Abstract
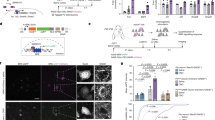
Animal microRNAs (miRNAs) are gene regulatory factors that prevent the expression of specific messenger RNA targets by binding to their 3′ untranslated region1,2,3. The Caenorhabditis elegans lsy-6 miRNA (for lateral symmetry defective) is required for the left/right asymmetric expression of guanyl cyclase (gcy) genes in two chemosensory neurons termed ASE left (ASEL) and ASE right (ASER)4,5. The asymmetric expression of these putative chemoreceptors in turn correlates with the functional lateralization of the ASE neurons6. Here we find that a mutation in the die-1 zinc-finger transcription factor disrupts both the chemosensory laterality and left/right asymmetric expression of chemoreceptor genes in the ASE neurons. die-1 controls chemosensory laterality by activating the expression of lsy-6 specifically in ASEL, but not in ASER, where die-1 expression is downregulated through two sites in its 3′ untranslated region. These two sites are complementary to mir-273, a previously uncharacterized miRNA, whose expression is strongly biased towards ASER. Forced bilateral expression of mir-273 in ASEL and ASER causes a loss of asymmetric die-1 expression and ASE laterality. Thus, an inverse distribution of two sequentially acting miRNAs in two bilaterally symmetric neurons controls laterality of the nematode chemosensory system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993)
Wightman, B., Ha, I. & Ruvkun, G. Post-transcriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75, 855–862 (1993)
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004)
Johnston, R. J. & Hobert, O. A microRNA controlling left/right neuronal asymmetry in Caenorhabditis elegans. Nature 426, 845–849 (2003)
Hobert, O., Johnston, R. J. Jr & Chang, S. Left-right asymmetry in the nervous system: the Caenorhabditis elegans model. Nature Rev. Neurosci. 3, 629–640 (2002)
Pierce-Shimomura, J. T., Faumont, S., Gaston, M. R., Pearson, B. J. & Lockery, S. R. The homeobox gene lim-6 is required for distinct chemosensory representations in C. elegans. Nature 410, 694–698 (2001)
Chang, S., Johnston, R. J. Jr & Hobert, O. A transcriptional regulatory cascade that controls left/right asymmetry in chemosensory neurons of C. elegans. Genes Dev. 17, 2123–2137 (2003)
Heid, P. J. et al. The zinc-finger protein DIE-1 is required for late events during epithelial cell rearrangement in C. elegans. Dev. Biol. 236, 165–180 (2001)
Bulyk, M. L. Computational prediction of transcription-factor binding site locations. Genome Biol. 5, 201 (2003)
Grad, Y. et al. Computational and experimental identification of C. elegans microRNAs. Mol. Cell. 11, 1253–1263 (2003)
Stark, A., Brennecke, J., Russell, R. B. & Cohen, S. M. Identification of Drosophila microRNA targets. PLoS Biol. 1, 1–13 (2003)
Doench, J. G., Petersen, C. P. & Sharp, P. A. siRNAs can function as miRNAs. Genes Dev. 17, 438–442 (2003)
Muhr, J., Andersson, E., Persson, M., Jessell, T. M. & Ericson, J. Groucho-mediated transcriptional repression establishes progenitor cell pattern and neuronal fate in the ventral neural tube. Cell 104, 861–873 (2001)
Hobert, O. PCR fusion-based approach to create reporter gene constructs for expression analysis in transgenic C. elegans. Biotechniques 32, 728–730 (2002)
Bargmann, C. I. & Horvitz, H. R. Chemosensory neurons with overlapping functions direct chemotaxis to multiple chemicals in C. elegans. Neuron 7, 729–742 (1991)
Yu, S., Avery, L., Baude, E. & Garbers, D. L. Guanylyl cyclase expression in specific sensory neurons: a new family of chemosensory receptors. Proc. Natl Acad. Sci. USA 94, 3384–3387 (1997)
Palmer, R. E., Inoue, T., Sherwood, D. R., Jiang, L. I. & Sternberg, P. W. Caenorhabditis elegans cog-1 locus encodes GTX/Nkx6.1 homeodomain proteins and regulates multiple aspects of reproductive system development. Dev. Biol. 252, 202–213 (2002)
Acknowledgements
We thank C. Antonio for the ot100 allele, J.Hardin for die-1 reagents, Q. Chen and S. Narula for expert technical assistance, J. Tien for help with mapping ot26, I. Greenwald for advice, L. Johnston, A. Lanjuin, P. Sengupta, I. Greenwald and F. Slack for reading the manuscript. S.C. was funded by an NIH predoctoral fellowship, R.J.J. by an NSF predoctoral fellowship, and C.F.-J. by a American Heart Association predoctoral fellowship. O.H. was supported by grants from the NIH, the McKnight Foundation and the Irma T.Hirschl Trust.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Data
This file contains additional Methods, references and legends to Supplementary (DOC 72 kb)
Supplementary Figure 1
ot26 is an allele of the Zn finger transcription factor DIE-1 which acts in ASE to affect gcy gene expression. (JPG 28 kb)
Supplementary Figure 2
die-1 controls lim-6 expression. (JPG 27 kb)
Supplementary Figure 3
Identification of mir-273 complementary sites and the structure of the mir-273 precursor. (JPG 126 kb)
Supplementary Figure 4
Bilateral expression of mir-273 disrupts ASE laterality. (JPG 32 kb)
Rights and permissions
About this article
Cite this article
Chang, S., Johnston, R., Frøkjær-Jensen, C. et al. MicroRNAs act sequentially and asymmetrically to control chemosensory laterality in the nematode. Nature 430, 785–789 (2004). https://doi.org/10.1038/nature02752
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02752
This article is cited by
-
Differential expression of microRNAs in the human fetal left and right cerebral cortex
Molecular Biology Reports (2020)
-
Integrated microRNA and mRNA analysis in the pathogenic filamentous fungus Trichophyton rubrum
BMC Genomics (2018)
-
Transcriptional and Epigenetic Control of Mammalian Olfactory Epithelium Development
Molecular Neurobiology (2018)
-
miRNAs as Biomarkers and Therapeutic Targets in Non-Small Cell Lung Cancer: Current Perspectives
Targeted Oncology (2017)
-
Maintenance of postmitotic neuronal cell identity
Nature Neuroscience (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.