Abstract
The finished sequence of human chromosome 10 comprises a total of 131,666,441 base pairs. It represents 99.4% of the euchromatic DNA and includes one megabase of heterochromatic sequence within the pericentromeric region of the short and long arm of the chromosome. Sequence annotation revealed 1,357 genes, of which 816 are protein coding, and 430 are pseudogenes. We observed widespread occurrence of overlapping coding genes (either strand) and identified 67 antisense transcripts. Our analysis suggests that both inter- and intrachromosomal segmental duplications have impacted on the gene count on chromosome 10. Multispecies comparative analysis indicated that we can readily annotate the protein-coding genes with current resources. We estimate that over 95% of all coding exons were identified in this study. Assessment of single base changes between the human chromosome 10 and chimpanzee sequence revealed nonsense mutations in only 21 coding genes with respect to the human sequence.
Similar content being viewed by others
Main
We report here the final clone map and sequence assembly of human chromosome 10. This metacentric chromosome accounts for 4.5% of the genome and is best known for harbouring the PTEN tumour suppressor gene and the RET proto-oncogene.
With the human genome sequence in hand, the task ahead is to identify the different units of genetic information embedded in the sequence and understand their function both at the molecular and cellular level. In this study we address the former by reporting a comprehensive annotation of manually inspected gene structures and their correlation to sequence variation and other features of the genomic sequence. The annotation process is assessed by comparative analysis to the genome sequence of two rodents, Mus musculus and Rattus norvegicus, and three fishes, Tetraodon nigroviridis, Fugu rubripes and Danio rerio. Finally, we report our preliminary findings on the distribution of single base differences along human chromosome 10 in comparison to the chimpanzee genome.
The clone map and finished sequence
A clone map spanning the euchromatic regions of the short (p) and long (q) arm of human chromosome 10 was assembled by restriction fingerprint and sequence-tagged site (STS) content analysis1. We identified clones by screening approximately 85 genomic equivalents of P1-derived artificial chromosome (PAC), bacterial artificial chromosome (BAC), yeast artificial chromosome (YAC), cosmid and fosmid libraries. The tiling path consists of 1,144 minimally overlapping clones (Supplementary Table S1) organized into 12 contigs (Table 1). Contig-1340 spans the entire p arm and harbours the boundary between euchromatin and heterochromatin with the proximal 250 kilobases (kb) extending into pericentromeric satellite repeats. In a detailed study of this region two further clones carrying satellite 3 sequences, contig-2069, were identified and mapped by pulse field gel electrophoresis (PFGE) ∼50 kb proximal to contig-1340 (ref. 2). Similarly, contig-43 harbours the q-arm boundary with the proximal 240 kb composed of satellite repeats3. In addition, clone RP11-745D9, 94% of which consists of α-satellite repeats, was arbitrarily placed proximal to contig-43 as it was suggested to map to human chromosome 10 (E. Eichler, personal communication). A 9.75-megabase (Mb) PFGE map spanning the chromosome 10 centromere4 places the core α-satellite block (D10Z1) ∼0.2 Mb distal to contig-102 and 1 Mb proximal to contig-43.
In contrast with the p arm, nine gaps remain in the clone map of the q arm. Five of them are clustered within a ∼4-Mb region (Table 1 and Fig. 1; see also Supplementary Fig. S1 for a detailed view). Our inability to walk across these five gaps was due to the extensive segmental duplications in this region (Fig. 2); however, we obtained a size estimate by fluorescent in situ hybridization (FISH) of RP11-172C24 (AL512595) and RP11-13E1 (AC013284) on metaphase chromosomes. We sized the remaining four gaps by FISH with clones immediately flanking each gap to extended DNA fibres. No gap was estimated to be larger than 50 kb in size. Altogether the euchromatic gaps account for no more than 840 kb (Table 1). Finally, we defined the location of both telomeres. Clone RP11-631M1 (AL713922) ends ∼20 kb away from the telomeric repeats of the p arm based on the telomeric half-YAC XX-YAC22O3 (http://www.wistar.upenn.edu/Riethman/). At the end of the q arm (qtel), clone XX-YAC2136 (BX322534) contains part of the telomeric repeat block.
The current sequence assembly (v1.1) and that used in the analysis presented in this study (v1.0) are available at http://www.sanger.ac.uk/HGP/Chr10. Tracks from top to bottom are: (1) the scale bar (in Mb); (2) the sequence map of human chromosome 10 represented by a black solid bar interrupted by clone and sequence gaps (grey); (3) syntenic blocks in the mouse (top track) and the rat (bottom track) where blocks are colour-coded per chromosome and labelled with the chromosome number and coordinates (Mb) (for example, 2: 3.1–11.3 Mb; unordered sequence contigs are tagged as random); (4) predicted CpG islands (brown); (5) regions of sequence homology to Fugu (blue), zebrafish (dark blue) and Tetraodon (dark pink); and (6) protein-coding genes. Gene names are in dark blue for the known and black for the novel CDS.
For a larger version of this figure please download the pdf.
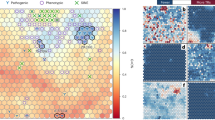
a, Interchromosomal duplications across chromosome 10 showing blocks of 10 kb or greater. Duplicated regions are colour-coded per chromosome and indicated as lines (thickness is proportional to physical distance). Each chromosome other than 10 is represented by an open black rectangle with a black line representing the centromere. b, 10q11:10q22:10q23.1:10q23.3 intrasegmental duplications. Top row, ideogram of chromosome 10; second row, positions of members of the six main gene families outside 10q11 (colour-coded per gene class; arrows indicate transcription); third row, intrachromosomal duplications across the whole chromosome showing blocks of 10 kb or greater; bottom row, enlarged view of the 10q11 region. Yellow bars represent sequence (bottom row for sequences submitted in reverse orientation) contigs between AL358394 and AL589794 (complete clone list in Supplementary Fig. S1). From left to right the members (where parentheses indicate members only appearing in the enlarged section) of each family are: GLUD family, GLUDP5, (GLUDP7, GLUDP8, GLUDP6, GLUDP2), GLUDP3, GLUD1; CTSLL family, (CTSLL5, CTSLL7, CTSLL2, CTSLL3, CTSLL4), CTSLL6, CTSLL1; ANXA family, (ANXA8L1, ANXA8L2, ANXA8), ANXA2P3, ANXA7, ANXA11; CTGLF family, (CTGLF10P, CTGLF1, CTGLF13P, CTGLF7, CTGLF11P, CTGLF6, CTGLF9P, CTGLF12P, CTGLF5, CTGLF4, CTGLF3), CTGLF2; BMS1L family, BMS1L, (BMSILP1, BMSILP2, BMSILP6, BMSILP5, BMSILP7), BMSILP4, BMSILP3; FAM25 family, (FAM25E, FAM25B, FAM25HP, FAM25G, FAM25C, FAM25D), FAM25A. ANXA7 and ANXA11 were included owing to their proximity to the 10q22 and 10q23.1 locus, respectively. Seven CTGLF1 paralogues (Supplementary Table S3) were annotated as novel genes for consistency but probably represent expressed pseudogenes.
Each clone of the tiling path was subjected to random subcloning and sequencing at either the Genome Therapeutics Corporation (GENE) or the Sanger Institute—the initial draft sequence of a few clones was carried out by other centres that are credited in the corresponding submissions to the EMBL/GenBank/DDBJ databases. We finished clones according to the international finishing standard (http://genome.wustl.edu/Overview/g16stand.php). Of the 1,144 clones in the human chromosome 10 tiling path, 221 and 913 were finished at GENE and the Sanger Institute, respectively, and three elsewhere (Supplementary Fig. S1). The remaining seven clones show persistent deletion of internal fragments. In total, we finished 131,666,441 base pairs (bp) in 18 sequence contigs; euchromatic coverage is estimated at 99.4%. Sequence accuracy was estimated as described in ref. 5 and found to exceed 99.99%. The sequence assembly comprises all known chromosome 10 messenger RNAs (RefSeq set) and STS markers from available radiation hybrid6 and genetic maps7,8 (T. Furey, personal communication). In addition, the integrity of the sequence map was independently assessed at the University of California, Santa Cruz, by alignment of fosmid and BAC paired end sequences (http://genome.cse.ucsc.edu/). Table 1 lists the size of each sequence contig, with the largest one spanning 44,693,577 bp.
The gene and protein index
The Sanger Institute has established a standardized annotation pipeline (outlined at http://vega.sanger.ac.uk/) in which gene structures are drawn on the basis of human interpretation of the combined supportive evidence generated during sequence analysis. Annotation of the human chromosome 10 sequence resulted in a total of 1,787 gene structures that we then classified, as described in ref. 9, into: (1) 654 ‘known’ genes; (2) 162 ‘novel genes’; (3) 219 ‘novel transcripts’; (4) 322 ‘putative genes’; and (5) 430 ‘pseudogenes’. Pseudogenes were further subdivided into processed (371) and unprocessed (59).
Excluding the pseudogenes, human chromosome 10 is a chromosome with an average gene density (10.4 genes Mb-1). The 1,357 genes span 66,309,730 bp in total (mean 51,335 bp per gene). Therefore, 50.6% of the analysed sequence is transcribed, matching the figure reported for chromosome 22 (51%), which is gene-rich, but appearing elevated in comparison with chromosomes 6, 7, 14 and 20 (42.2%, 46.5%, 43.6% and 42.4%, respectively), which have gene densities similar to chromosome 10. The latter suggests that the human chromosome 10 genes have on average a larger genomic span than those on chromosomes 6, 7, 14 and 20. Gene size along human chromosome 10 varies enormously, with the two extremes being CTNNA3 (1,776,209 bp) and CALML5 (859 bp). Exons account for only 2.3% of the sequence and the mean exon size is 313 bp. The longest and shortest exons annotated in this study have a length of 9,763 (SH3MD1) and 3 bp (CDH23), respectively. CDH23 is also the gene with most exons (69) on this chromosome. Table 2 summarizes the features of each gene class.
Alternative splicing is a major contributor to the complexity of the human transcriptome. We annotated a total of 4,204 transcripts for 1,357 gene structures (Table 2). No splice variants were annotated on the basis of alternative polyadenylation sites. Approximately 73% of the protein-coding genes have more than one transcript and 5.8 on average. For ADD3 we annotated 22 variants. Note that the use of partial expressed sequences (for example, expressed sequence tags (ESTs)) may result in the annotation of more than one transcript per splice variant. Given this caveat, our analysis suggests a significantly higher level of alternative splicing compared with previous estimates10. Approximately 50% of the 3,456 transcripts (known and novel) do not seem to encode a protein. Annotation of these transcripts is largely based on ESTs, many of which may correspond to aberrant transcripts. Their precise role is largely unknown but they may be part of the machinery of transcriptional regulation (for example, via nonsense-mediated decay). Nevertheless, there are 1,837 transcripts with an open reading frame (ORF). Of the 342 genes with at least two transcripts having a complete ORF (that is, possess both a 5′ and a 3′ untranslated region (UTR)), 312 encode at least two distinct peptides.
Identification of transcription start sites and promoter regions remains a challenge in the annotation process. First, we scanned the human chromosome 10 unmasked sequence and predicted a total of 1,025 CpG islands (Fig. 1), which are known to be associated with the 5′ end of an estimated 60% of human genes11.We then used Eponine12 to predict transcription start sites and FirstEF13 to predict regions that encompass the promoter and 5′ exon. In total, FirstEF and Eponine predicted 1,801 (rank = 1, score ≥0.8) and 2,800 features, respectively (Supplementary Fig. S1). Notably, 62% of FirstEF and 96% of Eponine features directly overlapped CpG islands, suggesting a heavy bias towards this feature in both algorithms. The distribution of CpG islands, FirstEF and Eponine hits was also assessed relative to the first exon of each of the 4,635 annotated transcripts using a window of 5,000 bp upstream and 1,000 bp downstream of the exon. Table 3 summarizes the results obtained per gene class. For example, in the ‘known’ class, 1,544 (49.6%) and 1,124 (36.1%) transcripts had a FirstEF and Eponine feature, respectively. Not surprisingly, 89.9% of FirstEF and 96.7% of Eponine transcripts were also associated with a CpG island. Note that Eponine predicts multiple transcription start sites per transcript (4.24 on average).
Regulation of gene expression by antisense transcription is a recognized mechanism with examples reported in species ranging from bacteria to mammals14,15,16. We observed widespread occurrence of overlapping coding genes (either strand) in human chromosome 10 (101 pairs in total). In 38 cases one of the genes is fully contained within an intron of the other, typically on the opposite strand (34). For example, the second intron of the splice variant LIPA-004 encompasses four members of the IFIT gene cluster (Supplementary Table S3 and Fig. S2) and appears to be transcribed from the same bidirectional promoter as IFIT5 (IFIT cluster member). Interestingly, the LIPA-001 and -002 variants do not overlap any IFIT gene, whereas LIPA-003 and LIPA-005 both overlap with IFIT2 and IFIT4 (Supplementary Fig. S2). Mutations in LIPA can cause Wolman and cholesteryl ester storage disease (Online Mendelian Inheritance in Man (OMIM) 278000). Among partially overlapping pairs (opposite strands), 34% involve the respective 5′ exons, which is indicative in each case of a bidirectional promoter. We also searched for non-coding transcripts located on the opposite strand of coding genes. There are 67 antisense transcripts overlapping 63 coding genes. The two most common patterns observed were intragenic with partial overlap of one exon, and partial overlap with the most 5′ exon of the coding gene. For ZNF32, we found two antisense transcripts (ZNF32OS1 and ZNF32OS2).
Finally, we looked at the distribution of known protein domains in both human chromosome 10 (this study) and the whole genome (Ensembl v.17.33.1) proteome using InterProScan. At the gene level, 70.6% of the human chromosome 10 peptides have at least one InterPro match with a Pfam domain and 32% are multidomain (1.37 distinct InterPros on average). Supplementary Table S2 shows the top 24 domains in chromosome 10 alongside their genome-wide ranking, suggesting that this chromosome is enriched in peptides with a lipase (IPR000734), aldo/keto reductase (IPR001395) or alpha/beta hydrolase (IPR000073) domain. BLASTP analysis (e-values <10-15) showed that all six genes encoding the peptides carrying the IPR001395 domain are clustered (AKR1C cluster) at position 4.8–5.3 Mb, whereas there are two lipase clusters, LIP (90.0–91.0 Mb; six members) and PNLIP (117.9–118.1 Mb; four members). In total, we found 42 gene clusters along human chromosome 10 (Supplementary Table S3).
Genomic landscape
The average G + C and repeat content of human chromosome 10 are 41.58% and 43.66%, respectively. The distribution of the main classes of repeats (Supplementary Table S4), the G + C and CpG density plots all seem to follow the known genome-wide trends. For example, the G + C content fluctuates along the chromosome, peaks at the qtel and is positively correlated to gene density (Supplementary Fig. S1). Large genes tend to be located adjacent to or within gene-poor regions, for example PRKG1 and PCDH15 (interval 52.0–59.0 Mb), whereas regions of high gene density seem enriched in short interspersed elements (SINEs).
Segmental duplications are an important feature of the genomic landscape, being an integral part of the evolutionary machinery. Figure 2a illustrates the interchromosomal duplications along human chromosome 10 (see also ref. 17 and http://humanparalogy.cwru.edu/SDD). Figure 2b shows the extensive segmental duplications within a 5 Mb region at 10q11 with sequences further dispersed at 10q22, 10q23.1 and 10q23.2. Using the draft genome sequence Crosier and colleagues18 reported that 10q11 has been subject to multiple rounds of local duplications in at least the last 30–40 million years (Myr); they also showed that a 10q11:10q23 paracentric inversion occurred after the divergence of orang-utan from other great apes and hypothesized that the 10q22 locus resulted from chromosome-specific duplicative transposition. Bryce and colleagues19 characterized three cathepsin-L-like paralogues, which are expressed pseudogenes, and reported FISH signals at 10q11, 10q22 and 10q23, a pattern previously seen with the GLUD paralogues20 (Fig. 2b). The duplication of the CTSL locus between chromosomes 9 and 10 was estimated to have occurred some 40 Myr ago19. We identified four and three additional CTSLL and GLUD pseudogenes, respectively (Fig. 2b), consistent with the local duplication events involving BMS1L (known as KIAA0187)18 and CTGLF1 (this study). In total, we annotated 7 BMS1L and 11 CTGLF1 paralogues (Fig. 2b; functional genes in dark blue). Retroposition of a truncated KIAA1099 (CENTG2; chromosome 2) mRNA gave rise to a processed pseudogene on chromosome 10 (ref. 18). However, this pseudogene forms the 3′ exon of CTGLF1, suggesting that this gene resulted from a fusion between an ancestral gene and the CENTG2 pseudogene, and the retroposition event predated all segmental duplications. Note that there is also evidence of transcripts combining exons of CTGLF1 and BMS1L paralogues. Interestingly, we predicted a novel gene, FAM25A (based on mouse complementary DNA AK008614 and without similarity to any known protein), with seven human chromosome 10 paralogues (Fig. 2b) that follow the pattern of GLUD, BMS1L, CTGLF and CTSLL. In addition to these five types of low copy repeats, the segmental duplications in 10q11:10q22:10q23 seem to have impacted on the number of genes on this chromosome. Notably, 31% of all the functionally related gene clusters are located within 10q11 and 10q23 (Supplementary Fig. S1 and Table S3).
The average recombination rate across the chromosome is 1.32 cM Mb-1 (Fig. 3). Note that ref. 7 used the draft human chromosome 10 sequence (inflated by ∼8%) and thus obtained a lower figure (1.21 cM Mb-1). The rate of male recombination is higher than the female rate near the telomeres, whereas between D10S211 and D10S575 the female rate is higher (Fig. 3). This comparison also indicates the presence of two female-specific recombination hotspots (Fig. 3, arrows). As expected, pericentromeric regions display a low rate of recombination, more than twofold below the chromosome average. In particular, the region between D10S1247 and D10S1783 has a rate of 0.3 cM Mb-1 and contains the only human chromosome 10 recombination ‘desert’21. The region of extensive segmental duplications at 10q11 shows a low rate of recombination (0.72 cM Mb-1). Thus, meiotic recombination is unlikely to have been the driving force in generating these duplications. Our analysis confirmed the recombination ‘jungle’ between D10S1782–D10S1651, which we extended to D10S212 (3.4 cM Mb-1); we refuted the one between D10S189 and D10S1728 (2.3 cM Mb-1), and identified a new one between D10S1154 and D10S552 (4.55 cM Mb-1).
The position of each genetic marker on the female, male and sex-averaged genetic map is indicated. Female-specific recombination hotspots are indicated by thick arrows (left, D10S1732–D10S208; right, D10S599–D10S676). The location of markers flanking recombination deserts (blue arrows) and jungles (black arrows) is shown. The asterisk indicates the location of the refuted jungle (D10S189–D10S1728).
Comparative sequence analysis
Many of the functional units in present-day vertebrate genomes have been conserved through evolution. Coding regions are highly conserved across all vertebrates, whereas non-coding regions are conserved among more closely related species. The genome sequences of three fishes (Tetraodon, Fugu and zebrafish) and two rodents (mouse and rat) were publicly available at the time of analysis. We compared the sequence of human chromosome 10 to each of the above species and searched for conserved regions with coding potential—distinguishing functional non-coding elements above background sequence conservation requires the use of additional genomes22. We then correlated the obtained hits in each species with the annotated genes. As expected, each of the fish genomes showed higher specificity (> 0.82; that is, 82% of sequences conserved in fish overlapped human annotated exons) than the rodents (> 0.29), whereas the highest specificity was obtained using all genomes together (0.96). Sensitivity was higher using a rodent genome (0.7; that is, 70% of all annotated exons had matches in rodent) than a fish genome (0.4).
Of the 1,787 gene structures (16,765 exons), 84% (78% of exons) had at least one exon supported by a conserved region in one of the other genomes and 52% (32% of exons) in all genomes. Note that the figures given for exons are underestimated owing to lower sequence conservation in the untranslated regions. Protein-coding genes are highly conserved (98.5%; 85% of exons). In contrast, 61% (29% of exons) and 45% (27% of exons) of novel transcripts and putative genes, respectively, have at least one match. Furthermore, only 21% of novel transcripts and 8% of putative genes show conservation with a fish (91% for protein-coding genes). Typically, novel transcripts were annotated on the basis of solid experimental evidence (that is, human mRNA) and may represent either genes that have evolved more rapidly or non-coding RNAs.
On the basis of specificity, regions conserved in all six species can serve as a measure of completeness of the gene annotation process that occurred independently of the comparative analysis. We found 5,604 such evolutionarily conserved regions (ECRs) of which 5,292 mapped inside annotated exons (including pseudogenes). Of the remaining 312 ECRs, 142 were intergenic and 170 intragenic. On inspection, we found 79 of these ECRs with supportive evidence to annotate a missed exon, most of which were part of a pseudogene (79%). The remaining 233 ECRs provide the basis to estimate that we have annotated at least 95.8% of all conserved coding exons on human chromosome 10. This is a conservative estimate as 131 of these ECRs are located in introns and may represent conserved non-exonic sequences. Interestingly, 54 (41%) of them are associated with just four genes: C10orf11 (26; also known as CDA017), EBF3 (20), TCF7L2 (5) and PAX2 (3). All but C10orf11 (unknown function) are transcription factors. Figure 4 shows a MultiPipMaker23 alignment of the orthologous EBF3 loci and the relative position of ECRs in the human gene. Note that sequence identity is often higher in ECRs than in exons.
The human early B-cell factor 3 (EBF3) gene is represented by the arrow at the top. Alignments are displayed using MultiPipMaker23. In the top panel, the first track shows the location of the ECRs (blue lines) across the human locus, whereas the following four tracks show regions conserved in mouse, rat, Fugu and zebrafish, respectively (green, aligned regions; orange, aligned regions with at least 70% nucleotide identity and no gap over 100 bp). The bottom panel shows a detailed view of the three regions with the highest number of ECRs. Vertical black and grey numbered boxes represent coding and UTR exons, respectively. The scale at the right indicates the percentage of sequence identity. Physical distance is given in kilobases (kb).
Sequence variation
During the human chromosome 10 project we discovered 35,882 single nucleotide polymorphisms (SNPs) by sequence alignment in regions of clone overlaps. In total, we mapped 143,364 SNPs (dbSNP release 115) to the chromosome 10 sequence. Supplementary Fig. S1 shows the density plots for randomly discovered24 and all SNPs across the chromosome.
There are 5,864 (4.1%) exonic and 65,973 (46%) intronic SNPs. Of the 1,821 SNPs in coding exons 984 are non-synonymous. MSMB has the most polymorphic coding region with 43 SNPs kb-1; it encodes a protein with inhibin-like activity and its expression is decreased in prostate cancer25.
We also considered 729,553 human–chimpanzee single base differences (SBDs) remapped on the current assembly of human chromosome 10. These were high-confidence sequence differences originally identified by aligning 14 million shotgun reads of the chimpanzee genome, generated jointly by the Whitehead Institute and Washington University Genome Centers, to the human genome sequence assembly (NCBI build 31). We first removed all human–chimpanzee SBDs that co-localized with known human SNPs. Supplementary Fig. S1 shows the density plot of the remaining 703,338 SBDs. Of those, 55.3% are intergenic, 42.9% intronic and 1.8% exonic. The highest density of human–chimpanzee SBDs, fourfold greater than the average level, was observed in a 200-kb gene-poor region at 19.43–19.63 Mb. We then examined the 12,710 human–chimpanzee SBDs that lie in exons of the 816 human coding genes. Of those, 3,972 were in coding regions and can be subdivided further into 2,273 synonymous, 1,678 non-synonymous and 21 nonsense with respect to the human sequence. For each gene we calculated the rate of evolution of non-synonymous (Ka) and synonymous (Ks) changes, and the ratio Ka/Ks, which provides a measure of evolutionary selection. Supplementary Table S5 lists the 1,413 transcripts with at least one coding human–chimpanzee SBD sorted on the Ka/Ks value. There are only 29 transcripts (21 genes) that have a Ka/Ks value ≥1, whereas there are 484 without non-synonymous SBDs. Note that several caveats apply in this type of analysis owing to the incomplete nature of both the chimpanzee data and the list of human SNPs; we used the number of intronic human–chimpanzee SBDs per base in comparison to the chromosome average of 0.005 as a possible estimate of coverage. The gene with most non-synonymous human–chimpanzee SBDs is MKI67, an antigen identified by monoclonal antibody Ki-67, which appears to be fast evolving in humans (Ka/Ks = 1.038507; SNP data). The expression pattern of MKI67 in gastric and other cancers is under investigation as this gene is expressed in proliferating cells. Interestingly, a nonsense human–chimpanzee SBD is present in both of its coding transcripts. Among the 21 genes with nonsense human-chimpanzee SBDs notable examples are the serotonin receptor HTR7 (the neurotransmitter serotonin is thought to be involved in cognition and behaviour), PSAP (prosaposin; involved in variant Gaucher's disease and metachromatic leukodystrophy) and the developmental gene NODAL.
Medical implications
At the time of writing there were 85 disease loci reported on human chromosome 10 (http://www.ncbi.nlm.nih.gov/omim/), a 47% increase since 1999 (ref. 26). Several of these loci account for multiple disease phenotypes caused by mutations in the same gene; notable examples are FGFR2 (OMIM 176943), PTEN (OMIM 601728) and the proto-oncogene RET (OMIM 164761). Since PTEN was first shown to be mutated in brain, breast and prostate cancers27, there has been an explosion of reported mutations (110 germline and 332 somatic mutations)28 and disease phenotypes29. Human chromosome 10 harbours several other genes involved in tumorigenesis; for example, deregulation of TLX1, NFKB2 or BMI1 caused by chromosomal translocations or amplifications has been detected in lymphatic neoplasms. Mapping of allelic imbalances and functional studies suggest the presence of additional tumour-related genes. The finished and annotated sequence is key in the process of cloning these and other hitherto unidentified disease-associated genes.
The prompt release of both the clone and sequence map resources throughout the project has accelerated the cloning of many disease-causing genes. To this end we recently showed as part of the European ADLTE consortium that mutations in the LGI1 gene cause autosomal dominant lateral temporal epilepsy30. Notably, we found that the FRA10A folate-sensitive fragile site is located close to LGI1 and its expression is associated with the expansion of a polymorphic CGG repeat located at the 5′ UTR of FRA10AC1, a gene encoding a novel nuclear protein31. There are seven fragile sites mapping to human chromosome 10 (ref. 32).
The challenge ahead is to unravel the molecular basis of common disease. An increasing number of susceptibility loci for complex diseases is being mapped to human chromosome 10, including metabolic diseases such as type I diabetes (IDDM10 and IDDM17), psychiatric disorders such as schizophrenia, or neurodegenerative illnesses such as Alzheimer's disease26,32. In a case control study of morbidly obese and healthy individuals Boutin and colleagues33 identified a SNP in the GAD2 gene that increases the risk for obesity as well as a protective haplotype. Studies so far have mainly focused on candidate genes. The construction of a comprehensive haplotype map of the human genome is well underway34, making it possible to undertake a systematic approach to scanning the genome for associations to disease-related and other phenotypes.
Methods
The mapping and sequencing methods used in the assembly of the bacterial clone and sequence map of chromosome 10, respectively, as well as the tools of the gene annotation pipeline are described in refs 1 and 35 (see ref. 36 for detailed protocols). Manual annotation of gene structures followed the guidelines agreed in the human annotation workshop (HAWK; http://www.sanger.ac.uk/HGP/havana/hawk.shtml), whereas gene symbols were approved where possible by the HUGO Gene Nomenclature Committee37. Protein translations were analysed with InterProScan (http://www.ebi.ac.uk/interpro/scan.html), which was run via the Ensembl protein annotation pipeline, to obtain Pfam, Prosite, Prints and Profiles domain matches.
Alignments for inter- and intrachromosomal duplications were performed with WU-BLASTN (http://blast.wustl.edu) using the current sequence assembly of chromosome 10 and the NCBI34 build for the rest of the genome. All sequences were repeat masked with RepeatMasker (http://repeatmasker.genome.washington.edu) and low-quality alignments (e-value >10-30, sequence identity <90%, length <80 bp) were discarded. For intrachromosomal duplications, self matches were discarded. For interchromosomal duplications, the sequence was split into 400-kb segments. Adjacent matches in the same orientation were joined together as described by ref. 38. Only blocks of 10 kb or greater were retained.
The following sequence assemblies were used for comparative analysis: M. musculus NCBI build 30 (Mouse Genome Sequencing Consortium; http://www.ensembl.org/Mus_musculus/resources.html); R. norvegicus version 2.0 (Rat Genome Sequencing Consortium; http://www.hgsc.bcm.tmc.edu/projects/rat); D. rerio Assembly version 1 (Sanger Institute; http://www.sanger.ac.uk/Projects/D_rerio); F. rubripes version 2 (International Fugu Genome Consortium; http://www.fugu-sg.org/project/info.html); and T. nigroviridis version 6 (Genoscope and Whitehead Institute for Genome Research; http://www.genoscope.cns.fr/externe/tetraodon/Ressource.html). The repeat-masked sequence of chromosome 10 was aligned to the mouse and rat genome sequences using BLASTZ39 and the resulting matches were post-processed by axtBest and subsetAxt (W. J. Kent; http://www.soe.ucsc.edu/~kent/src) as described elsewhere35. Alignment of the three fish genome sequences to chromosome 10 was performed with WU-TBLASTX using the scoring matrix, parameters and filtering strategy applied by Exofish40. Overlapping alignments to different sequences were merged to produce contiguous regions of sequence conservation, analogous to the ECRs or ‘Ecores’, reported by Exofish.
SNPs in sequence overlaps were identified using a modification of the SSAHA software41. The chromosome 10 SNPs (dbSNP release 115) were mapped to the sequence assembly of this chromosome (this study) first with SSAHA and then Cross-Match. Of the approximately 14 million chimpanzee reads (http://www.genome.gov/11008056) mapped onto the human sequence assembly (NCBI build 31), those mapping to chromosome 10 were remapped to our sequence assembly and used to identify human–chimpanzee SBDs.
References
Bentley, D. R. et al. The physical maps for sequencing human chromosomes 1, 6, 9, 10, 13, 20 and X. Nature 409, 942–943 (2001)
Guy, J. et al. Genomic sequence and transcriptional profile of the boundary between pericentromeric satellites and genes on human chromosome arm 10p. Genome Res. 13, 159–172 (2003)
Guy, J. et al. Genomic sequence and transcriptional profile of the boundary between pericentromeric satellites and genes on human chromosome arm 10q. Hum. Mol. Genet. 9, 2029–2042 (2000)
Jackson, M. S., See, C. G., Mulligan, L. M. & Lauffart, B. F. A 9.75-Mb map across the centromere of human chromosome 10. Genomics 33, 258–270 (1996)
Felsenfeld, A., Peterson, J., Schloss, J. & Guyer, M. Assessing the quality of the DNA sequence from the Human Genome Project. Genome Res. 9, 1–4 (1999)
Deloukas, P. et al. A physical map of 30,000 human genes. Science 282, 744–746 (1998)
Kong, A. et al. A high-resolution recombination map of the human genome. Nature Genet. 31, 241–247 (2002)
Broman, K. W., Murray, J. C., Sheffield, V. C., White, R. L. & Weber, J. L. Comprehensive human genetic maps: individual and sex-specific variation in recombination. Am. J. Hum. Genet. 63, 861–869 (1998)
Deloukas, P. et al. The DNA sequence and comparative analysis of human chromosome 20. Nature 414, 865–871 (2001)
International Human Genome Sequencing Consortium, Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
Antequera, F. & Bird, A. Number of CpG islands and genes in human and mouse. Proc. Natl Acad. Sci. USA 90, 11995–11999 (1993)
Down, T. A. & Hubbard, T. J. Computational detection and location of transcription start sites in mammalian genomic DNA. Genome Res. 12, 458–461 (2002)
Davuluri, R. V., Grosse, I. & Zhang, M. Q. Computational identification of promoters and first exons in the human genome. Nature Genet. 29, 412–417 (2001)
Wagner, E. G. & Simons, R. W. Antisense RNA control in bacteria, phages, and plasmids. Annu. Rev. Microbiol. 48, 713–742 (1994)
Eddy, S. R. Non-coding RNA genes and the modern RNA world. Nature Rev. Genet. 2, 919–929 (2001)
Kiyosawa, H. et al. Antisense transcripts with FANTOM2 clone set and their implications for gene regulation. Genome Res. 13, 1324–1334 (2003)
Bailey, J. A. et al. Recent segmental duplications in the human genome. Science 297, 1003–1007 (2002)
Crosier, M. et al. Human paralogs of KIAA0187 were created through independent pericentromeric-directed and chromosome-specific duplication mechanisms. Genome Res. 12, 67–80 (2002)
Bryce, S. D. et al. A novel family of cathepsin L-like (CTSLL) sequences on human chromosome 10q and related transcripts. Genomics 24, 568–576 (1994)
Deloukas, P., Dauwerse, J. G., Moschonas, N. K., van Ommen, G. J. & van Loon, A. P. Three human glutamate dehydrogenase genes (GLUD1, GLUDP2, and GLUDP3) are located on chromosome 10q, but are not closely physically linked. Genomics 17, 676–681 (1993)
Yu, A. et al. Comparison of human genetic and sequence-based physical maps. Nature 409, 951–953 (2001)
Thomas, J. W. et al. Comparative analyses of multi-species sequences from targeted genomic regions. Nature 424, 788–793 (2003)
Schwartz, S. et al. PipMaker—a web server for aligning two genomic DNA sequences. Genome Res. 10, 577–586 (2000)
Sachidanandam, R. et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409, 928–933 (2001)
Vanaja, D. K., Cheville, J. C., Iturria, S. J. & Young, C. Y. Transcriptional silencing of zinc finger protein 185 identified by expression profiling is associated with prostate cancer progression. Cancer Res. 63, 3877–3882 (2003)
Deloukas, P., French, L., Meitinger, T. & Moschonas, N. K. Report of the third international workshop on human chromosome 10 mapping and sequencing 1999. Cytogenet. Cell Genet. 90, 1–12 (2000)
Li, J. et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275, 1943–1946 (1997)
Bonneau, D. & Longy, M. Mutations of the human PTEN gene. Hum. Mutat. 16, 109–122 (2000)
Eng, C. PTEN: one gene, many syndromes. Hum. Mutat. 22, 183–198 (2003)
Morante-Redolat, J. M. et al. Mutations in the LGI1/Epitempin gene on 10q24 cause autosomal dominant lateral temporal epilepsy. Hum. Mol. Genet. 11, 1119–1128 (2002)
Sarafidou, T. et al. Folate-sensitive fragile site FRA10A is due to an expansion of a CGG-repeat in a novel gene FRA10AC1, encoding a nuclear protein. Genomics (in the press)
Moschonas, N. K. in Encyclopedia of the Human Genome Vol. 1, 618–625 (Nature Publishing Group, London, 2003)
Boutin, P. et al. GAD2 on chromosome 10p12 is a candidate gene for human obesity. PLoS Biol. 1, 1–11 (2003)
The International HapMap Consortium. The International HapMap project. Nature 426, 789–796 (2003)
Mungall, A. J. et al. The DNA sequence and analysis of human chromosome 6. Nature 425, 805–811 (2003)
Dunham, I. (ed.) Genome Mapping and Sequencing (Horizon Scientific, Wymondham, UK)
Wain, H. M., Lush, M., Ducluzeau, F. & Povey, S. Genew: the human gene nomenclature database. Nucleic Acids Res. 30, 169–171 (2002)
Cheung, J. et al. Genome-wide detection of segmental duplications and potential assembly errors in the human genome sequence. Genome Biol. 4, R25 (2003)
Schwartz, S. et al. Human–mouse alignments with BLASTZ. Genome Res. 13, 103–107 (2003)
Roest Crollius, H. et al. Estimate of human gene number provided by genome-wide analysis using Tetraodon nigroviridis DNA sequence. Nature Genet. 25, 235–238 (2000)
Ning, Z., Cox, A. J. & Mullikin, J. C. SSAHA: a fast search method for large DNA databases. Genome Res. 11, 1725–1729 (2001)
Acknowledgements
We thank the Fugu, Mouse and Rat Genome Sequencing Consortia; Genoscope and Whitehead Institute for Genome Research (Tetraodon genome data); the members of the zebrafish project at the Sanger Institute; the groups at the Washington University Genome Sequencing Center and the Whitehead Institute for early access to the chimpanzee whole-genome shotgun data; the laboratories of H. Riethman and M. S. Jackson for the provision of telomeric and pericentromeric clones, respectively; H. M. Wain, E. A. Bruford, C. C. Talbot, V. K. Khodiyar, M. J. Lush, M. W. Wright and S. Povey for assigning official gene symbols; the EMBL, GenBank and Ensembl database groups; I. Dunham for critical reading of the manuscript; and the Wellcome Trust for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Table 1
Bacterial clones in the chromosome 10 tiling path. (DOC 31 kb)
Supplementary Table 2
Most common InterPro domains in the chromosome 10 proteome. (DOC 43 kb)
Supplementary Table 3
Gene clusters on chromosome 10. (DOC 84 kb)
Supplementary Table 4
Interspread repeats. (DOC 36 kb)
Supplementary Table 5
Human transcripts with human-chimpanzee Single Base Differences (h-cSBDs). (DOC 3320 kb)
Supplementary Figure 1
The human chromosome 10 sequence and its features. Detailed view of all map and sequence features (tile path, G+C, CpG, Repeats, SNPs, Conserved regions, Gene families, Gene structures). (PDF 11747 kb)
Supplementary Figure 2
Overlapping genes on opposite stands. The LIPA gene and the IFIT gene cluster on chromosome 10. (PDF 176 kb)
Rights and permissions
About this article
Cite this article
Deloukas, P., Earthrowl, M., Grafham, D. et al. The DNA sequence and comparative analysis of human chromosome 10. Nature 429, 375–381 (2004). https://doi.org/10.1038/nature02462
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02462
This article is cited by
-
Autosomal recessive congenital cataract is associated with a novel 4-bp splicing deletion mutation in a novel C10orf71 human gene
Human Genomics (2023)
-
A perspective on molecular signalling dysfunction, its clinical relevance and therapeutics in autism spectrum disorder
Experimental Brain Research (2022)
-
Intellectual disability: dendritic anomalies and emerging genetic perspectives
Acta Neuropathologica (2021)
-
Genetic association and functional analysis of rs7903456 in FAM35A gene and hyperuricemia: a population based study
Scientific Reports (2018)
-
Homeobox NKX2-3 promotes marginal-zone lymphomagenesis by activating B-cell receptor signalling and shaping lymphocyte dynamics
Nature Communications (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.