Abstract
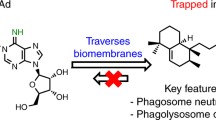
Fifty million new infections with Mycobacterium tuberculosis occur annually, claiming 2–3 million lives from tuberculosis worldwide1. Despite the apparent lack of significant genetic heterogeneity between strains of M. tuberculosis2,3, there is mounting evidence that considerable heterogeneity exists in molecules important in disease pathogenesis. These differences may manifest in the ability of some isolates to modify the host cellular immune response, thereby contributing to the observed diversity of clinical outcomes4,5,6,7. Here we describe the identification and functional relevance of a highly biologically active lipid species—a polyketide synthase-derived phenolic glycolipid (PGL) produced by a subset of M. tuberculosis isolates belonging to the W-Beijing family8 that show ‘hyperlethality’ in murine disease models. Disruption of PGL synthesis results in loss of this hypervirulent phenotype without significantly affecting bacterial load during disease. Loss of PGL was found to correlate with an increase in the release of the pro-inflammatory cytokines tumour-necrosis factor-α and interleukins 6 and 12 in vitro. Furthermore, the overproduction of PGL by M. tuberculosis or the addition of purified PGL to monocyte-derived macrophages was found to inhibit the release of these pro-inflammatory mediators in a dose-dependent manner.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Tiruviluamala, P. & Reichman, L. B. Tuberculosis. Annu. Rev. Publ. Health 23, 403–426 (2002)
Sreevatsan, S. et al. Restricted structural gene polymorphism in the Mycobacterium tuberculosis complex indicates evolutionarily recent global dissemination. Proc. Natl Acad. Sci. USA 94, 9869–9874 (1997)
Fleischmann, R. D. et al. Whole-genome comparison of Mycobacterium tuberculosis clinical and laboratory strains. J. Bacteriol. 184, 5479–5490 (2002)
North, R. J., Ryan, L., LaCource, R., Mogues, T. & Goodrich, M. E. Growth rate of mycobacteria in mice as an unreliable indicator of mycobacterial virulence. Infect. Immun. 67, 5483–5485 (1999)
Manca, C. et al. Mycobacterium tuberculosis CDC1551 induces a more vigorous host response in vivo and in vitro, but is not more virulent than other clinical isolates. J. Immunol. 162, 6740–6746 (1999)
Manca, C. et al. Virulence of a Mycobacterium tuberculosis clinical isolate in mice is determined by failure to induce Th1 type immunity and is associated with induction of IFN-α/β. Proc. Natl Acad. Sci. USA 98, 5752–5757 (2001)
Valway, S. E. et al. An outbreak involving extensive transmission of a virulent strain of Mycobacterium tuberculosis. N. Engl. J. Med. 338, 633–639 (1998)
Bifani, P. J., Mathema, B., Kurepina, N. E. & Kreiswirth, B. N. Global dissemination of the Mycobacterium tuberculosis W-Beijing family strains. Trends Microbiol. 10, 45–52 (2002)
Glynn, J. R., Whiteley, J., Bifani, P. J., Kremer, K. & van Soolingen, D. Worldwide occurrence of Beijing/W strains of Mycobacterium tuberculosis: a systematic review. Emerg. Infect. Dis. 8, 843–849 (2002)
Cole, S. T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998)
Manca, C. et al. Differential monocyte activation underlies strain specific M. tuberculosis pathogenesis. Infect. Immun. (in the press)
Cox, J. S., Chen, B., McNeil, M. & Jacobs, W. R. Jr Complex lipid determines tissue-specific replication of Mycobacterium tuberculosis in mice. Nature 402, 79–83 (1999)
Sirakova, T. D., Thirumala, A. K., Dubey, V. S., Sprecher, H. & Kolattukudy, P. E. The Mycobacterium tuberculosis pks2 gene encodes the synthase for the hepta- and octamethyl-branched fatty acids required for sulfolipid synthesis. J. Biol. Chem. 276, 16833–16839 (2001)
Constant, P. et al. Role of the pks15/1 gene in the biosynthesis of phenolglycolipids in the Mycobacterium tuberculosis complex. Evidence that all strains synthesize glycosylated p-hydroxybenzoic methly esters and that strains devoid of phenolglycolipids harbor a frameshift mutation in the pks15/1 gene. J. Biol. Chem. 277, 38148–38158 (2002)
Marmiesse, M. et al. Macro-array and bioinformatic analyses reveal mycobacterial ‘core’ genes, variation in the ESAT-6 gene family and new phylogenetic markers for the Mycobacterium tuberculosis complex. Microbiol. 150, 483–496 (2004)
Kolattukudy, P. E., Fernandes, N. D., Azad, A. K., Fitzmaurice, A. M. & Sirakova, T. D. Biochemistry and molecular genetics of cell-wall lipid biosynthesis in mycobacteria. Mol. Microbiol. 24, 263–270 (1997)
Vergne, I. I. & Daffe, M. Interaction of mycobacterial glycolipids with host cells. Front. Biosci. 3, 865–876 (1998)
Hunter, S. W. & Brennan, P. J. A novel phenolic glycolipid from Mycobacterium leprae possibly involved in immunogenicity and pathogenicity. J. Bacteriol. 147, 728–735 (1981)
Mehra, V., Brennan, P. J., Rada, E., Convit, J. & Bloom, B. R. Lymphocyte suppression in leprosy induced by unique M. leprae glycolipid. Nature 308, 194–196 (1984)
Fournie, J.-J., Adams, E., Mullins, R. J. & Basten, A. Inhibition of human lymphoproliferative responses by mycobacterial phenolic glycolipids. Infect. Immun. 57, 3653–3659 (1989)
Vachula, K., Holzer, T. J. & Andersen, B. R. suppression of monocyte oxidative responses by phenolic glycolipid 1 of Mycobacterium leprae. J. Immunol. 142, 1696–1701 (1989)
Silva, C. L., Faccioli, L. H. & Foss, N. T. Suppression of human monocyte cytokine release by phenolic glycolipid-1 of Mycobacterium leprae. Int. J. Lepr. 61, 107–108 (1993)
Hashimoto, K. et al. Mycobacterium leprae infection in monocyte-derived dendritic cells and its influence on antigen-presenting function. Infect. Immun. 70, 5167–5176 (2002)
Ng, V. et al. Role of the cell wall phenolic glycolipid-1 in the peripheral nerve predilection of Mycobacterium leprae. Cell 103, 511–524 (2000)
Stover, C. K. et al. New use of BCG for recombinant vaccines. Nature 351, 456–460 (1991)
Gordon, S. Alternative activation of macrophages. Nature Rev. Immunol. 3, 23–35 (2003)
Garbe, T. R. et al. Transformation of mycobacterial species using hygromycin resistance as selectable marker. Microbiol. 140, 133–138 (1994)
Pelicic, V. et al. Efficient allelic exchange and transposon mutagenesis in Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 94, 10955–10960 (1997)
O'Gaora, P. et al. Mycobacteria as immunogens: development of expression vectors for use in multiple mycobacterial species. Med. Princ. Prac. 6, 91–96 (1997)
Slayden, R. A. & Barry, C. E. III in Mycobacterium tuberculosis protocols (eds Parish, T. & Stoker, N. G.) 229–245 (Humana, New Jersey, 2001)
Acknowledgements
The authors wish to thank J. Gonzales and M. Goodwin for their assistance with animal studies and NMR spectroscopy, respectively. G.K. is supported by grants from the NIH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Reed, M., Domenech, P., Manca, C. et al. A glycolipid of hypervirulent tuberculosis strains that inhibits the innate immune response. Nature 431, 84–87 (2004). https://doi.org/10.1038/nature02837
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02837
This article is cited by
-
Non-specific effects of inactivated Mycobacterium bovis oral and parenteral treatment in a rabbit scabies model
Veterinary Research (2024)
-
Selective delipidation of Mycobacterium bovis BCG retains antitumor efficacy against non-muscle invasive bladder cancer
Cancer Immunology, Immunotherapy (2023)
-
Mycobacterium tuberculosis and its clever approaches to escape the deadly macrophage
World Journal of Microbiology and Biotechnology (2023)
-
Early alveolar macrophage response and IL-1R-dependent T cell priming determine transmissibility of Mycobacterium tuberculosis strains
Nature Communications (2022)
-
Inhibition of the Niemann-Pick C1 protein is a conserved feature of multiple strains of pathogenic mycobacteria
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.