Abstract
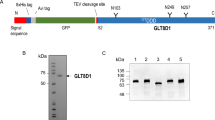
Lipid transfer proteins are important in membrane vesicle biogenesis and trafficking, signal transduction and immunological presentation processes1,2,3. The conserved and ubiquitous mammalian glycolipid transfer proteins (GLTPs) serve as potential regulators of cell processes mediated by glycosphingolipids, ranging from differentiation and proliferation to invasive adhesion, neurodegeneration and apoptosis4,5. Here we report crystal structures of apo-GLTP (1.65 Å resolution) and lactosylceramide-bound (1.95 Å) GLTP, in which the bound glycosphingolipid is sandwiched, after adaptive recognition, within a previously unknown two-layer all-α-helical topology. Glycosphingolipid binding specificity is achieved through recognition and anchoring of the sugar-amide headgroup to the GLTP recognition centre by hydrogen bond networks and hydrophobic contacts, and encapsulation of both lipid chains, in a precisely oriented manner within a ‘moulded-to-fit’ hydrophobic tunnel. A cleft-like conformational gating mechanism, involving two interhelical loops and one α-helix of GLTP, could enable the glycolipid chains to enter and leave the tunnel in the membrane-associated state. Mutation and functional analyses of residues in the glycolipid recognition centre and within the hydrophobic tunnel support a framework for understanding how GLTPs acquire and release glycosphingolipids during lipid intermembrane transfer and presentation processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rogers, D. P. & Bankaitis, V. A. Phospholipid transfer proteins and physiological functions. Int. Rev. Cytol. 197, 35–81 (2000)
Wirtz, K. W. A. Phospholipid transfer proteins revisited. Biochem. J. 324, 353–360 (1997)
Zhou, D. et al. Editing of CD1d-bound lipid antigens by endosomal lipid transfer proteins. Science 303, 523–527 (2004)
Hakomori, S. The glycosynapse. Proc. Natl Acad. Sci. USA 99, 225–232 (2002)
Dwek, R. A., Butters, T. D., Platt, F. M. & Zitzmann, N. Targeting glycosylation as a therapeutic approach. Nature Rev. Drug Discov. 1, 65–75 (2002)
Metz, R. J. & Radin, N. S. Purification and properties of a cerebroside transfer protein. J. Biol. Chem. 257, 12901–12907 (1982)
Abe, A. & Sasaki, T. Purification and some properties of the glycolipid transfer protein from pig brain. J. Biol. Chem. 260, 11231–11239 (1985)
Brown, R. E., Jarvis, K. L. & Hyland, K. J. Purification and characterization of glycolipid transfer protein from bovine brain. Biochim. Biophys. Acta 1044, 77–83 (1990)
Yamada, K., Abe, A. & Sasaki, T. Specificity of the glycolipid transfer protein from pig brain. J. Biol. Chem. 260, 4615–4621 (1985)
Lin, X., Mattjus, P., Pike, H. M., Windebank, A. J. & Brown, R. E. Cloning and expression of glycolipid transfer protein from bovine and porcine brain. J. Biol. Chem. 275, 5104–5110 (2000)
Brodersen, P. et al. Knockout of Arabidopsis accelerated-cell-death11 encoding a sphingosine transfer protein causes activation of programmed cell death and defense. Genes Dev. 16, 490–502 (2002)
Mattjus, P., Turcq, B., Pike, H. M., Molotkovsky, J. G. & Brown, R. E. Glycolipid intermembrane transfer is accelerated by HET-C2, a filamentous fungus gene product involved in the cell–cell incompatibility response. Biochemistry 42, 535–542 (2003)
Tsujishita, Y. & Hurley, J. H. Structure and lipid transport mechanism of a StAR-related domain. Nature Struct. Biol. 7, 408–414 (2000)
Roderick, S. L. et al. Structure of human phosphatidylcholine transfer protein in complex with its ligand. Nature Struct. Biol. 9, 507–511 (2002)
Min, K. C., Kovall, R. A. & Hendrickson, W. A. Crystal structure of α-tocopherol transfer protein bound to it ligand: implications for ataxia with vitamin E deficiency. Proc. Natl Acad. Sci. USA 100, 14713–14718 (2003)
Gadola, S. D. et al. Structure of human CD1b with bound ligands at 2.3 Å, a maze of alkyl chains. Nature Immunol. 3, 721–726 (2002)
Zajonc, D. M., Elsliger, M. A., Teyton, L. & Wilson, I. A. Crystal structure of CD1a in complex with a sulfatide self antigen at a resolution of 2.15 Å. Nature Immunol. 4, 808–815 (2003)
Mahfoud, R. et al. Identification of a common sphingolipid-binding domain in Alzheimer, prion, and HIV-1 proteins. J. Biol. Chem. 277, 11292–11296 (2002)
Schubert Wright, C., Zhao, Q. & Rastinejad, F. Structural analysis of lipid complexes of GM2-activator protein. J. Mol. Biol. 331, 951–964 (2003)
Weis, W. I. & Drickamer, K. Structural basis of lectin–carbohydrate interactions. Annu. Rev. Biochem. 65, 441–473 (1996)
Feinberg, H., Mitchell, D. A., Drickamer, K. & Weis, W. I. Structural basis for selective recognition of oligosaccharides by DC-SIGN and DC-SIGNR. Science 294, 2163–2166 (2001)
Wimley, W. W. & White, S. H. Experimentally determined hydrophobicity scale for proteins at membrane interfaces. Nature Struct. Biol. 3, 842–848 (1996)
Killian, J. A. & von Heijne, G. How proteins adapt to a membrane–water interface. Trends Biochem. Sci. 25, 429–434 (2000)
Doublié, S. Preparation of selenomethionyl proteins for phase determination. Methods Enzymol. 276, 523–530 (1997)
Li, X.-M., Momsen, M. M., Brockman, H. L. & Brown, R. E. Lactosylceramide: effect of acyl chain structure on phase behavior and molecular packing. Biophys. J. 83, 1535–1546 (2002)
Hendrickson, W. A. Determination of macromolecular structures from anomalous diffraction of synchrotron radiation. Science 254, 51–58 (1991)
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999)
Lamzin, V. S. & Wilson, K. S. Automated refinement of protein models. Acta Crystallogr. D 49, 129–149 (1993)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1997)
Acknowledgements
We thank the personnel at SBC beamline 19BM of the Advanced Photon Source beamline staff for assistance with data collection from multiwavelength anomalous dispersion; A. Serganov for technical support; X. Lin, T. Chung and H. Pike for their contributions to the cloning and expression of the recombinant human GLTP; X.-M. Li for synthesizing and purifying N-18:1 lactosylceramide; A. J. Windebank for help with DNA sequencing at the Mayo Molecular Biology Core Facility; T. Burghardt for help with recording near-ultraviolet CD spectra; and S. Venyaminov in the Franklyn Prendergast laboratory for recording the far-ultraviolet CD spectra. This research was supported by NIH and the Hormel Foundation. Use of the ANL SBC beamlines at the APS was supported by the US Department of Energy, Office of Energy Research.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Table S1
X-ray data collection and refinement statistics. (RTF 47 kb)
Supplementary Figure S1
A two-layer topology of the γ-helices. (PDF 106 kb)
Supplementary Figure S2
Structure of the lactosylceramide-GLTP complex. (PDF 1910 kb)
Supplementary Figure S3
Schematic showing the hydrogen bonding between lactosylceramide and protein side chains in the complex. (PDF 98 kb)
Supplementary Figure S4
Glycolipid transfer activities of wtGLTP and representative GLTPs with point mutations. (PDF 220 kb)
Supplementary Figure S5
Superposition of wild type GLTP and the D48V mutant. (PDF 917 kb)
Supplementary Figure S6
Far UV CD spectra of GLTP mutants. (PDF 116 kb)
Supplementary Figure S7
Near UV CD spectra of GLTP mutants. (PDF 118 kb)
Supplementary Figure S8
Electron density map, sequence and secondary structure elements for apo-GLTP. (PDF 2448 kb)
Supplementary Figure S9
Omit electron density map for a ligand. (PDF 733 kb)
Rights and permissions
About this article
Cite this article
Malinina, L., Malakhova, M., Teplov, A. et al. Structural basis for glycosphingolipid transfer specificity. Nature 430, 1048–1053 (2004). https://doi.org/10.1038/nature02856
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02856
This article is cited by
-
Mapping QTL for leaf pigment content at dynamic development stage and analyzing Meta-QTL in rice
Euphytica (2021)
-
Non-vesicular trafficking by a ceramide-1-phosphate transfer protein regulates eicosanoids
Nature (2013)
-
Structural motifs recurring in different folds recognize the same ligand fragments
BMC Bioinformatics (2009)
-
Human glycolipid transfer protein (GLTP) genes: organization, transcriptional status and evolution
BMC Genomics (2008)
-
Taxi service for lipids
Nature (2007)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.