Abstract
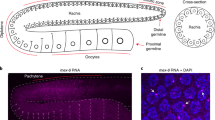
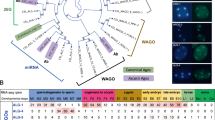
In many organisms, introducing double-stranded RNA (dsRNA) causes the degradation of messenger RNA that is homologous to the trigger dsRNA—a process known as RNA interference. The dsRNA is cleaved into short interfering RNAs (siRNAs), which hybridize to homologous mRNAs and induce their degradation1. dsRNAs vary in their ability to trigger RNA interference: many mRNA-targeting dsRNAs show weak phenotypes, and nearly all mRNAs of the Caenorhabditis elegans nervous system are refractory to RNA interference2,3,4. C. elegans eri-1 was identified in a genetic screen for mutants with enhanced sensitivity to dsRNAs. Here we show that eri-1 encodes an evolutionarily conserved protein with domains homologous to nucleic-acid-binding and exonuclease proteins. After exposure to dsRNA or siRNAs, animals with eri-1 mutations accumulate more siRNAs than do wild-type animals. C. elegans ERI-1 and its human orthologue degrade siRNAs in vitro. In the nematode worm, ERI-1 is predominantly cytoplasmic and is expressed most highly in the gonad and a subset of neurons, suggesting that ERI-1 siRNase activity suppresses RNA interference more intensely in these tissues. Thus, ERI-1 is a negative regulator that may normally function to limit the duration, cell-type specificity or endogenous functions of RNA interference.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hannon, G. J. RNA interference. Nature 418, 244–251 (2002)
Kamath, R. S. et al. Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 421, 231–237 (2003)
Timmons, L., Court, D. L. & Fire, A. Ingestion of bacterially expressed dsRNAs can produce specific and potent genetic interference in Caenorhabditis elegans. Gene 263, 103–112 (2001)
Tavernarakis, N., Wang, S. L., Dorovkov, M., Ryazanov, A. & Driscoll, M. Heritable and inducible genetic interference by double-stranded RNA encoded by transgenes. Nature Genet. 24, 180–183 (2000)
McIntire, S. L., Reimer, R. J., Schuske, K., Edwards, R. H. & Jorgensen, E. M. Identification and characterization of the vesicular GABA transporter. Nature 389, 870–876 (1997)
Beitel, G. J., Tuck, S., Greenwald, I. & Horvitz, H. R. The Caenorhabditis elegans gene lin-1 encodes an ETS-domain protein and defines a branch of the vulval induction pathway. Genes Dev. 9, 3149–3162 (1995)
Zuo, Y. & Deutscher, M. P. Exoribonuclease superfamilies: structural analysis and phylogenetic distribution. Nucleic Acids Res. 29, 1017–1026 (2001)
Li, Z., Pandit, S. & Deutscher, M. P. 3′ exoribonucleolytic trimming is a common feature of the maturation of small, stable RNAs in Escherichia coli. Proc. Natl Acad. Sci. USA 95, 2856–2861 (1998)
Ghosh, S. & Deutscher, M. P. Oligoribonuclease is an essential component of the mRNA decay pathway. Proc. Natl Acad. Sci. USA 96, 4372–4377 (1999)
Dominski, Z., Yang, X. C., Kaygun, H., Dadlez, M. & Marzluff, W. F. A 3′ exonuclease that specifically interacts with the 3′ end of histone mRNA. Mol. Cell 12, 295–305 (2003)
Kipp, M. et al. SAF-box, a conserved protein domain that specifically recognizes scaffold attachment region DNA. Mol. Cell. Biol. 20, 7480–7489 (2000)
Hamdan, S., Carr, P. D., Brown, S. E., Ollis, D. L. & Dixon, N. E. Structural basis for proofreading during replication of the Escherichia coli chromosome. Structure Fold. Des. 10, 535–546 (2002)
Zamore, P. D., Tuschl, T., Sharp, P. A. & Bartel, D. P. RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 101, 25–33 (2000)
Elbashir, S. M., Martinez, J., Patkaniowska, A., Lendeckel, W. & Tuschl, T. Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J. 20, 6877–6888 (2001)
Caplen, N. J., Parrish, S., Imani, F., Fire, A. & Morgan, R. A. Specific inhibition of gene expression by small double-stranded RNAs in invertebrate and vertebrate systems. Proc. Natl Acad. Sci. USA 98, 9742–9747 (2001)
Simmer, F. et al. Loss of the putative RNA-directed RNA polymerase RRF-3 makes C. elegans hypersensitive to RNAi. Curr. Biol. 12, 1317–1319 (2002)
Tabara, H. et al. The rde-1 gene, RNA interference, and transposon silencing in C. elegans. Cell 99, 123–132 (1999)
Tabara, H., Yigit, E., Siomi, H. & Mello, C. C. The dsRNA binding protein RDE-4 interacts with RDE-1, DCR-1, and a DExH-box helicase to direct RNAi in C. elegans. Cell 109, 861–871 (2002)
Winston, W. M., Molodowitch, C. & Hunter, C. P. Systemic RNAi in C. elegans requires the putative transmembrane protein SID-1. Science 295, 2456–2459 (2002)
Sijen, T. et al. On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 107, 465–476 (2001)
Gu, T., Orita, S. & Han, M. Caenorhabditis elegans SUR-5, a novel but conserved protein, negatively regulates LET-60 Ras activity during vulval induction. Mol. Cell. Biol. 18, 4556–4564 (1998)
Acknowledgements
We thank J. Roig, G. Lenz and J. Avruch for reagents and advice; A. Frand for critically evaluating our manuscript; and members of the Ruvkun laboratory for advice and discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
41586_2004_BFnature02302_MOESM2_ESM.pdf
Supplementary figure 2: eri-1 is differentially spliced. Northern blot analysis of RNA from wildtype and eri-1(mg366) eggs. (PDF 307 kb)
Rights and permissions
About this article
Cite this article
Kennedy, S., Wang, D. & Ruvkun, G. A conserved siRNA-degrading RNase negatively regulates RNA interference in C. elegans. Nature 427, 645–649 (2004). https://doi.org/10.1038/nature02302
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02302
This article is cited by
-
RNA interference-mediated knockdown of odorant-binding protein 2 and MP58 gene causes mortality in Myzus persicae
International Journal of Tropical Insect Science (2022)
-
Mating can initiate stable RNA silencing that overcomes epigenetic recovery
Nature Communications (2021)
-
Harnessing the power of genetics: fast forward genetics in Caenorhabditis elegans
Molecular Genetics and Genomics (2021)
-
RNAi technology for plant protection and its application in wheat
aBIOTECH (2021)
-
trans-Fatty acids facilitate DNA damage-induced apoptosis through the mitochondrial JNK-Sab-ROS positive feedback loop
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.