Abstract
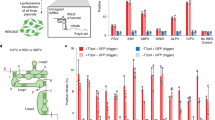
Recent studies on the control of specific metabolic pathways in bacteria have documented the existence of entirely RNA-based mechanisms for controlling gene expression. These mechanisms involve the modulation of translation, transcription termination or RNA self-cleavage through the direct interaction of specific intracellular metabolites and RNA sequences1,2,3,4. Here we show that an analogous RNA-based gene regulation system can effectively be designed for mammalian cells via the incorporation of sequences encoding self-cleaving RNA motifs5 into the transcriptional unit of a gene or vector. When correctly positioned, the sequences lead to potent inhibition of gene or vector expression, owing to the spontaneous cleavage of the RNA transcript. Administration of either oligonucleotides complementary to regions of the self-cleaving motif or a specific small molecule results in the efficient induction of gene expression, owing to inhibition of self-cleavage of the messenger RNA. Efficient regulation of transgene expression is shown in a variety of mammalian cell lines and live animals. In conjunction with other emerging technologies6, this methodology may be particularly applicable to the development of gene regulation systems tailored to any small inducer molecule, and provide a novel means of biological sensing in vivo that may have an important application in the regulated delivery of protein therapeutics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Winkler, W., Nahvi, A. & Breaker, R. R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 419, 952–956 (2002)
Winkler, W. C., Nahvi, A., Roth, A., Collins, J. A. & Breaker, R. R. Control of gene expression by a natural metabolite-responsive ribozyme. Nature 428, 281–286 (2004)
Mandal, M. & Breaker, R. R. Adenine riboswitches and gene activation by disruption of a transcription terminator. Nature Struct. Mol. Biol. 11, 29–35 (2004)
Cech, T. R. RNA finds a simpler way. Nature 428, 263–264 (2004)
Cech, T. R. Nobel lecture. Self-splicing and enzymatic activity of an intervening sequence RNA from Tetrahymena. Biosci. Rep. 10, 239–261 (1990)
Silverman, S. K. Rube Goldberg goes (ribo)nuclear? Molecular switches and sensors made from RNA. RNA 9, 377–383 (2003)
Ory, D. S., Neugeboren, B. A. & Mulligan, R. C. A stable human-derived packaging cell line for production of high titer retrovirus/vesicular stomatitis virus G pseudotypes. Proc. Natl Acad. Sci. USA 93, 11400–11406 (1996)
Rojas, A. A. et al. Hammerhead-mediated processing of satellite pDo500 family transcripts from Dolichopoda cave crickets. Nucleic Acids Res. 28, 4037–4043 (2000)
Ferbeyre, G., Smith, J. M. & Cedergren, R. Schistosome satellite DNA encodes active hammerhead ribozymes. Mol. Cell. Biol. 18, 3880–3888 (1998)
Hertel, K. J. et al. Numbering system for the hammerhead. Nucleic Acids Res. 20, 3252 (1992)
Ruffner, D. E., Stormo, G. D. & Uhlenbeck, O. C. Sequence requirements of the hammerhead RNA self-cleavage reaction. Biochemistry 29, 10695–10702 (1990)
Chowrira, B. M., Pavco, P. A. & McSwiggen, J. A. In vitro and in vivo comparison of hammerhead, hairpin and hepatitis delta virus self-processing ribozyme cassettes. J. Biol. Chem. 269, 25856–25864 (1994)
Zillmann, M., Limauro, S. E. & Goodchild, J. In vitro optimization of truncated stem-loop II variants of the hammerhead ribozyme for cleavage in low concentrations of magnesium under non-turnover conditions. RNA 3, 734–747 (1997)
Conaty, J., Hendry, P. & Lockett, T. Selected classes of minimised hammerhead ribozyme have very high cleavage rates at low Mg2+ concentration. Nucleic Acids Res. 27, 2400–2407 (1999)
Hermann, T. & Westhof, E. Aminoglycoside binding to the hammerhead ribozyme: a general model for the interaction of cationic antibiotics with RNA. J. Mol. Biol. 276, 903–912 (1998)
Jenne, A. et al. Rapid identification and characterization of hammerhead-ribozyme inhibitors using fluorescence-based technology. Nature Biotechnol. 19, 56–61 (2001)
Murray, J. B. & Arnold, J. R. Antibiotic interactions with the hammerhead ribozyme:tetracyclines as a new class of hammerhead inhibitor. Biochem. J. 317, 855–860 (1996)
Stage, T. K., Hertel, K. J. & Uhlenbeck, O. C. Inhibition of the hammerhead ribozyme by neomycin. RNA 1, 95–101 (1995)
Tor, Y., Hermann, T. & Westhof, E. Deciphering RNA recognition: aminoglycoside binding to the hammerhead ribozyme. Chem. Biol. 5, R277–R283 (1998)
von Ahsen, U., Davies, J. & Schroeder, R. Antibiotic inhibition of group I ribozyme function. Nature 353, 368–370 (1991)
Braasch, D. A. & Corey, D. R. Novel antisense and peptide nucleic acid strategies for controlling gene expression. Biochemistry 41, 4503–4510 (2002)
Morcos, P. A. Achieving efficient delivery of morpholino oligos in cultured cells. Genesis 30, 94–102 (2001)
Aszalos, A., Lemanski, P., Robison, R., Davis, S. & Berk, B. Identification of antibiotic 1037 as toyocamycin. J. Antibiot. (Tokyo) 19, 285 (1966)
Contag, P. R., Olomu, I. N., Stevenson, D. K. & Contag, C. H. Bioluminescent indicators in living mammals. Nature Med. 4, 245–247 (1998)
Gossen, M. & Bujard, H. Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc. Natl Acad. Sci. USA 89, 5547–5551 (1992)
Rivera, V. M. et al. A humanized system for pharmacologic control of gene expression. Nature Med. 2, 1028–1032 (1996)
Suhr, S. T., Gil, E. B., Senut, M. C. & Gage, F. H. High level transactivation by a modified Bombyx ecdysone receptor in mammalian cells without exogenous retinoid X receptor. Proc. Natl Acad. Sci. USA 95, 7999–8004 (1998)
Wang, Y., O'Malley, B. W. Jr, Tsai, S. Y. & O'Malley, B. W. A regulatory system for use in gene transfer. Proc. Natl Acad. Sci. USA 91, 8180–8184 (1994)
Breaker, R. R. Engineered allosteric ribozymes as biosensor components. Curr. Opin. Biotechnol. 13, 31–39 (2002)
Wilson, D. S. & Szostak, J. W. In vitro selection of functional nucleic acids. Annu. Rev. Biochem. 68, 611–647 (1999)
Acknowledgements
We thank K. Salehi-Ashtlani and J. Szostak for helpful discussions, Y. Tang and R. Weissleder for help with imaging experiments performed during the early course of the work, and M. Chung for her technical assistance. This work was supported by grants from AMGEN and L'Association Francaise contre les Myopathies (AFM). R.C.M. is an AMGEN consultant and equity holder.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Funding for this work was provided by AMGEN corporation and L'Association Francaise contre les Myopathies (AFM). R.C.M. holds a non-paying consultant position and AMGEN equity.
Supplementary information
Supplementary Table
Survey of ability of different ‘self-cleaving’ ribozymes to function in mammalian cells. Including a list of references. (PDF 106 kb)
Rights and permissions
About this article
Cite this article
Yen, L., Svendsen, J., Lee, JS. et al. Exogenous control of mammalian gene expression through modulation of RNA self-cleavage. Nature 431, 471–476 (2004). https://doi.org/10.1038/nature02844
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02844
This article is cited by
-
Control of mammalian gene expression by modulation of polyA signal cleavage at 5′ UTR
Nature Biotechnology (2024)
-
Ribo-On and Ribo-Off tools using a self-cleaving ribozyme allow manipulation of endogenous gene expression in C. elegans
Communications Biology (2023)
-
Regulated control of gene therapies by drug-induced splicing
Nature (2021)
-
High-throughput identification of synthetic riboswitches by barcode-free amplicon-sequencing in human cells
Nature Communications (2020)
-
Optoribogenetic control of regulatory RNA molecules
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.