Abstract
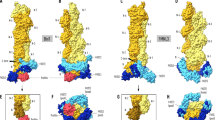
In the unactivated Limulus sperm, a 60-µm-long bundle of actin filaments crosslinked by the protein scruin is bent and twisted into a coil around the base of the nucleus. At fertilization, the bundle uncoils and fully extends in five seconds to support a finger of membrane known as the acrosomal process. This biological spring is powered by stored elastic energy and does not require the action of motor proteins or actin polymerization1. In a 9.5-Å electron cryomicroscopic structure of the extended bundle, we show that twist, tilt and rotation of actin–scruin subunits deviate widely from a ‘standard’ F-actin filament. This variability in structural organization allows filaments to pack into a highly ordered and rigid bundle in the extended state and suggests a mechanism for storing and releasing energy between coiled and extended states without disassembly.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Tilney, L. G. Actin filaments in the acrosomal reaction of Limulus sperm. J. Cell Biol. 64, 289–310 (1975)
Schmid, M. F., Jakana, J., Matsudaira, P. & Chiu, W. Imaging frozen, hydrated acrosomal bundle from Limulus sperm at 7 Å resolution with a 400 kV electron cryomicroscope. J. Mol. Biol. 230, 384–386 (1993)
DeRosier, D., Tilney, L. & Flicker, P. A change in the twist of the actin-containing filaments occurs during the extension of the acrosomal process in Limulus sperm. J. Mol. Biol. 137, 375–389 (1980)
Shin, J. H., Mahadevan, L., So, P. T. & Matsudaira, P. Bending stiffness of a crystalline actin bundle. J. Mol. Biol. 337, 255–261 (2004)
Shin, J. H., Mahadevan, L., Waller, G. S., Langsetmo, K. & Matsudaira, P. Stored elastic energy powers the 60-microm extension of the Limulus polyphemus sperm actin bundle. J. Cell Biol. 162, 1183–1188 (2003)
Gardel, M. L. et al. Elastic behavior of cross-linked and bundled actin networks. Science 304, 1301–1305 (2004)
Sherman, M. B. et al. The three-dimensional structure of the Limulus acrosomal process: a dynamic actin bundle. J. Mol. Biol. 294, 139–149 (1999)
Schmid, M. F. Cross-correlation and merging of crystallographic reflections derived from cryoelectron micrographs of 3D crystals: application to the Limulus acrosomal bundle. J. Struct. Biol. 144, 195–208 (2003)
Schmid, M. F., Agris, J. M., Jakana, J., Matsudaira, P. & Chiu, W. Three-dimensional structure of a single filament in the Limulus acrosomal bundle: scruin binds to homologous helix–loop–beta motifs in actin. J. Cell Biol. 124, 341–350 (1994)
Galkin, V. E. et al. The location of ubiquitin in Lethocerus arthrin. J. Mol. Biol. 325, 623–628 (2003)
Galkin, V. E. et al. The bacterial protein SipA polymerizes G-actin and mimics muscle nebulin. Nature Struct. Biol. 9, 518–521 (2002)
Galkin, V. E. et al. The utrophin actin-binding domain binds F-actin in two different modes: implications for the spectrin superfamily of proteins. J. Cell Biol. 157, 243–251 (2002)
Orlova, A. et al. Probing the structure of F-actin: cross-links constrain atomic models and modify actin dynamics. J. Mol. Biol. 312, 95–106 (2001)
Galkin, V. E., Orlova, A., Lukoyanova, N., Wriggers, W. & Egelman, E. H. Actin depolymerizing factor stabilizes an existing state of F-actin and can change the tilt of F-actin subunits. J. Cell Biol. 153, 75–86 (2001)
McGough, A., Pope, B., Chiu, W. & Weeds, A. Cofilin changes the twist of F-actin: implications for actin filament dynamics and cellular function. J. Cell Biol. 138, 771–781 (1997)
Jiang, W., Baker, M. L., Ludtke, S. J. & Chiu, W. Bridging the information gap: computational tools for intermediate resolution structure interpretation. J. Mol. Biol. 308, 1033–1044 (2001)
Kabsch, W., Mannherz, H. G., Suck, D., Pai, E. F. & Holmes, K. C. Atomic structure of the actin:DNase I complex. Nature 347, 37–44 (1990)
Holmes, K. C., Popp, D., Gebhard, W. & Kabsch, W. Atomic model of the actin filament. Nature 347, 44–49 (1990)
Dominguez, R. & Graceffa, P. Solution properties of TMR-actin: when biochemical and crystal data agree. Biophys. J. 85, 2073–2074 (2003)
Graceffa, P. & Dominguez, R. Crystal structure of monomeric actin in the ATP state. Structural basis of nucleotide-dependent actin dynamics. J. Biol. Chem. 278, 34172–34180 (2003)
Sablin, E. P. et al. How does ATP hydrolysis control actin's associations? Proc. Natl Acad. Sci. USA 99, 10945–10947 (2002)
Holmes, K. C., Angert, I., Kull, F. J., Jahn, W. & Schroder, R. R. Electron cryo-microscopy shows how strong binding of myosin to actin releases nucleotide. Nature 425, 423–427 (2003)
Borovikov, Y. S. et al. Fluorescence depolarization of actin filaments in reconstructed myofibers: the effect of S1 or pPDM-S1 on movements of distinct areas of actin. Biophys. J. 86, 3020–3029 (2004)
Otterbein, L. R., Graceffa, P. & Dominguez, R. The crystal structure of uncomplexed actin in the ADP state. Science 293, 708–711 (2001)
Egelman, E. H. Actin allostery again? Nature Struct. Biol. 8, 735–736 (2001)
Egelman, E. H., Francis, N. & DeRosier, D. J. F-actin is a helix with a random variable twist. Nature 298, 131–135 (1982)
Way, M., Sanders, M., Garcia, C., Sakai, J. & Matsudaira, P. Sequence and domain organization of scruin, an actin-cross-linking protein in the acrosomal process of Limulus sperm. J. Cell Biol. 128, 51–60 (1995)
Bullitt, E. S., DeRosier, D. J., Coluccio, L. M. & Tilney, L. G. Three-dimensional reconstruction of an actin bundle. J. Cell Biol. 107, 597–611 (1988)
Sanders, M., Way, M., Sakai, J. & Matsudaira, P. Characterization of the actin crosslinking properties of the scruin-calmodulin complex from the acrosomal process of Limulus sperm. J. Biol. Chem. 271, 2651–2657 (1996)
Mahadevan, L. & Matsudaira, P. Motility powered by supramolecular springs and ratchets. Science 288, 95–100 (2000)
Acknowledgements
This research is supported by the NCRR and NIGMS of NIH. We thank M. Baker for assistance in the helixhunter and foldhunter searches, and M. Dougherty for advice on graphical display.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary figure 1
Bundle Packing (DOC 441 kb)
Supplementary figure 2
Actin coordinates expressed as density (DOC 426 kb)
Supplementary movie
Transformation between the F-actin structure and the actin found in the bundle (MP4 2455 kb)
Rights and permissions
About this article
Cite this article
Schmid, M., Sherman, M., Matsudaira, P. et al. Structure of the acrosomal bundle. Nature 431, 104–107 (2004). https://doi.org/10.1038/nature02881
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02881
This article is cited by
-
Methyl labeling and TROSY NMR spectroscopy of proteins expressed in the eukaryote Pichia pastoris
Journal of Biomolecular NMR (2015)
-
Direct visualization of secondary structures of F-actin by electron cryomicroscopy
Nature (2010)
-
Structural polymorphism in F-actin
Nature Structural & Molecular Biology (2010)
-
Fission yeast IQGAP arranges actin filaments into the cytokinetic contractile ring
The EMBO Journal (2009)
-
The structure of bacterial ParM filaments
Nature Structural & Molecular Biology (2007)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.