence supporting these gene predictions is that they are strongly enriched on chromosomes that have not yet been annotated by human experts (Supplementary Table SI10). The novel gene predictions have relatively small size (average coding sequence (CDS) of 469 bp), which may have caused them to be eliminated by systematic annotation procedures. They provide a rich resource to help complete the human gene catalogue.
Tetraodon gene evolution
We measured rates of sequence divergence between fish and mammals to estimate the relative speed with which functional and non-functional sequences evolve in these lineages. We used fourfold degenerate (4D) site substitutions in orthologous proteins as a proxy for neutral nucleotide mutations, an approach that has been shown to be robust across entire genomes2. To optimize further the selection of sites used for comparison, we only considered the 5,802 proteins that are identified as orthologues in all pairwise comparisons between human, mouse, Tetraodon and Takifugu. The average neutral nucleotide substitution rate, inferred using the REV model22,23, shows that the divergence between Tetraodon and Takifugu is about twice as fast per year as between human and mouse (Table 5), or between mouse and rat3.
We were interested to see whether this higher mutation rate is also seen in protein sequences. Pairwise comparison of all possible combinations of the 5,802 four-way orthologous proteins clearly indicates that proteins between the two puffer fish are more divergent than between the two mammals, despite the shorter evolutionary time that has elapsed (Fig. 3). This is confirmed by the fact that the average frequency of non-synonymous mutations (leading to an amino acid change, Ka) between C. intestinalis and human proteins is lower than between Ciona and Tetraodon (see Methods).
Comparisons were performed with 2,289 proteins that are orthologous between the chordate C. intestinalis and all four vertebrates—Tetraodon, Takifugu, human and mouse (asterisks)—and with 5,802 proteins orthologous between all four vertebrates only, between fish and mammals (triangles) or between the two fish (circles), and between the two mammals (squares). As expected, all vertebrates show the same distribution profile compared to Ciona and both fish show the same distribution profile compared to mammals. Surprisingly, the distribution profile of the comparison between the two fish and between the two mammals is also very similar, despite the much shorter evolutionary time since the tetraodontiform radiation.
Independent of the overall rate of change, the ratio of non-synonymous to synonymous changes (Ka/Ks ratio) is much higher between the two puffer fish than between human and mouse (Supplementary Table SI11 and Supplementary Information), suggesting that protein evolution is proceeding more rapidly along the puffer fish lineage. The reasons for this faster tempo of protein change are unknown, although it is likely to be positively correlated with the higher rate of neutral mutation.
Genome evolution
Genome-wide sequence provides a rare opportunity to address key evolutionary questions in a global fashion, circumventing biases due to small sequence and gene samples. In this respect, the combination of long-range linkage in the Tetraodon sequence and its evolutionary divergence from the mammalian lineage at 450 Myr ago makes it possible to explore overall genome evolution in the vertebrate clade.
Evidence for whole-genome duplication
The occurrence of WGD in the ray-finned fish lineage is a hotly debated question due both to the cataclysmic nature of such an event and to the difficulty in establishing that it actually occurred24,25,26. Definitive proof of WGD requires identifying certain distinctive signatures in long-range genome organization, which has previously been impossible to address with the data available.
It is expected that after WGD the resulting polyploid genome gradually returns to a diploid state through extensive gene deletion, with only a small proportion of duplicated copies ultimately retained as sources of functional innovation26. Paralogous chromosomes will thus each retain only a small subset of their initially common gene complement and then will be broken into smaller segments by genomic rearrangements. WGD will thus leave two distinctive signs for considerable periods before eventually fading.
The first distinctive sign is duplicated genes on paralogous chromosomes. In the absence of chromosomal rearrangement it would be simple to recognize two paralogous chromosomes arising from a WGD from the genome-wide distribution of duplicate genes: the chromosomes would each contain one member from many duplicated gene pairs occurring in the same order along their length. The difficulty is that this neat picture will eventually be blurred by interchromosomal rearrangement, which will disrupt the 1:1 correspondence between chromosomes, and intrachromosomal rearrangement, which will disrupt gene ordering along chromosomes.
We analysed the genome-wide distribution of duplicated gene pairs to see whether a strong correspondence between chromosomes could be detected. We identified 1,078 and 995 pairs of duplicated genes in the Tetraodon and Takifugu genomes, respectively, using conservative criteria (see Supplementary Information). On the basis of the frequencies of silent mutations (Ks) between copies, ∼75% are ‘ancient’ duplications that arose before the Tetraodon–Takifugu speciation (Fig. 4a).
a, Distribution of Ks values of duplicated genes in Tetraodon (left) and Takifugu (right) genomes. Duplicated genes broadly belong to two categories, depending on their Ks value being below or higher than 0.35 substitutions per site since the divergence between the two puffer fish (arrows). b, Global distribution of ancient duplicated genes (Ks > 0.35) in the Tetraodon genome. The 21 Tetraodon chromosomes are represented in a circle in numerical order and each line joins duplicated genes at their respective position on a given pair of chromosomes.
The chromosomal distribution of these ancient duplicates follows a striking pattern characteristic of a WGD. Genes on one chromosome segment have a strong tendency to possess duplicate copies on a single other chromosome (Fig. 4b). The correspondence is not a perfect 1:1 match owing to interchromosomal exchange, but it is vastly stronger than expected by chance (Supplementary Table SI12). As expected from a WGD, all chromosomes are involved. Remarkably, some duplicate chromosome pairs such as Tetraodon chromosome 9 (Tni9) and Tni11 have remained largely undisturbed by chromosome translocations since the duplication event. In other cases, one chromosome has links to two or three others, suggestive of either fusion or fragmentation (for example, Tni13 matches Tni5 and Tni19).
The second distinctive sign, which is an even more powerful signature of genome duplication, comes from comparison with a related species carrying a genome that did not undergo the WGD. Such a comparison was recently used to prove the existence of an ancient WGD in the yeast Saccharomyces cerevisiae based on comparison with a second yeast species Kluyveromyces waltii that diverged before the WGD27,28. Although two ancient paralogous regions typically retained only a few genes in common, they could be readily recognized because they showed a characteristic 2:1 mapping with interleaving; that is, they both showed conserved synteny and local order to the same region of the K. waltii genome with the S. cerevisiae genes interleaving in alternating stretches. Such regions were called blocks of DCS (doubly conserved synteny). Whereas the first distinctive sign of WGD depends only on a minority of duplicated genes, the DCS signature considers all genes for which orthologues can be found in the related species.
We used 6,684 Tetraodon genes localized on individual chromosomes that possess an orthologue in either human or mouse to create a high-resolution synteny map (Fig. 5 and Supplementary Fig. S11, respectively). The map contains 900 syntenic groups composed of at least two consecutive genes (average 6.1; maximum 55) having orthologues on the same human chromosome; the syntenic groups include 76% of Tetraodon–human orthologues. The synteny map with mouse contains 1,011 syntenic groups, probably reflecting the higher degree of chromosomal rearrangement in the rodent lineage2.
a, For each Tetraodon chromosome, coloured segments represent conserved synteny with a particular human chromosome. Synteny is defined as groups of two or more Tetraodon genes that possess an orthologue on the same human chromosome, irrespective of orientation or order. Tetraodon chromosomes are not in descending order by size because of unequal sequence coverage. The entire map includes 5,518 orthologues in 900 syntenic segments. b, On the human genome the map is composed of 905 syntenic segments. See Supplementary Information for the synteny map between Tetraodon and mouse (Supplementary Fig. S11).
The synteny map typically associates two regions in Tetraodon with one region in human. Using precise criteria (see Methods) we defined DCS blocks for Tetraodon relative to human; in contrast to the yeast study, strict conservation of gene order within DCSs was not required. Notably, most (79.6%) orthologous genes in syntenic groups can be assigned to 90 DCS blocks (Fig. 6). As in S. cerevisiae27, we see the distinctive interleaving pattern expected from WGD followed by massive gene loss. Analysis of the interleaving pattern shows that the gene loss occurred through many small deletions in a balanced fashion over the two Tetraodon sister chromosomes (average balance 42% and 58% of retention; Supplementary Information); this is consistent with the results in yeast.
Blocks of synteny along human chromosomes map to two (or three) Tetraodon chromosomes in an interleaving pattern. Small boxes represent groups of syntenic orthologous genes enclosed in larger boxes that define the boundaries of 110 DCS blocks. Black circles indicate human centromeres. A region of human chromosomes Xq and 16q are shown in detail with individual Tetraodon orthologous genes depicted on either side.
These two analyses provide definitive evidence that the Tetraodon genome underwent a WGD sometime after its divergence from the mammalian lineage. The first test used only the ∼3% of genes that represent duplicated gene pairs retained from the WGD. The second test used the pattern of 2:1 mapping with interleaving involving ∼80% of orthologues between Tetraodon and human.
The presence of supernumerary HOX clusters in zebrafish7, Tetraodon (see Supplementary Figure 8) and many other percomorphs29 but not in the bichir Polypterus senegalus30 indicates that the event has affected most teleosts but not all actinopterygians. This timing early in the teleost lineage is in agreement with recent evolutionary analyses in Takifugu that estimated the divergence time for most duplicated gene pairs at ∼320–350 Myr ago31,32.
The analyses above also shed light on the rate of intra- and interchromosomal exchange. The synteny analysis shows extensive syntenic segments in which gene content has been well preserved but gene order has been extensively scrambled (striking examples include conserved synteny of Tni20 with human chromosome 4q (Hsa4q) and Tni1 with HsaXq); this is consistent with observations in zebrafish33. The duplication analysis within Tetraodon also shows that the chromosomal correspondence of duplicated gene pairs has been extensively preserved, whereas local gene order has been largely scrambled. Both analyses thus indicate that a relatively high degree of intrachromosomal rearrangement and a relatively low degree of interchromosomal exchange have taken place in the Tetraodon lineage.
Ancestral genome of bony vertebrates
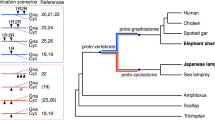
We then sought to use the correspondence between the Tetraodon and human genomes to attempt to reconstruct the karyotype of their osteichthyan (bony vertebrate) ancestor. The DCS blocks define Tetraodon regions that arose from duplication of a common ancestral region. Notably, the DCS blocks largely fall into 12 simple patterns: eight cases involving the interleaving of two current Tetraodon chromosomes and four cases involving three current Tetraodon chromosomes (Fig. 7 and Table 6). The first group represents cases in which the ancestral chromosomes have remained largely untouched by interchromosomal exchange; the second group represents cases in which one major translocation has occurred.
The 110 DCS blocks identified on the human genome are grouped according to their composition in terms of Tetraodon chromosomes, thus delineating 12 ancestral chromosomes containing 90 DCS blocks. The order of DCSs within an ancestral chromosome is arbitrary. The 20 blocks denoted by the letters U, V, W and Z (Supplementary Information) could not be assigned to an ancestral chromosome because each has a unique composition, probably due to rearrangements in the human or Tetraodon genome. Colour codes are as in Fig. 6.
The distribution of Tetraodon orthologues in the human genome (shown as an Oxford grid in Supplementary Fig. S12) provides a detailed record that can be used to partially reconstruct the history of rearrangements in both lineages. We considered the expected distribution resulting from various types of interchromosomal rearrangements, assuming a relatively high degree of intrachromosomal shuffling (Fig. 8; see also Supplementary Information). We found that only ten large-scale interchromosomal events suffice to largely explain the data, connecting an ancestral vertebrate karyotype of 12 chromosomes to the modern Tetraodon genome of 21 chromosomes (Fig. 9). Eleven of the Tetraodon chromosomes appear to have undergone no major interchromosomal rearrangement. For example, 13 DCS blocks in human are composed of interleaved syntenic groups mapping to Tni9 and Tni11, which are presumed to be derived from a common ancestral chromosome denoted chromosome K (AncK; Fig. 7). The orthologue distribution between the two chromosomes (Fig. 8) confirms that they derive by duplication from AncK (Fig. 9). In a more complex case, Tni13 is systematically interleaved with Tni5 (AncE) or Tni19 (AncF), but Tni5 and Tni19 are never interleaved together; the orthologue distribution among the three chromosomes (Fig. 8) implies that the duplication partners of Tni5 and Tni19 fused soon after the WGD to give rise to Tni13 (Fig. 9). The overall model is consistent with a complete WGD, in that it accounts for all Tetraodon chromosomes.
Model of chromosome duplication followed by the four simplest chromosome rearrangements: (1) no rearrangement; (2) two different duplicate copies fused recently; (3) two different duplicate copies fused early after the duplication; (4) a duplicate chromosome fragmented very recently. In each model, the distribution of human orthologues from a given chromosomal region on two or three duplicate Tetraodon chromosomal regions is expected to be different (each dot is an orthologue, positioned in the human genome on the vertical axis and in the Tetraodon genome on the horizontal axis). The distinction between early or late events follows the assumption that intrachromosomal shuffling progressively redistributes genes over a given chromosome. A recent fusion would thus bring together two sets of genes that appear compartmented on their respective segments, whereas an ancient fusion shows the same pattern except that genes have been redistributed over the length of the fused chromosome. It should be noted that a fifth case exists, consisting of a chromosome break early after duplication but it is not represented here. The lower panel shows excerpts of data illustrating the four types of event. The complete Oxford grid is shown in Supplementary Fig. SI12.
The ten major rearrangements (two ancient fusions, three recent fusions, one ancient and one recent fission, and three ancient translocations) are deduced by fitting the distribution of orthologues to the four simple theoretical models of chromosome evolution. The order between events is arbitrary although the approximate timeline differentiates between ancient and recent events respectively before and after the dashed line. Arrowheads point to the direction of three ancient translocations.
Several lines of evidence support the historical reconstitution presented here. First, the pairing of Tetraodon chromosomes agrees with the independently derived distribution of duplicated genes in the genome (Fig. 4b). Second, centric fusions of the three largest chromosomes are consistent with cytogenetic studies34, and the recent timing of the fusion leading to Tni1 is supported by cytogenetic studies showing its absence in Takifugu35. Third, the modal value for the haploid number of chromosomes in teleosts is 24 (refs 36–38), consistent with a WGD of an ancestral genome composed of 12 chromosomes.
The analysis also sheds light on genome evolution in the human lineage, with the interleaving patterns on human chromosomes delineating the mosaic of ancestral segments in the human genome (Figs 6 and 10). The results are consistent with and extend several known cases of rearrangements in the human lineage. The model correctly shows the recent fusion of two primate chromosomes leading to Hsa2 (ref. 39) occurring at the junction between two ancestral segments (D2 and D3; Fig. 6) in 2q13.2-2q14.1. It shows HsaXp and HsaXq to be of different origins (corresponding to AncD and AncH, respectively), consistent with the fact that HsaXp is known to be absent in non-placental mammals40. The map indicates that most of HsaXq and Hsa5q were once part of the same chromosome, but that the tip of HsaXq (Xq28) originates from a different ancestral segment and is thus a later addition. Some pairs of human chromosomes show similar or identical compositions, suggesting that they derived by fission from the same ancestral chromosome, with examples being Hsa13–Hsa21 and Hsa12–Hsa22; the latter case is consistent with cytogenetic studies showing that a fission occurred in the primate lineage41.
The composition of Tetraodon chromosomes is based on their duplication pattern (Fig. 9), whereas the composition of human chromosomes is based on the distribution of orthologues of Tetraodon genes (Fig. 6). A vertical line in Tetraodon chromosomes denotes regions where sequence has not yet been assigned. With 90 blocks in human compared with 44 in Tetraodon, the complexity of the mosaic of ancestral segments in human chromosomes underlines the higher frequency of rearrangements to which they were submitted during the same evolutionary period.
The results show a major difference in the evolutionary forces shaping the Tetraodon and the human genomes (Fig. 10). Whereas 11 Tetraodon chromosomes did not undergo interchromosomal exchange over 450 Myr, only one human chromosome (Hsa14) was similarly undisturbed. Hsa7 is an extreme case, with contributions from six ancestral chromosomes. A possible explanation for the difference may be the massive integration of transposable elements in the human genome. The presence of transposable elements may increase the overall frequency of chromosome breaks, as well as the likelihood that a chromosome break fails to disrupt a gene (by increasing the size of intergenic intervals). It will be interesting to see whether teleosts that carry many more transposable elements (such as zebrafish) show a higher frequency of interchromosomal exchanges.
Conclusion
The purpose of sequencing the Tetraodon genome was to use comparative analysis to illuminate the human genome in particular and vertebrate genomes in general. The Tetraodon sequence, which has been made freely available during the course of this project, has already had a major impact on human gene annotation. It has provided the first clear evidence of a sharply lower human gene count6 and has been used in the annotation of several human chromosomes42,43,44,45. Here, we show that it suggests an additional ∼900 predicted genes in the human genome. Given its compact size, the Tetraodon genome will probably also prove valuable in identifying key conserved regulatory features in intergenic and intronic regions.
In addition, the Tetraodon genome provides fundamental insight into genome evolution in the vertebrate lineage. First, the analysis here shows that Tetraodon is the descendant of an ancient WGD that most probably affected all teleosts. Together with the recent demonstration of an ancient WGD in the yeast lineage, this suggests that WGD followed by massive gene loss may be an extremely important mechanism for eukaryote genome evolution—perhaps because it allows for the neofunctionalization of entire pathways rather than simply individual genes. There remains a fierce debate about whether one or more earlier WGD events occurred in early vertebrate evolution25,46,47,48,49,50, with no direct and conclusive evidence found so far51,52. The examples of yeast and Tetraodon show that ultimate proof will probably best come from the sequence of a related non-duplicated species. An obvious candidate is amphioxus, as its non-duplicated status is supported by the presence of many single-copy genes (including one HOX cluster53) instead of two or more in vertebrates, and it is among our closest non-vertebrate relatives based on anatomical and evolutionary observations.
Second, the remarkable preservation of the Tetraodon genome after WGD makes it possible to infer the history of vertebrate chromosome evolution. The model suggests that the ancestral vertebrate genome was comprised of 12 chromosomes, was compact, and contained not significantly fewer genes than modern vertebrates (inasmuch as the WGD and subsequent massive gene loss resulted in only a tiny fraction of duplicate genes being retained). The explosion of transposable elements in the mammalian lineage, subsequent to divergence from the teleost lineage, may have provided the conditions for increased interchromosomal rearrangements in mammals; in contrast, the Tetraodon genome underwent much less interchromosomal rearrangement.
With the availability of additional vertebrate genomes (dog, marsupial, chicken, medaka, zebrafish and frog are underway), it will be possible to explore intermediate nodes such as the last common ancestor of amniotes, of sarcopterygians and of actinopterygians, and to gain an increasingly clearer picture of the early vertebrate ancestor. Because the early vertebrate genome is ‘closer’ to current invertebrates, this should in turn facilitate comparison between vertebrate and invertebrate evolution.
Methods
Sequencing, assembly and data access
Sequencing was performed as described previously for Genoscope54 and the Broad Institute1,2. Approximately 4.2 million plasmid reads were cloned and sequenced from DNA extracted from two wild Tetraodon fish and passed extensive checks for quality and source, representing approximately 8.3-fold sequence coverage of the Tetraodon genome. To alleviate problems due to polymorphism, the assembly proceeded in four stages: (1) reads from a single fish were assembled by Arachne as described previously10,11; (2) reads from the second individual were added to increase sequencing depth; (3) scaffolds were constructed using plasmid and BAC paired reads; and (4) contigs from a separate assembly combining both individuals were added if they did not overlap with the first assembly. The final assembly can be downloaded from the EMBL/GenBank/DDBJ databases under accession number CAAE01000000. Full-length Tetraodon cDNAs have been submitted under accession numbers CR631133–CR735083. Ultracontigs organized in chromosomes are available from http://www.genoscope.org/tetraodon. This site also contains an annotation browser and further information on the project.
Gene annotation
Protein-coding genes were predicted by combining three types of information: alignments with proteins and genomic DNA from other species, Tetraodon cDNAs, and ab initio models. All alignments with genomic DNA from human and mouse were performed with Exofish as described previously6, whereas a new Exofish method was developed to align Takifugu genomic DNA. Proteins predicted from human and mouse were also matched using Exofish and a selected subset was then aligned using Genewise. The integration of these data sources was performed with GAZE14. A specific GAZE automaton was designed, and parameters were adjusted on a training set of 184 manually annotated Tetraodon genes. See Supplementary Information for details.
Evolution of coding and non-coding DNA
To identify orthologous genes between human, mouse, Tetraodon, Takifugu and Ciona, their predicted proteomes were compared using the Smith–Waterman algorithm and reciprocal best matches were considered as orthologous genes between two species. However, only those genes that were reciprocal best matches between four or five species, and only sites that were aligned between the four or five genes, were further considered to compute the percentage identity, Ka, Ks and fourfold degenerate sites by the PBL method applying Kimura's two-parameter model55,56,57. See Supplementary Information for details.
Genome duplication
A core set of Tetraodon duplicated genes was identified by an all-against-all comparison of Tetraodon predicted protein using Exofish. Only proteins that matched a single other protein by reciprocal best match were considered further and realigned by the Smith–Waterman algorithm to compute Ka and Ks values. Duplicates with a Ks > 0.35 (the amount of neutral substitution since the Tetraodon–Takifugu divergence) were considered ‘ancient’ and used to calculate P-values for chromosome pairing (Supplementary Table SI12). Rules for classifying alternating patterns of syntenic groups along human chromosomes in DCS blocks included the following criteria: number of genes in syntenic groups, number of syntenic groups in the DCS region, number of Tetraodon chromosomes that alternate, and number of times the same combination of Tetraodon chromosomes occur in the human genome. See Supplementary Information for details.
Ancestral genome reconstruction
One category of DCS with the following definition encompassed most orthologues: “alternating series of i syntenic groups that belong to two (i > = 2) or three (i > = 3) Tetraodon chromosomes. The series may only be interrupted by groups from categories ‘unassigned singletons’ or ‘background singletons’. A given combination of two or three Tetraodon chromosomes must appear at least twice in the human genome”. These DCS blocks showed 12 recurring combinations of Tetraodon chromosomes, and were thus further classified in 12 groups labelled A to L. Each of the 12 groups, consisting of at least two DCS blocks with the same combination of alternating Tetraodon chromosomes, represents a proto-chromosome from the ancestral bony vertebrate (Osteichthyes). A model was then designed to account for the possible fates of chromosomes after duplication of the ancestral genome in the teleost lineage (Fig. 8). The model only deals with orthologous gene distribution between two genomes. It is simply based on the postulate that interchromosomal shuffling of genes within a genome increases with time, which is a measure to distinguish between ancient and recent events (for example, chromosome fusions or fissions). The two-dimensional distribution of 7,903 Tetraodon–human orthologues (Oxford Grid, Supplementary Fig. S12) was then confronted to the model and all 21 Tetraodon chromosomes could be grouped in pairs or triplets and assigned to a given type of event. See Supplementary Information for details.
References
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002)
Rat Genome Sequencing Project Consortium. Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428, 493–521 (2004)
Aparicio, S. et al. Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 297, 1301–1310 (2002)
Hedges, S. B. The origin and evolution of model organisms. Nature Rev. Genet. 3, 838–849 (2002)
Roest Crollius, H. et al. Estimate of human gene number provided by genome-wide analysis using Tetraodon nigroviridis DNA sequence. Nature Genet. 25, 235–238 (2000)
Amores, A. et al. Zebrafish hox clusters and vertebrate genome evolution. Science 282, 1711–1714 (1998)
Robinson-Rechavi, M., Marchand, O., Escriva, H. & Laudet, V. An ancestral whole-genome duplication may not have been responsible for the abundance of duplicated fish genes. Curr. Biol. 11, R458–R459 (2001)
Taylor, J. S., Braasch, I., Frickey, T., Meyer, A. & Van de Peer, Y. Genome duplication, a trait shared by 22000 species of ray-finned fish. Genome Res. 13, 382–390 (2003)
Batzoglou, S. et al. ARACHNE: a whole-genome shotgun assembler. Genome Res. 12, 177–189 (2002)
Jaffe, D. B. et al. Whole-genome sequence assembly for mammalian genomes: Arachne 2. Genome Res. 13, 91–96 (2003)
Roest Crollius, H. et al. Characterization and repeat analysis of the compact genome of the freshwater pufferfish Tetraodon nigroviridis. Genome Res. 10, 939–949 (2000)
Bouneau, L. et al. An active non-LTR retrotransposon with tandem structure in the compact genome of the pufferfish Tetraodon nigroviridis. Genome Res. 13, 1686–1695 (2003)
Howe, K. L., Chothia, T. & Durbin, R. GAZE: a generic framework for the integration of gene-prediction data by dynamic programming. Genome Res. 12, 1418–1427 (2002)
Hatfield, D. L. Selenium: Its Molecular Biology and Role in Human Health (Kluwer, Dordrecht, 2001)
Boulay, J. L., O'Shea, J. J. & Paul, W. E. Molecular phylogeny within type I cytokines and their cognate receptors. Immunity 19, 159–163 (2003)
Mulder, N. J. et al. InterPro: an integrated documentation resource for protein families, domains and functional sites. Brief. Bioinform. 3, 225–235 (2002)
Dehal, P. et al. The draft genome of Ciona intestinalis: insights into chordate and vertebrate origins. Science 298, 2157–2167 (2002)
Zdobnov, E. M. & Apweiler, R. InterProScan—an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17, 847–848 (2001)
Harris, M. A. et al. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res. 32(Database issue), D258–D261 (2004)
Torrents, D., Suyama, M., Zdobnov, E. & Bork, P. A genome-wide survey of human pseudogenes. Genome Res. 13, 2559–2567 (2003)
Tavaré, S. Some probabilistic and statistical problems in the analysis of DNA sequences. Lect. Math. Life Sci. 17, 57–86 (1986)
Gu, X. & Li, W. H. A general additive distance with time-reversibility and rate variation among nucleotide sites. Proc. Natl Acad. Sci. USA 93, 4671–4676 (1996)
Holland, P. W. H. Introduction: gene duplication in development and evolution. Semin. Cell Dev. Biol. 10, 515–516 (1999)
Martin, A. Is tetralogy true? Lack of support for the “one-to-four” rule. Mol. Biol. Evol. 18, 89–93 (2001)
Wolfe, K. H. Yesterday's polyploids and the mystery of diploidization. Nature Rev. Genet. 2, 333–341 (2001)
Kellis, M., Birren, B. W. & Lander, E. S. Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae. Nature 428, 617–624 (2004)
Dietrich, F. S. et al. The Ashbya gossypii genome as a tool for mapping the ancient Saccharomyces cerevisiae genome. Science 304, 304–307 (2004)
Prohaska, S. J. & Stadler, P. F. The duplication of the Hox gene clusters in teleost fishes. Theor. Biosci. 123, 89–110 (2004)
Chiu, C. H. et al. Bichir HoxA cluster sequence reveals surprising trends in ray-finned fish genomic evolution. Genome Res. 14, 11–17 (2004)
Vandepoele, K., De Vos, W., Taylor, J. S., Meyer, A. & Van de Peer, Y. Major events in the genome evolution of vertebrates: paranome age and size differ considerably between ray-finned fishes and land vertebrates. Proc. Natl Acad. Sci. USA 101, 1638–1643 (2004)
Christoffels, A. et al. Fugu genome analysis provides evidence for a whole-genome duplication early during the evolution of ray-finned fishes. Mol. Biol. Evol. 21, 1146–1151 (2004)
Woods, I. G. et al. A comparative map of the zebrafish genome. Genome Res. 10, 1903–1914 (2000)
Fischer, C. et al. Karyotype and chromosomal localization of characteristic tandem repeats in the pufferfish Tetraodon nigroviridis. Cytogenet. Cell Genet. 88, 50–55 (2000)
Grutzner, F. et al. Classical and molecular cytogenetics of the pufferfish Tetraodon nigroviridis. Chromosome Res. 7, 655–662 (1999)
Ohno, S., Wolf, U. & Atkin, N. B. Evolution from fish to mammals by gene duplication. Hereditas 59, 169–187 (1968)
Ojima, Y. in Chromosomes in Evolution of Eukaryotic Groups (eds Sharma, A. K. & Sharma, A.) 111–145 (CRC Press, Boca Raton, 1983)
Naruse, K. et al. A medaka gene map: the trace of ancestral vertebrate proto-chromosomes revealed by comparative gene mapping. Genome Res. 14, 820–828 (2004)
Yunis, J. J. & Prakash, O. The origin of man: a chromosomal pictorial legacy. Science 215, 1525–1530 (1982)
Graves, J. A., Gecz, J. & Hameister, H. Evolution of the human X—a smart and sexy chromosome that controls speciation and development. Cytogenet. Genome Res. 99, 141–145 (2002)
Richard, F., Lombard, M. & Dutrillaux, B. Reconstruction of the ancestral karyotype of eutherian mammals. Chromosome Res. 11, 605–618 (2003)
The chromosome 21 mapping and sequencing consortium, The DNA sequence of human chromosome 21. Nature 405, 311–319 (2000)
Deloukas, P. et al. The DNA sequence and comparative analysis of human chromosome 20. Nature 414, 865–871 (2001)
Collins, J. E. et al. Reevaluating human gene annotation: a second-generation analysis of chromosome 22. Genome Res. 13, 27–36 (2003)
Heilig, R. et al. The DNA sequence and analysis of human chromosome 14. Nature 421, 601–607 (2003)
Holland, P. W., Garcia-Fernandez, J., Williams, N. A. & Sidow, A. Gene duplications and the origins of vertebrate development. Development(suppl.), 125–133 (1994)
Spring, J. Vertebrate evolution by interspecific hybridisation–are we polyploid? FEBS Lett. 400, 2–8 (1997)
Friedman, R. & Hughes, A. L. Pattern and timing of gene duplication in animal genomes. Genome Res. 11, 1842–1847 (2001)
Hughes, A. L., da Silva, J. & Friedman, R. Ancient genome duplications did not structure the human Hox-bearing chromosomes. Genome Res. 11, 771–780 (2001)
Thornton, J. W. Evolution of vertebrate steroid receptors from an ancestral estrogen receptor by ligand exploitation and serial genome expansions. Proc. Natl Acad. Sci. USA 98, 5671–5676 (2001)
McLysaght, A., Hokamp, K. & Wolfe, K. H. Extensive genomic duplication during early chordate evolution. Nature Genet. 31, 200–204 (2002)
Panopoulou, G. et al. New evidence for genome-wide duplications at the origin of vertebrates using an amphioxus gene set and completed animal genomes. Genome Res. 13, 1056–1066 (2003)
Garcia-Fernandez, J. & Holland, P. W. Archetypal organization of the amphioxus Hox gene cluster. Nature 370, 563–566 (1994)
Artiguenave, F. et al. Genomic exploration of the hemiascomycetous yeasts: 2. Data generation and processing. FEBS Lett. 487, 13–16 (2000)
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980)
Li, W. H., Wu, C. I. & Luo, C. C. A new method for estimating synonymous and nonsynonymous rates of nucleotide substitution considering the relative likelihood of nucleotide and codon changes. Mol. Biol. Evol. 2, 150–174 (1985)
Pamilo, P. & Bianchi, N. O. Evolution of the Zfx and Zfy genes: rates and interdependence between the genes. Mol. Biol. Evol. 10, 271–281 (1993)
Acknowledgements
This work was supported by Consortium National de Recherche en Génomique. We thank T. Itami and S. Watabe for their gift of Takifugu blood samples; C. Nardon and M. Weiss for help with flow cytometry experiments; K. Howe for discussions regarding GAZE; R. Heilig for help with the annotation; the Centre Informatique National de l'Enseignement Supérieur for computer resources; and Gene-IT for assistance with the Biofacet software package.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Information
Tetraodon naming conventions, natural habitat and phylogeny (DOC 29 kb)
Supplementary Figure 1
Mitochondrial DNA sequence alignments for Tetraodon species identification (DOC 68 kb)
Supplementary Figure 2
Phylogenetic tree of Tetraodon family (DOC 64 kb)
Supplementary Figure 3
Flow cytometry results for genome size measurements (DOC 190 kb)
Supplementary Figure 4
Percentage G+C distribution and repeat content (DOC 59 kb)
Supplementary Figure 5
GAZE automaton and data transformation for genome annotation (DOC 87 kb)
Supplementary Figure 6
Distribution of exon and intron sizes (DOC 51 kb)
Supplementary Figure 7
Tetraodon catalogue of helical cytokines I and their receptors (DOC 277 kb)
Supplementary Figure 8
Tetraodon, Takifugu and zebrafish HOX gene clusters (DOC 58 kb)
Supplementary Figure 9
Gene Ontology annotation of proteins from 5 metazoan species (DOC 26 kb)
Supplementary Figure 10
Protein evolution in fish and mammals (DOC 144 kb)
Supplementary Figure 11
Synteny maps between Tetraodon and mouse (DOC 111 kb)
Supplementary Figure 12
Complete Oxford grid for Tetraodon-Human ortholog distribution (DOC 88 kb)
Supplementary Figure 13
Cladistic representation of chordate evolution (DOC 64 kb)
Supplementary Tables
Table 1) Sequencing statistics Table 2) Sequencing statistics per chromosome Table 3) Catalogue of transposable elements in the Tetraodon genome Table 4) Summary of evidence (coding segments) used to annotate the Tetraodon genome Table 5) Summary of evidence (signals) used to annotate the Tetraodon genome Table 6) Interpro domain content in four vertebrates and one urochrodate Table 7) Top 100 Interpro families in Tetraodon Table 8) Exofish analysis of five finished human chromosomes Table 9) Statistics on 904 new human genes Table 10) Distribution of the 904 new human genes on human chromosomes Table 11) Rates of DNA evolution in vertebrates Table 12) Expected probability that two Tetraodon chromosomes share the observed number of duplicated genes assuming a uniform distribution (DOC 506 kb)
Rights and permissions
About this article
Cite this article
Jaillon, O., Aury, JM., Brunet, F. et al. Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype. Nature 431, 946–957 (2004). https://doi.org/10.1038/nature03025
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03025
This article is cited by
-
IFNγ modulates the innate immune response via Toll-like receptors in green-spotted pufferfish (Tetraodon nigroviridis)
Aquaculture International (2024)
-
Costimulatory receptors in the channel catfish: CD28 family members and their ligands
Immunogenetics (2024)
-
A new genome assembly of an African weakly electric fish (Campylomormyrus compressirostris, Mormyridae) indicates rapid gene family evolution in Osteoglossomorpha
BMC Genomics (2023)
-
Reconstruction of hundreds of reference ancestral genomes across the eukaryotic kingdom
Nature Ecology & Evolution (2023)
-
Evolutionary differentiation of androgen receptor is responsible for sexual characteristic development in a teleost fish
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.